Spin Columns vs. Magnetic Beads: A Comprehensive Guide to ctDNA Extraction Methods
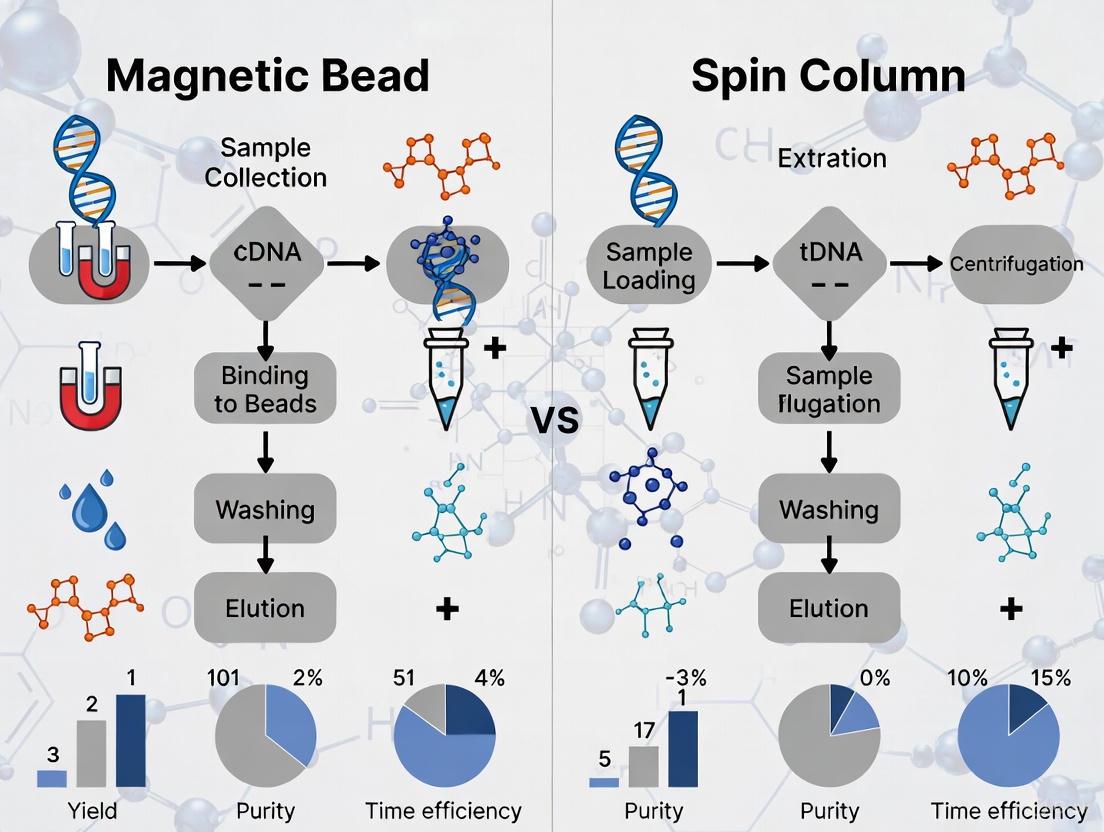
The choice between spin column and magnetic bead-based methods is critical for efficient circulating tumor DNA (ctDNA) extraction, directly impacting the sensitivity and reliability of downstream liquid biopsy applications.
Spin Columns vs. Magnetic Beads: A Comprehensive Guide to ctDNA Extraction Methods
Abstract
The choice between spin column and magnetic bead-based methods is critical for efficient circulating tumor DNA (ctDNA) extraction, directly impacting the sensitivity and reliability of downstream liquid biopsy applications. This article provides researchers, scientists, and drug development professionals with a foundational understanding of both techniques, detailing their methodological principles and practical workflows. It further offers guidance for troubleshooting and optimization, and presents a validated, comparative analysis of performance metrics—including yield, purity, scalability, and cost—to inform protocol selection for both research and clinical settings.
ctDNA Fundamentals: Biology and Extraction Principles
Circulating tumor DNA (ctDNA) refers to the fraction of cell-free DNA (cfDNA) in the bloodstream that originates directly from tumor cells. This complex biomarker is released into circulation through various mechanisms, including apoptosis, necrosis, and active release from tumor cells [1] [2]. In cancer patients, plasma obtained from a standard 10 mL blood draw typically contains approximately 5-10 ng/mL of ctDNA [1]. Understanding the fundamental biological properties of ctDNA—including its origin, fragment size, and circulation kinetics—is critical for developing effective liquid biopsy approaches. These intrinsic properties directly influence methodological decisions in ctDNA analysis, particularly in the comparison between magnetic bead-based and spin column-based extraction technologies [3] [4]. The preanalytical phase, including choice of extraction method, significantly impacts the yield, purity, and downstream analytical performance of ctDNA assays, making methodological comparisons essential for reliable liquid biopsy implementation in both research and clinical settings [3] [5] [6].
Core Biological Characteristics of ctDNA
Origin and Release Mechanisms
CtDNA originates from tumor cells through multiple release mechanisms that influence its characteristics. Apoptosis, or programmed cell death, represents a major release pathway that produces DNA fragments with a characteristic ladder-like pattern due to internucleosomal cleavage [2]. These apoptotic fragments typically display a peak size of approximately 167 base pairs, corresponding to the length of DNA wrapped around one nucleosome (147 bp) plus linker DNA [2] [7]. Necrosis provides another significant release mechanism, resulting from uncontrolled cell death in the adverse tumor microenvironment. Unlike apoptosis, necrosis produces larger, more randomly sized DNA fragments that can extend to many kilobases due to non-systematic DNA digestion [2]. Additionally, research suggests that viable tumor cells may actively release ctDNA through extracellular vesicles (such as exosomes) and other secretion mechanisms, although these pathways are less characterized [2] [7].
The relative contribution of each release mechanism varies depending on tumor type, location, and treatment status. Notably, tumors located behind anatomical barriers like the blood-brain barrier often show reduced ctDNA shedding, explaining the lower detection rates in primary brain cancers compared to other malignancies [7]. Furthermore, not all tumors shed detectable ctDNA into circulation, with some classified as "non-shedders" that present challenges for liquid biopsy approaches [8].
Fragment Size and Molecular Characteristics
CtDNA exhibits distinct fragmentation patterns that differentiate it from non-tumor cfDNA. While ctDNA fragments typically range from 70-200 base pairs [1], they are generally shorter than cfDNA derived from healthy cells. Specifically, ctDNA fragments average approximately 145 base pairs compared to the 166 bp average for non-tumor cfDNA [8]. This size difference arises from variations in nucleosome protection and DNA degradation processes between tumor and normal cells.
The fragment size distribution has profound implications for extraction efficiency and assay sensitivity. Smaller fragments (<100 bp) predominate in certain biological fluids like urine, where ctDNA must pass through renal filtration barriers [4] [2]. Recovery of these shorter fragments varies significantly between extraction methods, with some technologies demonstrating preferential capture of specific size ranges [4]. The variant allele frequency (VAF) of ctDNA is typically low, often falling below 1%, and is influenced by factors including cancer type, tumor stage, metabolic activity, and clearance rates [1] [8].
Half-Life and Clearance Dynamics
CtDNA demonstrates remarkably rapid clearance from circulation, with a half-life ranging from 16 minutes to 2.5 hours [1] [5] [7]. This brief window of detectability enables ctDNA to provide real-time, dynamic information about tumor status and treatment response [8]. Clearance occurs primarily through hepatic metabolism, renal excretion, and nuclease degradation in the bloodstream [7]. The rapid turnover rate means ctDNA levels can reflect changes in tumor burden almost in real time, offering a significant advantage over traditional imaging modalities that detect anatomical changes over longer timeframes [8] [7].
Table 1: Fundamental Biological Properties of ctDNA
| Characteristic | Properties | Biological Significance | Analytical Implications |
|---|---|---|---|
| Origin | Tumor cells via apoptosis, necrosis, active release [1] [2] | Reflects tumor biology and cell death mechanisms | Informs extraction strategy based on release mechanism |
| Typical Fragment Size | 70-200 bp; average ~145 bp (shorter than non-tumor cfDNA) [1] [8] | Indicates nucleosome protection and degradation history | Impacts extraction efficiency and sequencing library preparation |
| Half-Life in Circulation | 16 minutes - 2.5 hours [1] [5] | Enables real-time monitoring of tumor dynamics | Influences blood collection-to-processing timelines |
| Typical Concentration | 5-10 ng/mL from 10 mL blood draw [1] | Varies with tumor burden, stage, and location | Determines required plasma input volume and assay sensitivity |
Comparative Analysis of ctDNA Extraction Methods
The two primary technologies for ctDNA extraction—magnetic beads and spin columns—operate on different separation principles that impact their performance characteristics. Spin column-based methods utilize a silica membrane housed in a column that selectively binds DNA under high-salt conditions. After sample application and washing, DNA is eluted in a low-salt buffer [9]. This approach is valued for its simplicity, speed, and established protocols [9]. However, the binding capacity of silica membranes may limit recovery from samples with very low DNA concentrations, and these methods may demonstrate reduced efficiency for shorter DNA fragments [4] [9].
Magnetic bead-based methods employ superparamagnetic particles coated with a DNA-binding surface. When added to processed samples, DNA binds to the beads and is separated using a magnetic field, followed by washing and elution steps [9] [10]. This technology offers advantages in scalability, automation compatibility, and processing of large sample volumes [9]. The method has demonstrated effectiveness in microfluidic platforms for early cancer detection, with one study reporting extraction of approximately 5.7 ng of ctDNA from every 10 μL of plasma input [10]. Magnetic bead systems typically show higher binding capacity and potentially better recovery of low-abundance targets, making them suitable for the minimal DNA amounts often encountered in ctDNA analysis [9].
Performance Comparison: Yield, Purity, and Fragment Size Recovery
Direct comparisons of ctDNA extraction methods reveal significant differences in performance metrics. A 2020 study comparing three extraction kits (QIAamp CNA, Maxwell RSC, and Zymo Quick) found that the spin column-based CNA kit consistently yielded the highest total ccfDNA, while the magnetic bead-based RSC kit demonstrated higher variant allelic frequencies (VAFs) in mutation detection despite lower total DNA yields [3]. This suggests that magnetic bead methods may provide superior enrichment of the tumor-derived fraction relative to total background DNA [3].
Fragment size recovery represents another critical differentiator between technologies. Studies demonstrate that conventional silica-based methods (including many spin columns) show decreased recovery efficiency for fragments below 50-100 nucleotides [4]. This limitation particularly impacts urine ctDNA analysis, where fragments are often shorter than in plasma [4]. Magnetic bead systems may offer advantages in capturing these shorter fragments, though performance varies between specific kits and protocols. For example, the MagMAX kit showed minimal recovery of fragments below 80 nt, while specialized approaches like Q Sepharose or hybridization capture demonstrated superior recovery of shorter fragments (60-90% for fragments as short as 25 nt) [4].
Table 2: Direct Comparison of Magnetic Bead vs. Spin Column Extraction Methods
| Performance Metric | Magnetic Bead Methods | Spin Column Methods | Experimental Evidence |
|---|---|---|---|
| Total DNA Yield | Variable; often lower total yield but higher target enrichment [3] | Generally higher total DNA yields [3] | CNA kit (spin column) yielded highest total ccfDNA in 21 cancer patient samples [3] |
| Variant Allele Frequency | Higher VAFs reported despite lower total yield [3] | Lower VAFs observed despite higher total yield [3] | RSC kit (magnetic beads) showed higher VAF in 3 of 4 mutation-positive samples [3] |
| Short Fragment Recovery | Technology-dependent; some show better sub-100 bp recovery [4] | Often reduced recovery of fragments <50-100 bp [4] | QIAamp (spin column) showed minimal recovery of fragments <150 bp vs. specialized methods [4] |
| Automation Potential | High; easily adapted to high-throughput automated systems [9] [5] | Limited; primarily manual processing with some semi-automated options [9] | Automated magnetic bead extraction successfully applied to 649 plasma samples [5] |
| Sample Input Volume | Flexible; better suited for large volume processing [9] | Limited by column capacity; typically 2-4 mL plasma [3] [9] | Magnetic bead-based ME kit enabled 8 mL plasma input vs. 2 mL for CNA spin column [3] |
Experimental Protocols and Methodological Considerations
Standardized Workflow for Comparative Studies
Inter-laboratory comparisons require standardized protocols to ensure reproducible results. A 2020 technical report involving four Swiss laboratories established a robust framework for comparing ctDNA extraction and sequencing methods [6]. The recommended workflow begins with blood collection in preservative tubes (such as Streck, PAXgene, or Norgen) when immediate processing isn't possible, as these tubes maintain sample integrity for up to 168 hours [5]. For K2EDTA tubes, plasma should be separated within 60 minutes of collection to prevent genomic DNA contamination from leukocyte lysis [5].
The processing protocol involves double centrifugation (e.g., 1,900 × g for 15-20 minutes followed by 16,000 × g for 10 minutes) to efficiently remove cells and debris [5]. Extraction then proceeds using either magnetic bead or spin column systems with identical plasma input volumes. For the QIAamp CNA spin column kit, the protocol involves: (1) adding 2 mL plasma with proteinase K and buffer ACL, (2) incubating at 60°C, (3) adding ethanol and applying to column, (4) multiple wash steps, and (5) elution in 20-100 μL AVE buffer [3]. For the Maxwell RSC magnetic bead system, the automated protocol includes: (1) loading 1 mL plasma with DNase-free water, (2) automated lysis, binding to magnetic particles, washing, and (4) elution in 50-65 μL elution buffer [3].
Assessment Methods for Extraction Efficiency
Post-extraction analysis should employ multiple complementary methods to evaluate extraction efficiency. Fluorometric quantification (e.g., Qubit) provides total DNA concentration but cannot distinguish fragment sizes or amplifiability [5]. qPCR assays targeting both single-locus (e.g., 74 bp PDGFRA sequence) and multi-locus (e.g., 60 bp Alu repeats) sequences assess amplifiable DNA content [5]. The ratio between long (e.g., 445 bp FLI1) and short qPCR amplicons indicates potential genomic DNA contamination [5].
Fragment size analysis via parallel capillary electrophoresis (e.g., Fragment Analyzer, Bioanalyzer) provides detailed size distribution profiles, confirming the characteristic ~167 bp peak and detecting high molecular weight contamination [3] [5]. Finally, mutation-specific detection using droplet digital PCR (ddPCR) validates the recovery of tumor-derived fragments by quantifying mutant alleles in samples with known variants [3]. This multi-faceted assessment approach ensures comprehensive evaluation of both quantity and quality metrics across compared methods.
Research Reagent Solutions Toolkit
Table 3: Essential Research Reagents and Kits for ctDNA Extraction Studies
| Reagent/Kits | Specific Examples | Function/Purpose | Performance Notes |
|---|---|---|---|
| Spin Column Kits | QIAamp Circulating Nucleic Acid Kit (Qiagen) [3] | Silica-membrane based ctDNA extraction | Higher total DNA yield; standard input 2-4 mL plasma [3] |
| Magnetic Bead Kits | Maxwell RSC ccfDNA Plasma Kit (Promega) [3] | Magnetic particle-based automated extraction | Higher VAF despite lower total yield; amenable to automation [3] |
| Large Volume Kits | QIAamp MinElute ccfDNA Kit (Qiagen) [3] | Enables larger plasma input volumes (8 mL) | Processes larger input volumes from multiple blood collection tubes [3] |
| Blood Collection Tubes | Cell-Free DNA BCT (Streck) [5] | Preserves blood samples for delayed processing | Stable cfDNA yield for up to 168 hours; prevents genomic DNA release [5] |
| Quantification Assays | β-actin ddPCR size assay [3] | Measures amplifiable DNA across fragment sizes | Quantifies 137, 420, and 1950 bp fragments to assess integrity [3] |
| Size Analysis Systems | Fragment Analyzer / Bioanalyzer [3] [5] | Determines fragment size distribution | Confirms ctDNA size profile (~167 bp peak) and detects contamination [3] |
Integrated Workflow and Decision Framework
The following diagram illustrates the key decision points and methodological considerations for selecting and implementing ctDNA extraction methods based on research objectives and sample characteristics:
The biological nature of ctDNA—with its characteristic short fragment size, low abundance, and rapid clearance—presents both challenges and opportunities for liquid biopsy applications. The comparative analysis of magnetic bead versus spin column extraction technologies reveals a complex performance landscape where neither method universally outperforms the other across all metrics. Rather, the optimal choice depends on specific research requirements: magnetic bead systems demonstrate advantages in automation potential, processing scalability, and target enrichment (as evidenced by higher VAFs), while spin column methods typically provide higher total DNA yields with simpler manual protocols [3] [9]. This methodological comparison underscores the importance of aligning extraction technology with research objectives, whether prioritizing sensitivity for low-abundance mutations, throughput for large-scale studies, or practicality for resource-limited settings. As ctDNA analysis continues to evolve toward standardized clinical implementation, understanding these fundamental relationships between biomarker biology and extraction methodology will remain essential for advancing liquid biopsy applications in precision oncology.
The Critical Impact of Pre-analytical Variables on ctDNA Yield
Circulating tumor DNA (ctDNA), a fraction of cell-free DNA (cfDNA) shed into the bloodstream by tumor cells, has emerged as a transformative biomarker in oncology liquid biopsies. The analytical journey of ctDNA begins long before sequencing, with pre-analytical factors critically determining the success of downstream applications. Research demonstrates that pre-analytical variables introduce significant challenges, including false positives, false negatives, and substantial variability in tumor signal analysis [11]. This guide objectively compares the performance of the two primary ctDNA extraction technologies—spin columns versus magnetic beads—by synthesizing experimental data from recent studies to inform method selection for research and clinical applications.
Experimental Comparisons of ctDNA Extraction Methods
Multiple recent studies have systematically compared commercial cfDNA/ctDNA extraction kits to evaluate their performance across critical parameters. The following table summarizes key experimental designs from these investigations:
| Study Reference | Kits Compared | Sample Type | Key Assessment Parameters |
|---|---|---|---|
| PMC9601152 (2022) [12] | 6 kits (2 spin column, 4 magnetic bead-based, incl. 1 automated) | Plasma from healthy donors (n=10) | DNA yield (Qubit), fragment size distribution (Bioanalyzer), reproducibility |
| Scientific Reports (2018) [13] | 7 kits (3 spin column-based, 4 magnetic beads-based) | Pooled control plasma sample (n=10 replicates per kit) | LMW cfDNA yield (ddPCR), fragment size distribution, LMW fraction |
Performance Data: Yield, Fragment Size, and Reproducibility
The following table synthesizes quantitative results from direct kit comparisons, highlighting technology-specific performance differences:
| Performance Metric | Spin Column-based Kits | Magnetic Bead-based Kits | Statistical Significance & Notes |
|---|---|---|---|
| Total DNA Yield | Higher yields reported; QIAamp Circulating Nucleic Acid Kit showed highest recovery in one study [12] | Lower yields observed in multiple comparisons [12] | Significant differences observed (up to 4.3-fold variation between kits) [12] |
| LMW DNA Yield | Kit A: 1,936 GEs/mL plasma (median) [13] | Kit E: 1,515 GEs/mL plasma (median) [13] | Significant difference (p = 9.46 × 10−5) [13] |
| Fragment Size Profile | All kits isolated predominantly mono-nucleosomal fragments (~166 bp) [12] | All kits isolated predominantly mono-nucleosomal fragments (~166 bp) [12] | Magnetic beads may offer better recovery of very short fragments [14] |
| LMW Fraction | Kit A: 89% (median) [13] | Kit E: 90% (median) [13] | No significant difference in LMW fraction between technologies [13] |
| Reproducibility | High reproducibility for leading spin column kits [12] | Variable performance across different bead-based kits [12] | Automated systems (e.g., MagNA Pure) showed high reproducibility [12] |
Detailed Experimental Protocols from Cited Studies
Plasma Sample Processing Protocol (PMC9601152)
The following workflow details the standardized sample processing method used in the 2022 comparative study [12]:
Key Materials: Sarstedt S-Monovettes 9 mL K3E (containing 1.6 mg K3 EDTA/mL blood), 15 mL Falcon tubes (BD), 1.5 mL Eppendorf Safe-Lock tubes [12].
cfDNA Extraction and Quantification Methods
The 2022 study performed extractions from 1 mL plasma following manufacturer protocols, with elution volumes varying by kit (12-100 μL) [12]. DNA was quantified in duplicate using:
- Fluorometric quantification: Qubit Fluorometer 3.0 with dsDNA HS Assay
- Fragment sizing: Agilent 2100 Bioanalyzer with High-Sensitivity DNA Kit
The 2018 study employed a multiplexed droplet digital PCR (ddPCR) assay with 5 short (67-75 bp) and 4 long (439-522 bp) amplicons to precisely quantify amplifiable LMW DNA concentration and fragment size distribution [13].
Blood Collection Tube Comparison Protocol
A 2018 study compared blood collection protocols using matched samples from 23 healthy volunteers [13]:
- EDTA tubes: Processed within 1 hour of venipuncture
- Cell-free DNA Blood Collection Tubes (BCT): Processed at 24 hours and 72 hours post-collection
- All samples were processed identically with cfDNA extracted using QIAamp Circulating Nucleic Acid kit
Findings: No significant differences in cfDNA yield, fragment size, or background noise were observed between immediate processing (EDTA) and delayed processing with specialized tubes (BCT) [13].
Technology Comparison: Principles and Applications
Mechanism of Action
- Spin Columns: Utilize selective DNA binding to a silica membrane under high-salt conditions [9]
- Magnetic Beads: Employ magnetic particles coated with a DNA-binding surface, separated via magnetic field [9]
The Scientist's Toolkit: Essential Research Reagents
| Category | Specific Products | Function & Application Notes |
|---|---|---|
| Extraction Kits (Spin Column) | QIAamp Circulating Nucleic Acid Kit (Qiagen) [12] | High-yield cfDNA extraction; manual processing |
| NucleoSpin Plasma XS (Macherey-Nagel) [12] | Low input volume (240 μL); high-sensitivity protocol | |
| Extraction Kits (Magnetic Bead) | MagMAX Cell-Free DNA Isolation Kit (Thermo Fisher) [12] | Magnetic bead-based; suitable for automation |
| MagNA Pure 24 Total NA Isolation Kit (Roche) [12] | Automated system; high reproducibility | |
| Blood Collection Tubes | EDTA Tubes [13] | Require processing within 1-2 hours |
| Cell-free DNA Blood Collection Tubes (BCT) [13] | Contain preservatives; enable room temperature storage for up to 72 hours | |
| Quantification & QC | Qubit Fluorometer with dsDNA HS Assay [12] | Fluorometric quantification; highly sensitive for low concentrations |
| Agilent Bioanalyzer with High-Sensitivity DNA Kit [12] | Fragment size distribution analysis | |
| Multiplexed ddPCR Assay [13] | Simultaneously quantifies amplifiable DNA and fragment size |
The choice between spin column and magnetic bead technologies for ctDNA extraction involves careful consideration of research priorities. Spin column methods generally provide superior DNA yield, making them suitable for applications where maximum DNA recovery is critical. Magnetic bead systems offer advantages in automation, scalability, and potentially better recovery of shorter DNA fragments, ideal for high-throughput settings. Beyond the extraction method itself, standardization of pre-analytical conditions—from blood collection to processing protocols—is equally critical for obtaining reliable, reproducible ctDNA results. Researchers should align their selection with specific application requirements, sample volume constraints, and throughput needs while implementing rigorous standardization across all pre-analytical phases.
Core Principles of Silica-Based DNA Binding
The isolation of circulating tumor DNA (ctDNA) from blood plasma is a critical step in liquid biopsy workflows, with silica-based methods forming the technological cornerstone. This guide provides an objective, data-driven comparison of the two predominant silica-based platforms: magnetic beads and spin columns. By synthesizing recent experimental findings on DNA yield, fragment size bias, purity, and suitability for automation, this analysis aims to equip researchers and drug development professionals with the evidence necessary to select the optimal extraction method for their specific application in ctDNA research.
Solid-phase extraction using silica matrices is the most widely adopted method for nucleic acid purification. The core principle hinges on the affinity between the negatively charged phosphate backbone of DNA and a positively charged silica surface in the presence of chaotropic salts [15]. These high-molarity salts, such as guanidine hydrochloride, disrupt the hydrogen-bonding network of water, dehydrate the DNA and silica surfaces, and thereby facilitate binding by reducing the energetic barrier between the two negatively charged entities [15].
While all silica methods operate on this foundational chemistry, the physical implementation of the silica surface—as a packed membrane in a spin column or as microbeads in a suspension—introduces significant practical divergences. These differences profoundly impact the efficiency of recovering the short, fragmented ctDNA from a background of wild-type cell-free DNA, which is paramount for sensitive downstream applications like next-generation sequencing (NGS) and digital PCR (dPCR) [3] [16] [17].
Comparative Experimental Data: Magnetic Beads vs. Spin Columns
Independent studies have systematically evaluated the performance of various commercial kits based on these two technologies. The following tables summarize key quantitative findings from these comparative experiments.
Table 1: Comparison of DNA Yield and Mutant Detection Performance
| Extraction Kit (Technology) | Average DNA Yield (ng/mL plasma) | Performance in Mutant Detection | Variant Allelic Frequency (VAF) |
|---|---|---|---|
| QIAamp CNA Kit (Spin Column) | Significantly higher yield [3] | More mutant copies/mL in 2 of 4 samples [3] | Lower in 3 of 4 samples [3] |
| Maxwell RSC ccfDNA Kit (Magnetic Beads) | Lower yield than CNA kit [3] | More mutant copies/mL in 2 of 4 samples [3] | Higher in 3 of 4 samples [3] |
| QIAamp MinElute ccfDNA Kit (Spin Column) | Not specified | Similar integrity and mutant levels vs. CNA and RSC kits [3] | Higher VAF than CNA kit [3] |
Table 2: Comparison of Fragment Size Recovery and Purity
| Performance Metric | Spin Column (Membrane) | Magnetic Beads |
|---|---|---|
| Recovery of Short Fragments (<300 bp) | Good [3] | Superior efficiency [16] [17] |
| Recovery of Long Fragments (>600 bp) | Better suited for variable-sized DNA, particularly high molecular weight fragments [16] [17] | Less efficient for long fragments [18] |
| Co-Extraction of Inhibitors | Higher risk, leading to lower purity [18] | Decreased co-extraction, resulting in excellent purity [18] |
Detailed Experimental Protocols
To ensure reproducibility and provide context for the data, this section outlines the standard methodologies employed in the cited comparison studies.
Protocol for Magnetic Bead-Based Extraction
The Maxwell RSC ccfDNA Plasma Kit was used as a representative magnetic bead method in a comparison of extraction techniques from cancer patient plasma [3]. The general workflow for automated systems like the QIAsymphony SP or Maxwell RSC involves:
- Lysis and Binding: Plasma samples are incubated with a lysis buffer containing proteinase K to digest proteins and release DNA. A binding buffer containing chaotropic salts and isopropanol is added to create conditions for DNA adsorption onto the silica surface of the magnetic beads [5] [19].
- Capture and Washing: An external magnetic field is applied to immobilize the bead-DNA complexes. The supernatant is discarded, and the beads are washed multiple times with an ethanol-based wash buffer to remove salts, proteins, and other contaminants [19].
- Elution: The purified DNA is eluted from the beads using a low-ionic-strength elution buffer (e.g., Tris-EDTA or nuclease-free water) at an elevated temperature, which disrupts the DNA-silica interaction [19] [15].
Protocol for Spin Column-Based Extraction
The QIAamp CNA Kit was used as a representative spin column method in the same study [3]. The standard manual protocol is as follows:
- Lysis and Binding: Plasma is lysed with a buffer containing chaotropic salts. The mixture is applied to the spin column, and through brief centrifugation, the DNA binds to the silica membrane under high-salt conditions [3].
- Washing: The membrane is washed multiple times with wash buffers containing ethanol to remove impurities. Between washes, the column is centrifuged to remove the flow-through [3].
- Elution: The purified DNA is eluted in a low-ionic-strength buffer that hydrates the DNA and disrupts its interaction with the silica membrane. A second elution step or an extended incubation time can be used to maximize DNA recovery [3].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents and Kits for ctDNA Extraction
| Item | Function in Workflow | Examples |
|---|---|---|
| Blood Collection Tubes with Stabilizers | Prevent leukocyte lysis and genomic DNA contamination, enabling delayed processing. | Streck, PAXgene, CellSave [5] [16] [17] |
| Magnetic Bead-Based Kits | Automated, high-throughput recovery of short DNA fragments. | Maxwell RSC ccfDNA Plasma Kit, SafeCAP 2.0, NucleoMag beads [3] [19] [20] |
| Spin Column-Based Kits | Reliable, manual purification of DNA; effective for variable fragment sizes. | QIAamp Circulating Nucleic Acid Kit, QIAamp MinElute ccfDNA Kit [3] |
| Chaotropic Salt-Based Binding Buffer | Drives DNA adsorption onto the silica surface by dehydrating molecules and neutralizing charge. | Guanidine HCl, sodium perchlorate [15] |
| Ethanol-Based Wash Buffer | Removes contaminants and salts from the silica matrix without eluting bound DNA. | Standard component in most commercial kits [19] |
| Low Ionic-Strength Elution Buffer | Disrupts DNA-silica interaction to release purified DNA into solution. | Tris-EDTA (TE) buffer, nuclease-free water [15] |
Workflow and Decision Pathway
The following diagram illustrates the core operational workflow for both silica-based methods and a logical path for selecting the appropriate technology based on research objectives.
Discussion and Technical Considerations
The experimental data indicates a clear trade-off between total DNA yield and the quality of the recovered ctDNA fraction. While spin column kits like the QIAamp CNA often produce a higher overall DNA yield, magnetic bead-based methods can provide a superior variant allelic frequency (VAF), which is critical for detecting low-abundance mutations [3]. This suggests that spin columns may co-purify more non-target, high molecular weight genomic DNA, thereby diluting the ctDNA signal.
The superior performance of magnetic beads in recovering the short, mono-nucleosomal DNA fragments that are characteristic of ctDNA is attributed to their high surface-area-to-volume ratio and the efficient suspension mixing during binding, which enhances the capture kinetics of small fragments [16] [17]. Furthermore, magnetic bead systems are inherently more amenable to automation on platforms like the QIAsymphony or KingFisher, reducing hands-on time, minimizing cross-contamination risk, and improving inter-laboratory reproducibility—a significant advantage for clinical diagnostics and multi-center trials [19] [20] [6].
Conversely, spin columns remain a robust, cost-effective choice for laboratories with lower sample throughput or where the simultaneous recovery of a broader size range of DNA fragments is desired [18].
Future Perspectives
The field of ctDNA extraction continues to evolve. Emerging technologies include magnetic ionic liquids (MILs) and nanowire networks, which promise even higher enrichment factors and recovery efficiencies for ctDNA [16] [17]. Furthermore, integrated microfluidic devices that combine extraction with subsequent analysis steps are under active development, aiming to create fully automated "lab-on-a-chip" solutions for liquid biopsy [16] [17]. The ongoing optimization of magnetic bead chemistry, such as the development of superparamagnetic beads with specialized silica coatings, continues to push the boundaries of sensitivity and workflow efficiency [19] [20].
The analysis of circulating tumor DNA (ctDNA) has emerged as a cornerstone of liquid biopsy, enabling non-invasive cancer diagnosis, tumor profiling, and therapeutic monitoring [21] [22]. The efficacy of these advanced molecular applications is fundamentally dependent on the upstream extraction of cell-free DNA (cfDNA), where the choice of binding matrix—silica membranes or functionalized magnetic particles—critically influences yield, purity, and workflow efficiency [23] [22]. This guide provides an objective, data-driven comparison of these two dominant solid-phase extraction methods, framing the discussion within the broader context of optimizing ctDNA research and clinical diagnostics. We evaluate performance metrics, detail experimental protocols, and present key reagent solutions to inform researchers and drug development professionals in selecting the most appropriate extraction methodology for their specific applications.
Performance Comparison: Quantitative Data Analysis
The following tables consolidate experimental data from published studies to compare the performance of silica membrane and magnetic particle-based methods across key parameters.
Table 1: Comprehensive Performance Metrics for DNA Extraction Methods
| Performance Parameter | Silica Membrane (Spin Column) | Functionalized Magnetic Particles | Supporting Evidence |
|---|---|---|---|
| Typical Extraction Time | ~25 minutes [24] | 6–15 minutes [25] [24] | SHIFT-SP method; PIBEX chip |
| DNA Binding Efficiency | ~50% yield reported in some comparisons [24] | Up to 98.2% binding efficiency [24]; ~96% recovery with optimized beads [24] | Optimized pH and tip-based binding |
| Elution Volume Flexibility | Limited by membrane size | Highly flexible; suitable for low elution volumes [24] | Aids in obtaining high-concentration eluates |
| Automation Compatibility | Limited; mostly manual | Excellent; highly amenable to automation [9] [22] | High-throughput validated systems |
| Throughput & Scalability | Moderate; suited for batch processing | High; ideal for parallel processing and large volumes [9] [26] | SPRI bead use in 8-strip tubes |
| Sample Loss Risk | Higher due to transfer steps | Lower; minimal handling and no centrifugation [25] [26] | Integrated PIBEX workflow |
| Size Selection Capability | Limited resolution | Excellent; highly tunable via bead-to-sample ratio [26] | SPRI bead cleanup for sequencing |
Table 2: Application-Specific Performance in ctDNA Workflows
| Application Aspect | Silica Membrane (Spin Column) | Functionalized Magnetic Particles | Notes & Context |
|---|---|---|---|
| Average cfDNA Yield | Variable; lower yield in some reports [23] | Higher, more consistent recovery rates [23] [22] | QIAamp Circulating Nucleic Acid Kit outperformed others in a comparative study [23] |
| Purity (gDNA Contamination) | Low gDNA contamination reported [23] | Minimal gDNA contamination; consistent fragment profile [22] | Critical for downstream sequencing accuracy |
| Fragment Size Profile | Reproducible profile | Reproducible mononucleosomal/~167 bp peak [22] | Preserves native cfDNA characteristics |
| Variant Detection Accuracy | Suitable for ddPCR/NGS | High concordance with expected variants in reference materials [22] | Essential for reliable liquid biopsy |
| Hands-On Time | Significant | Drastically reduced, especially in automated formats [25] [26] | PIBEX chip completes extraction in 15 min [25] |
Experimental Protocols and Methodologies
Silica Membrane-Based Extraction: Standard Spin Column Protocol
The fundamental principle of this method is the selective binding of DNA to a silica membrane in the presence of high-concentration chaotropic salts, which disrupt hydrogen bonding and facilitate DNA adsorption to the silica surface [9].
- Step 1: Sample Lysis and Binding. The plasma sample is mixed with a lysis/binding buffer containing chaotropic salts (e.g., guanidine hydrochloride). The lysate is then transferred to the spin column and centrifuged. During this step, DNA binds to the silica membrane, while contaminants pass through [9].
- Step 2: Washing. The membrane is washed twice with a buffer containing ethanol to remove salts, proteins, and other impurities without eluting the bound DNA. Centrifugation is performed after each wash [9] [23].
- Step 3: Elution. The DNA is eluted in a low-salt buffer (e.g., Tris-EDTA or nuclease-free water). The column is centrifuged, and the flow-through contains the purified DNA. A common limitation is the incomplete recovery of the elution buffer due to surface tension within the membrane pores [25].
Innovative Modification: Centrifugation-Free Microfluidic Chip To address workflow limitations, a pressure and immiscibility-based extraction (PIBEX) microfluidic chip was developed. This method replaces centrifugation with vacuum pressure and uses mineral oil as an immiscible fluid to displace residual buffers from the silica membrane, enabling efficient, centrifugation-free extraction within 15 minutes [25].
Functionalized Magnetic Particle-Based Extraction: Optimized Protocol
This method relies on magnetic silica beads functionalized with specific ligands to enhance nucleic acid binding. The process is driven by magnetic manipulation rather than centrifugation.
- Step 1: Binding Optimization. The sample is mixed with lysis/binding buffer and functionalized magnetic beads. Key parameters for maximizing yield include:
- pH: A lower pH (e.g., 4.1) reduces the negative charge on silica, minimizing electrostatic repulsion with DNA and significantly enhancing binding efficiency (up to 98.2%) compared to a higher pH [24].
- Mixing Mode: A "tip-based" method (repeated aspiration and dispensing) exposes beads to the sample more effectively than orbital shaking, achieving ~85% binding within 1 minute versus ~61% with shaking [24].
- Step 2: Magnetic Separation and Washing. A magnetic field is applied to concentrate the beads against the tube wall. The supernatant is discarded, and the beads are washed with an ethanol-based buffer. The tube is not moved during this step to prevent bead loss.
- Step 3: Elution. After a drying step to remove residual ethanol, the purified DNA is eluted from the beads using a low-salt elution buffer. Higher elution temperatures (e.g., 70-80°C) can improve elution efficiency and yield [24].
Advanced Application: ctDNA Extraction in a Microfluidic Platform A simulated study detailed a microfluidic device for extracting ctDNA from early-stage cancer patients using superparamagnetic (SPM) beads. The workflow involved: (1) Microfiltration: Using a channel with varying widths (5 µm and 2 µm) to separate ctDNA from larger impurities and thrombocytes based on size. (2) Binding and Mixing: ctDNA is mixed with SPM beads in a curved microchannel to enhance binding. (3) Magnetic Separation: A permanent magnet isolates bead-bound ctDNA from the solution. This simulation reported an average yield of 5.7 ng per 10 µL of plasma input [27].
Visualization of Workflows
The following diagrams illustrate the core procedural and logistical differences between the two extraction methods.
Procedural Workflow Comparison
Throughput and Handling Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents and Kits for DNA Extraction
| Reagent / Kit Name | Type | Primary Function | Key Characteristic |
|---|---|---|---|
| QIAamp Circulating Nucleic Acid Kit [23] | Silica Membrane | Manual cfDNA extraction from plasma/serum | High yield and purity; used as a benchmark in comparative studies |
| QIAamp MinElute ccfDNA Kit [23] | Silica Membrane | Small-volume elution from plasma | Designed for concentrated elution from small sample volumes |
| QIAsymphony DSP Circulating DNA Kit [23] | Silica Membrane (Automated) | Automated extraction on QIAsymphony | Reproducible, hands-off workflow for clinical settings |
| VERSANT Sample Preparation Kit [24] | Magnetic Silica Beads | Nucleic acid extraction for molecular diagnostics | Optimizable Boom method protocol; basis for SHIFT-SP |
| Dynabeads Silane DNA Kit [27] | Magnetic Silica Beads | DNA purification from various samples | Used in microfluidic simulation studies for ctDNA binding |
| SHIFT-SP Method [24] | Magnetic Silica Beads | Rapid, high-yield nucleic acid extraction | Optimized for speed (6-7 min) and binding efficiency (>96%) |
| CUTANA Quick Cleanup DNA Purification Kit [26] | SPRI Beads | DNA purification & size selection for NGS | Tunable bead-to-sample ratio for selective fragment recovery |
| AcroMetrix Multi-analyte ctDNA Plasma Control [22] | Quality Control | Assessing extraction efficiency & assay performance | Contains variants at defined VAFs (0%, 0.1%, 0.5%, 1%) |
| Seraseq ctDNA Complete Reference Material [22] | Quality Control | Validation of variant detection post-extraction | Multiplexed variants across multiple genes for NGS validation |
The comparative data and protocols presented herein demonstrate a clear technological evolution from traditional silica membranes toward functionalized magnetic particles for ctDNA extraction, particularly in demanding applications like liquid biopsy.
Silica membranes offer a robust, well-established, and straightforward methodology. However, their reliance on centrifugation, lower potential recovery rates, and limited scalability can be significant drawbacks in high-throughput or resource-limited settings [24] [9] [23].
In contrast, functionalized magnetic particles provide superior speed, significantly higher binding and elution efficiencies, and unparalleled compatibility with automation and miniaturized microfluidic systems [25] [24] [22]. The ability to functionalize the magnetic core with specific ligands (e.g., amino silanes) or to engineer porous silica coatings further enhances their binding capacity and application specificity [28] [29]. This makes them the binding matrix of choice for laboratories focusing on maximizing recovery from precious low-concentration samples, such as ctDNA from early-stage cancer patients, and for developing integrated, sample-to-answer diagnostic systems [27] [22].
In conclusion, the choice between silica membranes and magnetic particles is not merely a matter of preference but a strategic decision. For routine processing of a limited number of samples, silica membranes remain a viable option. However, for advanced ctDNA research and clinical diagnostics—where yield, reproducibility, throughput, and integration into automated workflows are paramount—the evidence strongly supports the adoption of functionalized magnetic particle-based extraction systems.
Methodologies in Practice: Protocols and Workflow Integration
The success of modern molecular biology, especially in sensitive applications like circulating tumor DNA (ctDNA) analysis for liquid biopsy, hinges on the quality of the extracted nucleic acids. The purification process must efficiently isolate DNA from challenging samples where the target, such as ctDNA, is often present in very low concentrations amidst a background of normal cell-free DNA [30] [21]. Among the various techniques available, spin column-based nucleic acid purification has become a cornerstone methodology in research and clinical laboratories worldwide [31]. This guide provides a detailed, step-by-step explanation of the spin column protocol for centrifugation and elution, while objectively comparing its performance to the increasingly prevalent magnetic bead-based method. Understanding the principles, advantages, and limitations of each technique is crucial for researchers and drug development professionals to select the optimal approach for their specific experimental and diagnostic goals.
Core Principles and Comparison
The Science Behind Spin Columns
The spin column method operates on the principle of selective binding of nucleic acids to a silica-based membrane under specific chemical conditions. The process relies on the use of chaotropic salts, which are high-concentration ions that disrupt the hydrogen-bonding network of water and cellular components [31]. This action serves multiple purposes: it denatures proteins, inactivates nucleases, and, most importantly, allows DNA molecules to dehydrate and form hydrogen bonds with the silica surface in the spin column membrane [31]. Once bound, contaminants such as proteins, salts, and other cellular impurities are removed through a series of wash steps using buffers that do not disrupt the DNA-silica interaction. The final elution step uses a low-salt buffer or nuclease-free water to rehydrate the DNA, breaking the hydrogen bonds and releasing the pure nucleic acids into the collection tube [31].
Spin Column vs. Magnetic Bead Extraction
While spin columns utilize a silica membrane in a column format, magnetic bead-based extraction employs microscale magnetic particles (typically 0.5–5 µm) coated with a DNA-binding surface, often also silica [9] [30]. In this method, DNA from the lysed sample binds to the beads, and a magnetic field is applied to separate the bead-DNA complexes from the rest of the sample mixture. Wash steps remove impurities, and purified DNA is finally eluted from the beads [9]. The fundamental difference lies not in the binding chemistry, which is often similar, but in the method of separation—centrifugation versus magnetic capture.
Table: Core Principle Comparison of Spin Column and Magnetic Bead Methods
| Feature | Spin Column | Magnetic Beads |
|---|---|---|
| Binding Surface | Silica membrane | Silica-coated magnetic particles |
| Separation Mechanism | Centrifugation | Magnetic field |
| Binding Chemistry | Chaotropic salts facilitate hydrogen bonding to silica [31] | Chaotropic salts or homobifunctional crosslinkers facilitate binding to silica or amine-coated beads [9] [30] |
| Typical Process | Sequential, batch processing of individual columns | Can be processed in batch or in automated liquid handlers |
| Elution Principle | Rehydration with low-salt buffer/water breaks hydrogen bonds [31] | Change in buffer conditions or pH breaks binding interaction [30] |
Performance and Experimental Data
Quantitative Performance Comparison
Direct comparative studies provide valuable insights into the performance of these two methods, particularly for challenging samples like cell-free DNA. A 2022 study introduced a magnetic bead-based cfDNA extraction method using a homobifunctional crosslinker (DMS) and reported a 56% higher extraction efficiency compared to a commercial spin-column kit (QIAamp kit) [30]. Furthermore, this magnetic bead method successfully extracted cfDNA from plasma within a rapid 10-minute processing time, highlighting a significant potential advantage in both yield and speed [30].
A larger 2025 study in Scientific Reports evaluating automated cfDNA extraction metrics analyzed 649 blood plasma samples and underscored that pre-analytical factors, including the choice of extraction method, are fundamental for accurate interpretation of results [5]. The study emphasized that the amount of cfDNA is generally limited and many downstream approaches require the assessment of individual molecules, making extraction yield and purity critical [5].
Table: Experimental Performance Data for cfDNA Extraction
| Parameter | Spin Column (QIAamp Kit) | Magnetic Beads (DMS Method) | Notes |
|---|---|---|---|
| Extraction Efficiency | Baseline | 56% higher than spin column [30] | Comparison based on yield from blood plasma |
| Processing Time | Varies by protocol (~30+ minutes) | ~10 minutes [30] | Magnetic method leverages instant binding mechanism |
| Scalability | Manual processing or dedicated systems | Highly amenable to automation [9] [5] | Magnetic beads are suited for high-throughput workflows |
| Suitable Sample Types | Moderate to high DNA concentration samples [9] | Challenging samples (e.g., low-yield cfDNA, soil, ancient samples) [9] | Magnetic beads often provide better recovery from low-yield samples [9] |
Workflow and Protocol Comparison
The experimental workflows for the two methods differ significantly in their mechanics, which directly impacts their ease of use, scalability, and potential for automation.
Detailed Spin Column Protocol
The spin column protocol is a multi-step process that leverages centrifugation for separation [31]:
- Step 1: Sample Preparation and Lysis. Cells or tissues are lysed using a buffer containing chaotropic salts and detergents. This cracks open the cells, denatures proteins, and releases nucleic acids into the solution [31].
- Step 2: Binding. The lysate is loaded onto the spin column. During the subsequent centrifugation, nucleic acids selectively bind to the silica-glass fiber membrane in the presence of high salinity, while contaminants flow through the membrane into the collection tube [31].
- Step 3: Washing. One or more wash buffers are applied to the column and centrifuged. These buffers are designed to remove residual impurities like proteins and salts without breaking the interaction between the DNA and the membrane [31]. A common mistake is skipping the final "drying spin," which can leave ethanol residue from the wash buffer that inhibits downstream enzymatic reactions [31].
- Step 4: Elution. The membrane is treated with nuclease-free water or a low-salt buffer (e.g., Tris-EDTA or 0.01 M sodium bicarbonate). This rehydrates the DNA, breaking the hydrogen bonds and releasing the purified nucleic acids. The elution buffer is centrifuged into a fresh, sterile collection tube [30] [31].
- Step 5: Collection. The final eluate containing pure, inhibitor-free DNA is ready for downstream applications such as PCR, qPCR, sequencing, or cloning [31].
Detailed Magnetic Bead Protocol
The magnetic bead protocol, as described for cfDNA extraction, uses magnetic separation [30]:
- Step 1: Sample Lysis. The sample (e.g., blood plasma) is mixed with a lysis buffer containing proteinase K and incubated at 60°C to break down proteins and release cfDNA.
- Step 2: Binding with Crosslinker. A homobifunctional crosslinker like dimethyl suberimidate (DMS) is added to the plasma. DMS immediately binds to DNA through either covalent or electrostatic bonding. Amine-conjugated magnetic beads are then added, which attach to the DMS-DNA complexes. This binding process is conducted at room temperature with gentle mixing [30].
- Step 3: Magnetic Separation. A magnet is placed near the tube, causing the magnetic beads with bound DMS-DNA complexes to aggregate against the wall. The supernatant, containing unwanted materials, is carefully removed with a pipette without the need for centrifugation [30].
- Step 4: Washing. The bead-DNA complexes are washed twice with a buffer like PBS (pH 7.4) to remove impurities. After each wash, the magnetic field is reapplied, and the supernatant is pipetted away [30].
- Step 5: Elution. The DMS-DNA complex-bound beads are treated with an elution buffer (e.g., 0.01 M sodium bicarbonate at pH 10.3). Vortexing and incubation at room temperature break the crosslinking, releasing the pure cfDNA into the solution. A final magnetic separation collects the beads, and the supernatant containing the purified cfDNA is collected [30].
Essential Research Reagent Solutions
The following table details key reagents and materials essential for executing the spin column and magnetic bead protocols, along with their primary functions.
Table: Essential Reagents and Materials for Nucleic Acid Extraction
| Item | Function | Example Use Case |
|---|---|---|
| Chaotropic Salts | Facilitate DNA binding to silica by denaturing proteins and creating high-salt conditions [31]. | Spin column binding buffer; some magnetic bead protocols. |
| Silica Membrane/Column | The solid phase that selectively binds DNA in the presence of chaotropic salts [31]. | Spin column-based purification kits. |
| Silica-coated Magnetic Beads | The mobile solid phase that binds DNA and allows for magnetic separation [9] [30]. | Magnetic bead-based automated or manual extraction. |
| Homobifunctional Crosslinker (e.g., DMS) | Acts as a bridge, binding to DNA and to amine-coated magnetic beads via its two reactive groups [30]. | Specialized magnetic bead protocols for enhanced cfDNA recovery. |
| Wash Buffer | Typically contains ethanol or alcohol; removes salts, proteins, and other contaminants without eluting DNA [31]. | Washing step in both spin column and magnetic bead protocols. |
| Elution Buffer | Low-salt buffer (TE) or water; rehydrates DNA, breaking hydrogen bonds with the silica surface [30] [31]. | Final elution of purified DNA in both methods. |
| Proteinase K | Enzyme that digests and inactivates nucleases and other proteins during lysis [30]. | Initial sample lysis step in both methods. |
The choice between spin column and magnetic bead-based DNA extraction methods is not a matter of one being universally superior, but rather of selecting the right tool for the specific application. The spin column protocol offers a proven, straightforward, and rapid method ideal for laboratories processing a moderate number of samples where DNA concentration is not a limiting factor. Its simplicity and reliability have made it a gold standard in many molecular biology labs [9] [31].
In contrast, magnetic bead-based extraction demonstrates clear advantages in scalability, automation potential, and efficiency, particularly for challenging samples like cfDNA where yield and recovery from low-concentration samples are paramount [9] [30]. The supporting experimental data, showing significantly higher extraction efficiency and faster processing times, makes a compelling case for its adoption in high-throughput settings and liquid biopsy workflows [30] [5].
For researchers and drug development professionals framing their work within the broader thesis of ctDNA extraction method comparison, the decision should be guided by the specific needs of the downstream application. When speed, simplicity, and cost are primary concerns for routine samples, spin columns remain an excellent choice. When maximizing recovery from precious, low-yield samples, processing large volumes, or integrating into fully automated workflows, magnetic beads offer a powerful and often superior alternative.
In the field of circulating tumor DNA (ctDNA) analysis, the purity and yield of extracted nucleic acids are paramount for the success of downstream applications like next-generation sequencing (NGS) and quantitative PCR. The choice of extraction methodology can significantly influence data reliability. This guide objectively compares two dominant techniques—magnetic bead-based protocols and traditional spin columns—focusing on their performance in binding efficiency, separation, and washing steps, to inform researchers and drug development professionals.
Performance Comparison: Magnetic Beads vs. Spin Columns
The following tables summarize key performance metrics and experimental data comparing magnetic bead and spin column technologies.
Table 1: Overall Performance and Workflow Comparison [32] [9] [33]
| Feature | Magnetic Beads | Spin Columns |
|---|---|---|
| Recovery Yield | 94–96% | 70–85% |
| DNA Size Range | 100 bp – 50 kb | 100 bp – 10 kb |
| Throughput | High (96-well & automation compatible) | Low (manual, single-tube focus) |
| Size Selection | Yes (via adjustable bead-to-sample ratio) | No |
| Automation Compatibility | Yes | No |
| Protocol Time | <15 minutes | 20–30 minutes |
| Cost per Sample | ~$0.90 | ~$1.75 |
| Environmental Impact | Lower (less plastic and reagent waste) | Higher |
Table 2: Experimental Performance Data for Key Workflow Steps [32] [34]
| Workflow Step | Magnetic Beads | Spin Columns |
|---|---|---|
| Binding Efficiency | High; binding capacity tends to be higher, especially for low-yield samples [9]. | Limited by the fixed surface area of the silica membrane; can be less effective for low-concentration samples [9]. |
| Separation | Rapid magnetic separation (~2 minutes); no centrifugation needed [32] [34]. | Requires multiple centrifugation steps; difficult to scale for batch processing [32]. |
| Washing Efficiency | Efficient removal of contaminants (salts, enzymes, primers) with 70-80% ethanol washes [34]. | Effective wash requires centrifugation; risk of membrane clogging [33]. |
| Elution | High-concentration elution in 20-50 µL; aided by warm elution buffer [32] [34]. | Larger minimum elution volume, resulting in lower final DNA concentration [33]. |
Table 3: Size Selection Capability of Magnetic Beads [32]
| Bead-to-Sample Ratio | DNA Fragment Size Retained |
|---|---|
| 0.6x | >500 bp |
| 0.8x | >300 bp |
| 1.0x | >100 bp |
| 1.8x | >50 bp |
Experimental Protocols
Detailed Magnetic Bead Protocol for PCR Clean-up
This protocol is adapted from established magnetic bead workflows for purifying DNA, such as post-PCR amplicons, and is the basis for the performance data cited [32] [34] [33].
Principle: Magnetic beads coated with a silica matrix bind DNA in the presence of a binding enhancer like polyethylene glycol (PEG) and salt. The beads are then immobilized using a magnet, and impurities are washed away before the purified DNA is eluted [32] [34].
Required Reagents and Equipment:
- Magnetic Beads: Silica- or carboxyl-coated magnetic beads (e.g., HighPrep PCR Beads, Sera-Mag SpeedBeads) [32] [35].
- Binding Enhancer: A solution containing PEG and salt is often pre-mixed with the beads [32].
- Wash Buffer: Freshly prepared 70-80% ethanol [34].
- Elution Buffer: Nuclease-free water or TE buffer [32] [34].
- Magnetic Stand: Suitable for tubes or 96-well plates [32].
- Pipettes and Tips.
Step-by-Step Methodology:
Binding:
- Transfer your DNA sample (e.g., PCR reaction) to a clean tube.
- Add a precise volume of thoroughly resuspended magnetic beads to the sample. A typical ratio for standard clean-up is 1.8x beads to sample volume [32] [33].
- Mix the bead-sample mixture thoroughly by pipetting or vortexing and incubate at room temperature for 5 minutes to allow DNA binding [32].
Magnetic Separation:
Washing:
- While the tube is still on the magnetic stand, add 200 µL of freshly prepared 80% ethanol to the bead pellet. Incubate for 30 seconds to 1 minute, then carefully pipette and discard the ethanol supernatant [32] [34].
- Repeat this wash step a second time for a total of two washes [34].
- After removing the second wash, ensure all residual ethanol is removed. Air-dry the bead pellet for 3-5 minutes at room temperature. Avoid over-drying, as this can reduce elution efficiency [32].
Elution:
- Remove the tube from the magnetic stand.
- Resuspend the dried beads completely in 20-50 µL of nuclease-free water or TE buffer. Using a pre-warmed (65°C) elution buffer can improve DNA yield [34].
- Incubate the resuspended beads at room temperature for 2 minutes to allow DNA to dissociate.
- Place the tube back on the magnetic stand for 1-2 minutes until the beads pellet and the solution is clear.
- Transfer the eluate, which now contains the purified DNA, to a new tube.
Principle: DNA binds to a silica membrane in the spin column under high-salt conditions. Contaminants are removed through a series of wash and centrifugation steps, and pure DNA is eluted in a low-salt buffer [9] [33].
- Binding: The sample is mixed with a binding buffer and loaded into the spin column. A centrifugation step (e.g., 30-60 seconds) binds DNA to the membrane while flow-through is discarded.
- Washing: The column is washed once or twice with a wash buffer (often ethanol-based) via centrifugation to remove impurities.
- Elution: A low-salt elution buffer or water is added to the column membrane. After a brief incubation, a final centrifugation step (1-2 minutes) elutes the purified DNA.
Workflow Visualization
The following diagram illustrates the core steps of the magnetic bead protocol.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 4: Key Materials for Magnetic Bead-Based Nucleic Acid Purification [32] [35] [34]
| Item | Function | Key Considerations |
|---|---|---|
| Magnetic Beads | Solid-phase support that reversibly binds nucleic acids; the core of the technology. | Choose coating (silica/carboxyl) and size based on application. Ensure consistent resuspension before use [35] [36]. |
| Magnetic Stand | Device that generates a magnetic field to immobilize beads during separation and wash steps. | Select a stand compatible with tube/plate format. A slanted design aids supernatant removal [32] [36]. |
| Binding Buffer | Creates conditions (e.g., with PEG and salt) that promote nucleic acid adsorption to the beads. | Critical for efficiency; often optimized and supplied with commercial bead kits [32] [34]. |
| Wash Buffer (Ethanol) | Removes contaminants like salts, proteins, and enzymes from the bead-bound DNA. | Must be freshly prepared at 70-80% concentration to prevent salt precipitation and ensure effective cleaning [34]. |
| Elution Buffer | Low-ionic-strength solution that disrupts DNA-bead interaction, releasing purified DNA. | Nuclease-free water or TE buffer. Pre-warming to 65°C can significantly increase yield [32] [34]. |
For ctDNA extraction and other sensitive molecular applications requiring high recovery and compatibility with advanced downstream analysis, magnetic bead technology offers a superior alternative to traditional spin columns. The data demonstrates clear advantages in yield, throughput, and cost-effectiveness. The flexibility of magnetic bead protocols, particularly the ability to perform fine size selection and seamless automation, makes them an indispensable tool for modern genomics and drug development pipelines.
The choice between magnetic bead and spin column technologies for circulating tumor DNA (ctDNA) extraction is a critical decision that directly impacts the success of downstream genomic analyses in cancer research and clinical diagnostics. As liquid biopsy applications expand from early cancer detection to minimal residual disease monitoring and therapy selection, matching the appropriate extraction method to specific sample types and processing volumes becomes increasingly important. This guide provides an objective comparison of these two dominant ctDNA extraction technologies, supported by experimental data, to help researchers and drug development professionals select the optimal approach for their specific experimental requirements.
Spin Column Technology
Spin column-based DNA extraction kits utilize a silica membrane housed in a column format that selectively binds DNA under high-salt conditions [9]. The process involves sample lysis to release nucleic acids, binding of DNA to the silica membrane in the presence of a binding buffer, multiple wash steps to remove contaminants and impurities, and final elution of purified DNA using a low-salt buffer [9]. This technology has been optimized over years for DNA yield and purity, making it suitable for most common applications including PCR, cloning, and sequencing [9].
Magnetic Bead Technology
Magnetic bead-based extraction employs paramagnetic particles coated with a DNA-binding surface [9]. Following sample lysis, DNA binds to the beads, and a magnetic field is applied to separate the bead-DNA complexes from the rest of the sample [9]. Subsequent wash steps remove impurities, followed by elution of purified DNA [9]. This method is particularly noted for its scalability and automation compatibility, making it advantageous for high-throughput settings [9].
Comparative Performance Analysis
Yield and Recovery Efficiency
Multiple studies have directly compared the performance of spin column and magnetic bead-based extraction methods. A comprehensive evaluation of seven commercial cfDNA extraction kits (three spin column-based and four magnetic beads-based) found significant variability in both cfDNA yield and fragment size distribution across different kits [13].
Table 1: Performance Comparison of Representative Extraction Kits
| Extraction Method | Kit Identifier | Median LMW DNA Yield (GEs/mL plasma) | LMW Fraction (%) | Methodology |
|---|---|---|---|---|
| Spin Column | Kit A | 1,936 | 89% | ddPCR [13] |
| Spin Column | Kit B | 1,760 | Not specified | ddPCR [13] |
| Magnetic Beads | Kit E | 1,515 | 90% | ddPCR [13] |
The highest median yield of low molecular weight (LMW) cfDNA was obtained using a spin column-based method (Kit A), with 1,936 genome equivalents (GEs) per mL of plasma and an LMW fraction of 89% [13]. Among magnetic beads-based methods, Kit E showed the highest yield of LMW DNA (median 1,515 LMW copies/mL plasma) with a comparable LMW fraction of 90% [13]. The yield difference between the top-performing spin column kit and the leading magnetic bead kit was statistically significant (t-test p = 9.46 × 10⁻⁵) [13].
Sample Input and Throughput Considerations
Sample input requirements and processing capabilities vary substantially between the two technologies:
Table 2: Workflow and Throughput Characteristics
| Parameter | Spin Column | Magnetic Bead |
|---|---|---|
| Maximum Throughput | Limited by manual processing | High, amenable to automation [9] |
| Sample Processing | Individual column processing [26] | 8-strip tubes and multi-channel pipettors [26] |
| Hands-on Time | Significant for large batches [26] | Reduced through parallel processing [26] |
| Size Selection Capability | Limited | Excellent for sequencing libraries [26] |
Magnetic bead systems demonstrate superior performance for high-throughput applications. As noted in CUT&RUN workflows, processing 48+ spin columns is "a daunting task," whereas SPRI bead-based purification allows researchers to work with 8-strip tubes and multi-channel pipettors, significantly improving efficiency [26]. The streamlined process reduces opportunities for human error, DNA shearing, and sample loss, thereby enhancing reproducibility [26].
Detailed Experimental Protocols
Spin Column Protocol for ctDNA Extraction
The standard protocol for spin column-based ctDNA extraction typically follows these steps:
Plasma Preparation: Collect blood in EDTA tubes or specialized cell-free DNA blood collection tubes (BCTs). Process EDTA tubes within 2-6 hours; BCTs can be stored for up to 7 days at room temperature [37]. Perform double centrifugation: initial slow spin (380-3,000 × g for 10 min at room temperature) to separate plasma from blood cells, followed by high-speed centrifugation (12,000-20,000 × g for 10 min at 4°C) to remove remaining cellular debris [37].
Sample Lysis: Mix plasma with a lysis buffer containing chaotropic salts (e.g., guanidine hydrochloride) and detergents to release nucleic acids and denature proteins.
DNA Binding: Add binding buffer to create high-salt conditions and transfer the mixture to the spin column. Centrifuge to facilitate DNA binding to the silica membrane [9].
Washing: Perform two wash steps using ethanol-based wash buffers to remove contaminants, proteins, and salts. Centrifuge after each wash to remove flow-through [9].
Elution: Add elution buffer (TE buffer or nuclease-free water) and centrifuge to collect purified ctDNA. Pre-heating the elution buffer to 60-70°C may improve yield [9].
Magnetic Beads Protocol for ctDNA Extraction
The standard protocol for magnetic bead-based ctDNA extraction includes:
Plasma Preparation: Identical to spin column protocol, using EDTA tubes or BCTs with double centrifugation [37].
Sample Lysis: Mix plasma with lysis buffer. Some protocols incorporate proteinase K digestion for improved yield.
DNA Binding: Add magnetic beads suspended in a binding buffer containing polyethylene glycol (PEG) and high salt concentrations. Incubate with mixing to allow DNA binding to the beads [26].
Magnetic Separation: Place the tube on a magnetic rack to capture beads. Discard the supernatant once the solution clears.
Washing: Wash beads with 70-85% ethanol while positioned on the magnetic rack. Remove wash solution completely.
Elution: Resuspend beads in elution buffer (TE or low-salt buffer) and incubate to release DNA. Apply magnetic separation and transfer the eluate containing purified ctDNA to a new tube [26].
Figure 1: Comparative Workflow Diagram: Spin Column vs. Magnetic Bead ctDNA Extraction
Application-Specific Recommendations
Matching Method to Sample Type
- Low DNA Concentration Samples: Magnetic bead-based kits often provide better recovery rates for samples with low DNA yields, as the binding capacity of magnetic beads tends to be higher compared to silica membranes [9].
- Challenging Sample Types: Magnetic bead methods are more adaptable for various sample types, including complex biological materials like soil or ancient specimens [9].
- High-Throughput Settings: Magnetic bead technology is preferable for automated, high-volume processing due to compatibility with liquid handling systems [9] [26].
- Standard Molecular Biology Applications: Spin column kits remain effective for labs processing moderate sample numbers with moderate to high DNA concentrations, particularly when cost considerations are paramount [9].
Volume Considerations
- Small Volume Processing: Both methods accommodate small sample volumes, though spin columns may have minimum volume requirements for efficient binding.
- Large Volume Processing: Magnetic bead systems excel at processing large sample volumes through scalable binding capacity, while spin columns are limited by membrane capacity [9].
- Blood Collection Considerations: For ctDNA analysis, collect 2 × 10 mL of blood for single-analyte liquid biopsy, though screening, minimal residual disease detection, whole genome sequencing, and multiple analyte testing may require larger plasma volumes [37].
Research Reagent Solutions
Table 3: Essential Materials for ctDNA Extraction and Analysis
| Reagent/Category | Example Products | Function/Application |
|---|---|---|
| Blood Collection Tubes | cfDNA (Streck), PAXgene Blood ccfDNA (Qiagen), cfDNA/cfRNA Preservative (Norgene), ImproGene cfDNA (Improve Medical), cfDNA (Roche) | Preserve blood cell integrity, prevent gDNA release, enable room temperature storage/transport [37] |
| Spin Column Kits | QIAamp Circulating Nucleic Acid Kit (Qiagen), Cobas ccfDNA Sample Preparation Kit | Silica membrane-based ctDNA isolation; demonstrate high LMW DNA yield [13] [37] |
| Magnetic Bead Kits | QIAamp MinElute ccfDNA Mini Kit (Qiagen), Maxwell RSC LV ccfDNA Kit (Promega), MagNa Pure 24 Total NA Isolation Kit (Roche) | Paramagnetic particle-based extraction; enable automation, high-throughput processing [37] |
| Quality Assessment Tools | Droplet Digital PCR (ddPCR), Agilent BioAnalyzer, Qubit Fluorometer | Quantify amplifiable DNA, assess fragment size distribution, verify sample quality [13] |
The selection between magnetic bead and spin column technologies for ctDNA extraction should be guided by specific research requirements, sample characteristics, and operational constraints. Spin column methods demonstrate advantages in DNA yield and established reliability for standard applications, while magnetic bead systems offer superior automation compatibility, throughput potential, and size selection capabilities. Researchers should consider implementing the quality assessment methods described herein to validate their extraction choice for specific applications, particularly when working with limited sample material or requiring high sensitivity for low-abundance variant detection. As liquid biopsy applications continue to evolve toward earlier cancer detection and minimal residual disease monitoring, the precise matching of extraction methodology to experimental needs remains fundamental to generating reliable, reproducible genomic data.
The analysis of circulating tumor DNA (ctDNA) has emerged as a cornerstone of liquid biopsy applications in precision oncology, enabling non-invasive tumor genotyping, treatment response monitoring, and minimal residual disease detection. A significant bottleneck in translating this powerful biomarker into widespread clinical practice lies in the pre-analytical phase, specifically the extraction of cell-free DNA (cfDNA) from plasma. Traditional manual methods, while reliable, are labor-intensive and subject to operator variability, creating limitations for large-scale studies and clinical implementation where processing dozens to hundreds of samples efficiently is paramount. This guide objectively compares the performance of manual spin-column-based extraction with automated magnetic bead-based platforms, providing researchers with experimental data to inform their scaling strategies from manual processing to high-throughput automation.
Methodological Principles: Spin Columns vs. Magnetic Beads
Fundamental Binding Chemistry
Despite their different physical implementations, both spin columns and magnetic beads often rely on a common underlying chemistry: the selective binding of DNA to a silica surface under high-salt conditions.
- Spin Column Technology: The manual QIAamp Circulating Nucleic Acid Kit (Qiagen) is a classic example of this method. The process involves lysing the sample to release DNA, adding a binding buffer to facilitate DNA adherence to a silica membrane housed within a spin column, performing wash steps to remove contaminants and impurities, and finally, using an elution buffer to release the purified DNA from the membrane [9].
- Magnetic Bead Technology: Methods like the QIAsymphony SP Circulating DNA Kit (Qiagen) and the Maxwell RSC LV ccfDNA Plasma Custom Kit (Promega) use magnetic particles coated with a silica-based DNA-binding surface. DNA from the lysed sample binds to these beads, and a magnetic field is applied to separate the bead-bound DNA from the rest of the sample. Subsequent wash steps remove impurities, and elution releases the purified DNA [9] [38].
Key Differentiator: The Path to Automation
The core distinction lies in the ease of automation. The process of magnetic separation is inherently more amenable to automation in a plate-based format than the centrifugation and liquid handling required for spin columns. This fundamental difference drives the significant divergence in throughput, hands-on time, and reproducibility between the two approaches.
To objectively evaluate these technologies, a pivotal study directly compared the manual QIAamp (QA) platform against two automated systems: the QIAsymphony (QS) and the Maxwell (MX) [38]. The experimental protocol and key findings are summarized below.
Experimental Protocol
- Sample Preparation: Blood samples from healthy donors and metastatic cancer patients were collected. Plasma was isolated from blood drawn in CellSave or EDTA tubes within specified timeframes (96h for CellSave, 24h for EDTA) through two centrifugation steps [38].
- cfDNA Isolation: For each platform, cfDNA was isolated from 2 mL of plasma according to the manufacturers' protocols, with minor modifications. The QS isolation included the addition of carrier RNA, while the MX protocol required a third plasma centrifugation step to eliminate residual leukocytes. All samples were eluted in a 60 μL volume [38].
- Performance Assessment: The isolated cfDNA was evaluated using multiple metrics:
- Total cfDNA Quantity: Assessed by TERT quantitative PCR (qPCR).
- Recovery Efficiency: Measured by qPCR analysis of spiked-in synthetic plant DNA.
- Genomic DNA Contamination: Determined using a β-actin fragmentation assay.
- ctDNA Quality: Evaluated by digital PCR (dPCR) for the detection of known somatic variants [38].
The following table synthesizes the quantitative and operational data from the aforementioned study and other relevant literature [38] [12].
Table 1: Direct comparison of manual spin-column and automated magnetic bead-based cfDNA extraction platforms.
| Performance Metric | QIAamp (QA) Manual Spin Column | QIAsymphony (QS) Automated Beads | Maxwell (MX) Automated Beads |
|---|---|---|---|
| Plasma Input Volume | 1.0–5.0 mL | 2.0–8.0 mL | 2.0–4.0 mL |
| Samples per Run | 24 | 96 | 16 (48 on RSC 48 instrument) |
| Handling Time per Run | 180–240 minutes | ~30 minutes | ~30 minutes |
| Estimated Cost per Sample | ~€20 | ~€24 | ~€20 |
| cfDNA Yield | High (Benchmark) | Comparable to QA | Lower than QA and QS |
| Recovery Efficiency | High (Benchmark) | Comparable to QA | Lower |
| Variant Allele Frequency (VAF) Accuracy | High (Benchmark) | Comparable to QA (no significant difference) | Comparable to QA (no significant difference) |
| Inter-Operator Reproducibility | Lower (Manual process) | Higher (Automated process) | Higher (Automated process) |
Key Findings and Interpretation
The data reveals a clear trade-off between performance and throughput. The manual QA and automated QS platforms showed comparable performance in critical metrics like cfDNA yield, recovery efficiency, and accuracy of variant detection [38]. This demonstrates that automation does not necessarily compromise quality. The MX platform, while automated, showed lower yields and recovery in this particular comparison [38]. Another independent study confirmed that spin-column-based methods like the QIAamp kits can produce high yields and reproducibility, though they noted these methods are often more costly and time-consuming than magnetic bead approaches [12].
Operationally, the advantage of automation is stark. The hands-on time for the automated systems is reduced to approximately 30 minutes, regardless of the number of samples being processed, compared to 3-4 hours for a batch of 24 samples with the manual method [38]. This translates to a dramatic increase in laboratory efficiency and a reduction in the risk of human error.
The Researcher's Toolkit: Essential Reagents and Platforms
Table 2: Key research reagent solutions for cfDNA extraction.
| Product Name | Manufacturer | Type | Key Function |
|---|---|---|---|
| QIAamp Circulating Nucleic Acid Kit | Qiagen | Manual Spin Column | High-yield manual cfDNA isolation; considered a "gold standard" [38] [12]. |
| QIAsymphony SP Circulating DNA Kit | Qiagen | Automated Magnetic Beads | High-throughput automated extraction on the QIAsymphony SP platform [38]. |
| Maxwell RSC LV ccfDNA Plasma Kit | Promega | Automated Magnetic Beads | Automated extraction on the benchtop Maxwell RSC instruments [38]. |
| MagNA Pure 24 Total NA Isolation Kit | Roche | Automated Magnetic Beads | Automated extraction on the MagNA Pure 24 system; shown to provide high yield and reproducibility [12]. |
| Cell-Free DNA BCTs (Streck) / PAXgene Blood ccfDNA Tubes (Qiagen) | Streck / Qiagen | Blood Collection Tubes | Preservative tubes that prevent leukocyte lysis and stabilize cfDNA, allowing for extended sample transport and storage [5]. |
Workflow Visualization: From Manual to Automated Processing
The following diagram illustrates the key steps and decision points in scaling ctDNA extraction from manual to high-throughput automated workflows.
The choice between manual spin-column and automated magnetic bead-based extraction is not a question of which technology is superior in a vacuum, but which is optimal for a specific research context. The manual QIAamp (spin-column) method remains a robust and reliable "gold standard" for laboratories with low sample volumes or limited access to automated instrumentation, as it delivers high yields and purity [38] [12].
For studies involving large patient cohorts, clinical trials, or routine clinical implementation where reproducibility, speed, and scalability are critical, automated magnetic bead-based platforms like the QIAsymphony are the clear choice. The experimental data confirms that these systems can match the quality of manual methods while drastically reducing hands-on time and increasing throughput [38]. When scaling up, researchers must also consider complementary pre-analytical factors, such as the use of preservative blood collection tubes (e.g., Streck tubes) that maintain sample integrity for longer periods, thereby providing the flexibility needed for processing large sample batches [5]. By aligning the extraction methodology with the project's scale and objectives, researchers can effectively harness the full potential of ctDNA in advancing precision oncology.
Optimizing Performance: Troubleshooting and Protocol Refinement
For researchers and drug development professionals working with circulating tumor DNA (ctDNA), obtaining sufficient DNA yield is not merely a technical concern but a fundamental determinant of assay success. The analysis of ctDNA presents unique challenges due to its exceptionally low concentration in blood, typically ranging from 1–10 ng/mL in plasma, and its rapid clearance from circulation with a half-life of just 16 minutes to 2.5 hours [1]. In precision oncology, where liquid biopsies enable non-invasive cancer monitoring and therapy response assessment, low DNA yield can compromise the detection of critical biomarkers, potentially leading to false negatives in mutation detection or inaccurate quantification of variant allele frequencies [1].
The choice between magnetic bead-based and spin column-based extraction methods significantly impacts yield, purity, and downstream application success. This guide provides an objective comparison of these technologies, supported by experimental data and detailed protocols, to help researchers optimize DNA recovery specifically for ctDNA and other challenging sample types. Understanding the causes of low yield and implementing appropriate solutions is essential for generating reliable, reproducible data in oncology research and diagnostic development.
Technical Comparison: Magnetic Bead vs. Spin Column Technologies
Fundamental Principles and Workflows
Spin column-based extraction relies on the selective binding of DNA to a silica membrane in a spin column under high-salt conditions [9]. The process involves sample lysis, binding to the silica membrane, washing to remove contaminants, and elution of purified DNA [9]. The centrifugation steps required at each stage make this method inherently difficult to automate, limiting its throughput potential [39].
Magnetic bead-based extraction utilizes paramagnetic particles coated with a DNA-binding surface [9]. After sample lysis, DNA binds to the beads in the presence of a binding buffer, often containing polyethylene glycol (PEG) and high salt concentrations—a principle known as Solid Phase Reversible Immobilization (SPRI) [26]. A magnetic field then immobilizes the beads while contaminants are removed through washing steps, followed by elution of pure DNA in a low-salt buffer [9] [26]. This magnetic separation principle enables full automation on robotic platforms, making it ideal for high-throughput laboratories [40] [41].
Table 1: Core Technology Comparison Between Methods
| Parameter | Spin Column | Magnetic Beads |
|---|---|---|
| Binding Principle | Silica membrane with high-salt conditions [9] | SPRI technology with magnetic particles [26] |
| Separation Method | Centrifugation [9] | Magnetic field [9] |
| Automation Potential | Low to moderate [39] | High [40] |
| Typical Elution Volume | 30–100 µL [42] | 20–50 µL [39] |
Performance Metrics and Experimental Data
Direct comparative studies reveal significant differences in performance characteristics between the two methods. Magnetic bead-based systems consistently demonstrate advantages in recovery yield, particularly for low-concentration samples and smaller DNA fragments commonly encountered in ctDNA analysis [39].
Table 2: Quantitative Performance Comparison Based on Experimental Data
| Performance Metric | Spin Column | Magnetic Beads | Experimental Context |
|---|---|---|---|
| Recovery Yield | 70–85% [39] | 94–96% [39] | PCR product cleanup [39] |
| DNA Size Range | 100 bp – 10 kb [39] | 100 bp – 50 kb [39] | Fragment retention capacity [39] |
| Cost per Sample | ~$1.75 [39] | ~$0.90 [39] | PCR cleanup for 1000 samples [39] |
| Throughput (Samples/Protocol Hour) | Low (manual processing) [40] | High (96-well & automation) [40] [39] | Processing efficiency comparison [40] |
| Size Selection Capability | No [39] | Yes (via bead ratio adjustment) [39] | SPRI bead technology [39] [26] |
Research applications requiring precise size selection, such as CUT&RUN sequencing libraries, particularly benefit from magnetic bead technology. The ability to adjust the bead-to-sample ratio allows researchers to selectively retain DNA fragments within specific size ranges, enabling efficient removal of adapter dimers and other unwanted fragments without additional purification steps [26].
Causes and Troubleshooting of Low DNA Yield
Common Causes Across Both Methods
Regardless of the extraction methodology, several pre-analytical and analytical factors can contribute to suboptimal DNA yield:
Pre-analytical Variables:
- Sample Quality: Clotted or hemolyzed blood samples can trap white blood cells or release nucleases that degrade DNA [43]. For ctDNA analysis, the use of EDTA blood collection tubes is essential, as heparin can inhibit downstream PCR reactions [43].
- Improper Storage: Delay in processing or multiple freeze-thaw cycles rapidly degrades DNA quality. Blood samples should be processed or refrigerated (4°C) shortly after collection [43].
- Biological Factors: Samples from pediatric, geriatric, or immunocompromised patients may inherently contain fewer white blood cells, resulting in lower DNA content [43].
Binding Efficiency Issues:
- Insufficient Lysis: Incomplete cell lysis prevents DNA release from cells. Optimization of lysis conditions (pH, temperature, duration) and use of detergents like Triton X-100 or SDS can enhance cell disruption [42].
- Inefficient Binding: Inadequate mixing of binding buffer with sample or suboptimal salt concentrations can reduce DNA binding to either silica membranes or magnetic beads [42].
Method-Specific Causes and Solutions
Table 3: Troubleshooting Low DNA Yield for Each Method
| Problem | Causes | Solutions | Applicable Method |
|---|---|---|---|
| Insufficient Sample Lysis | Inadequate lysis buffer, suboptimal conditions, short incubation [42] | Optimize lysis conditions (30 min at 56°C); use mechanical disruption [43] [42] | Both |
| Inefficient Binding | Inadequate buffer mixing; contaminant interference [42] | Ensure proper mixing; optimize pH and salt concentration; use chaotropic salts [42] | Both |
| Incomplete Washing | Residual impurities in final eluate [42] | Perform recommended wash steps thoroughly; ensure complete flow-through [42] | Both |
| Low Elution Volume | Suboptimal elution buffer volume or temperature [42] | Use higher elution volume; pre-warm elution buffer to 60–70°C [42] | Both |
| Membrane Blockage | Silica membrane clogging with particulate matter | Centrifuge samples before loading; avoid overloading column | Spin Column |
| Bead Handling Issues | Incomplete resuspension; over-drying of beads [42] | Vortex beads until fully dispersed; avoid over-drying during wash steps [42] | Magnetic Beads |
| Inadequate Magnetic Separation | Insufficient separation time; bead clumping | Increase separation time; ensure proper bead mixing | Magnetic Beads |
Workflow Optimization Strategies
For Spin Column Protocols:
- Increase Binding Efficiency: Pre-wash columns with wash buffer to remove potential contaminants that may interfere with binding [42].
- Maximize Elution Efficiency: Let the column incubate with elution buffer for several minutes before centrifugation to increase DNA recovery [42]. Using elution buffer heated to 60–70°C can significantly improve yield by reducing buffer viscosity and promoting DNA release from the silica membrane [42].
- Avoid Overloading: Exceeding the recommended sample input can lead to incomplete lysis and inefficient binding [42].
For Magnetic Bead Protocols:
- Optimize Bead Resuspension: Ensure complete resuspension of magnetic beads by gentle vortexing until fully dispersed and homogeneous to maximize binding surface area [42].
- Control Bead Drying: Avoid over-drying beads during wash steps, as this can cause clumping and reduce elution efficiency. Beads should be left slightly damp after ethanol removal [42].
- Optimize Incubation Times: Allow sufficient time (typically 5 minutes at room temperature) for complete DNA binding to beads before magnetic separation [39].
Research Reagent Solutions and Experimental Protocols
Essential Research Reagents
Table 4: Key Reagent Solutions for Nucleic Acid Extraction
| Reagent/Category | Function | Specific Examples |
|---|---|---|
| Lysis Buffers | Cell disruption and DNA release | Triton X-100, SDS-containing buffers [42] |
| Binding Enhancers | Promote DNA binding to matrices | Chaotropic salts (guanidine, ammonium sulfate) [42] |
| Wash Buffers | Remove contaminants while retaining DNA | Ethanol-based wash solutions [39] [33] |
| Elution Buffers | Release purified DNA from matrix | Low-salt buffers, TE buffer, nuclease-free water [42] [39] |
| Enzymatic Reagents | Digest proteins and remove RNA | Proteinase K, RNase A [43] |
| Magnetic Beads | DNA binding and magnetic separation | SPRI beads, HighPrep PCR beads [39] [26] |
Standardized Experimental Protocols
Magnetic Bead-Based Extraction Protocol (Adapted from HighPrep Protocol [39]):
- Binding: Add 1.8x volume of magnetic beads to the sample. Mix thoroughly by pipetting or vortexing.
- Incubation: Let stand for 5 minutes at room temperature to allow DNA binding.
- Separation: Place the sample on a magnetic stand until the solution clears and beads form a pellet (~2 minutes). Discard the supernatant.
- Washing: While tubes remain on the magnetic stand, add 200μL of 80% ethanol. Incubate for 30 seconds, then remove and discard ethanol. Repeat twice for a total of three washes.
- Drying: Air-dry beads for 3–5 minutes at room temperature. Avoid over-drying.
- Elution: Resuspend beads in 20–50μL nuclease-free water or TE buffer. Mix thoroughly and incubate for 2 minutes.
- Final Separation: Place on magnetic stand until solution clears. Transfer the eluate containing purified DNA to a new tube.
Spin Column-Based Extraction Protocol (Optimized for Maximum Yield):
- Lysis: Add appropriate lysis buffer to the sample. Incubate at 56°C for 30 minutes with occasional mixing.
- Binding: Add binding buffer to the lysate and mix thoroughly. Transfer the mixture to a spin column and centrifuge at ≥10,000 × g for 1 minute.
- Washing: Add wash buffer to the column and centrifuge at ≥10,000 × g for 1 minute. Discard the flow-through. Repeat with a second wash step.
- Additional Centrifugation: Centrifuge the empty column for an additional 1 minute to remove residual ethanol.
- Elution: Add pre-warmed (60–70°C) elution buffer to the center of the column membrane. Let stand for 2–5 minutes, then centrifuge at maximum speed for 1 minute.
Method Selection Guide for Research Applications
Application-Specific Recommendations
The optimal choice between magnetic bead and spin column technologies depends heavily on the specific research requirements and experimental constraints:
Choose Magnetic Beads When:
- Processing large sample volumes (96- or 384-well formats) requiring high throughput [40] [39]
- Automation is essential for workflow efficiency and reproducibility [40] [41]
- Working with low-concentration samples or challenging sample types where maximum recovery is critical [9] [43]
- Size selection capabilities are needed for applications like NGS library preparation [39] [26]
- Cost efficiency at scale is important, with magnetic beads offering approximately 50% cost reduction per sample compared to spin columns for high-throughput processing [39]
Choose Spin Columns When:
- Processing low to moderate numbers of samples where automation is not justified [40]
- Equipment limitations prevent investment in magnetic separation systems [9]
- Rapid, single-sample processing is required without batch processing needs [40]
- Budget constraints for initial setup outweigh long-term per-sample costs [40]
Decision Framework for ctDNA Research
For circulating tumor DNA applications specifically, magnetic bead-based methods generally offer significant advantages due to their superior recovery of low-abundance DNA fragments and compatibility with automated processing of multiple samples [1]. The ability to precisely control size selection through bead-to-sample ratios is particularly valuable for ctDNA analysis, where fragment size distribution can provide important diagnostic information [26].
When establishing a new ctDNA workflow, researchers should consider:
- Sample volume availability - Magnetic beads typically require smaller elution volumes, resulting in higher concentration samples suitable for low-input downstream applications [39].
- Downstream application requirements - Techniques like digital PCR and next-generation sequencing, commonly used in ctDNA analysis, benefit from the higher purity and yield provided by magnetic bead purification [1] [39].
- Quality control needs - Magnetic bead protocols typically generate more consistent results with lower coefficients of variation, essential for detecting subtle changes in ctDNA levels during therapy monitoring [44].
Workflow Diagram: DNA Extraction Methods
The following diagram illustrates the parallel workflows for magnetic bead and spin column-based DNA extraction methods, highlighting key steps where yield losses commonly occur and optimization strategies can be implemented:
Selecting the appropriate DNA extraction method and optimizing its implementation are critical decisions that significantly impact research outcomes, particularly in sensitive applications like ctDNA analysis. While spin columns offer simplicity and convenience for low-throughput applications, magnetic bead-based technologies provide superior recovery, automation compatibility, and cost efficiency at scale. By understanding the specific causes of low DNA yield and implementing the appropriate troubleshooting strategies outlined in this guide, researchers can significantly improve the quality and reliability of their nucleic acid extraction workflows, ultimately enhancing the validity of their downstream analytical results.
Optimizing Bead-to-Sample Ratios for Size Selection and Recovery
In the field of liquid biopsy and molecular biology, the purification and size selection of nucleic acids are critical steps that significantly impact the success of downstream applications. Among the various extraction methodologies, magnetic bead-based technology has emerged as a powerful alternative to traditional spin columns, offering superior control over fragment selection and enhanced recovery efficiencies. The performance of magnetic bead systems is fundamentally governed by the precise optimization of bead-to-sample ratios, a parameter that directly influences the size range of recovered DNA fragments and the overall yield. This principle of Solid Phase Reversible Immobilization (SPRI) relies on the controlled interaction between paramagnetic beads and DNA molecules in the presence of crowding agents like polyethylene glycol (PEG) and high-salt buffers [26].
The strategic manipulation of bead-to-sample ratios allows researchers to selectively target specific DNA fragment sizes, making this technology particularly valuable for applications working with fragmented DNA such as circulating tumor DNA (ctDNA) and cell-free DNA (cfDNA). Unlike spin columns which often exhibit inherent size bias and limited customization capabilities, magnetic bead systems provide researchers with a tunable parameter that can be optimized for specific experimental needs, from recovering ultra-short cfDNA fragments to selecting ideal insert sizes for next-generation sequencing (NGS) library preparation [45] [26]. This technical guide provides a comprehensive comparison of these competing technologies, with particular emphasis on the optimization strategies that maximize recovery and size selection performance for critical applications in clinical and research settings.
Technical Comparison: Magnetic Beads vs. Spin Columns
The fundamental differences between magnetic bead and spin column technologies extend beyond their operational mechanics to impact virtually every aspect of nucleic acid extraction, from yield and fragment size recovery to automation compatibility and cost efficiency.
Table 1: Comprehensive Performance Comparison of DNA Extraction Methods
| Feature | Magnetic Beads | Spin Columns |
|---|---|---|
| Recovery Yield | 94–96% [45] | 70–85% [45] |
| DNA Size Range | 100 bp – 50 kb [45] | 100 bp – 10 kb [45] |
| Size Selection Capability | Yes (via adjustable bead ratio) [45] [26] | No [45] |
| Throughput | High (96-well & automation compatible) [45] [26] | Low (manual processing) [45] |
| Automation Compatibility | Full compatibility [45] | Not compatible [45] |
| Cost per Sample | ~$0.90 [45] | ~$1.75 [45] |
| Protocol Time | <15 minutes [45] | 20–30 minutes [45] |
| Hands-on Time | Minimal [26] | Significant [26] |
The operational superiority of magnetic beads is particularly evident in their size selection capability, which allows researchers to precisely target specific DNA fragment ranges by simply adjusting the volumetric ratio of beads to sample. This flexibility enables applications such as selective recovery of mononucleosomal cfDNA fragments (~167 bp) while excluding shorter adapter dimers or longer genomic DNA contaminants [26]. In contrast, spin columns employ a fixed silica membrane with inherent pore sizes that create an uncontrolled size bias, often resulting in the preferential loss of shorter fragments that fail to bind effectively or longer fragments that may be sheared during binding/washing steps [45].
The throughput advantages of magnetic beads extend beyond simple multi-well processing to encompass full automation compatibility with systems such as the Thermo Fisher KingFisher Flex, Hamilton Micolab STAR, and Beckman Coulter Biomek i-Series [45]. This automation capability not only reduces hands-on time but also significantly enhances reproducibility by minimizing human error—a critical consideration for clinical applications and large-scale research studies [26]. Additionally, magnetic bead protocols generate less plastic waste due to compatibility with 96-well plate formats and bulk reagent packaging, offering both environmental and economic benefits compared to the single-use nature of spin columns [45].
Experimental Protocols for Performance Validation
Magnetic Bead-Based Cleanup and Size Selection Protocol
The following detailed methodology outlines the standard operating procedure for magnetic bead-based DNA cleanup, with specific emphasis on ratio adjustments for size selection:
- Binding: Add a precisely calculated volume of magnetic beads to the DNA sample. The bead-to-sample ratio typically ranges from 0.6x to 1.8x depending on the desired size selection (see Table 2 for specific ratios). Mix thoroughly by pipetting or vortexing to ensure complete homogenization [45].
- Incubation: Allow the mixture to stand at room temperature for 5 minutes to facilitate DNA binding to the beads through the SPRI mechanism. During this incubation, the PEG and salt conditions promote reversible binding of DNA to the magnetic particles [45] [26].
- Separation: Transfer the sample to a magnetic stand or plate and incubate until the solution clears and beads form a visible pellet (approximately 2 minutes). The supernatant contains unbound molecules that fall below the selected size threshold [45].
- Washing: While the tube remains on the magnetic stand, carefully remove and discard the supernatant. Wash the beads twice with 500 μL of freshly prepared 80% ethanol, incubating for 30 seconds during each wash to remove salts, enzymes, and other contaminants [45].
- Drying: Air dry the beads for 3-5 minutes at room temperature to ensure complete ethanol evaporation. Avoid over-drying, as this can reduce elution efficiency by making DNA resuspension more difficult [45].
- Elution: Resuspend the dried beads in 20-50 μL of nuclease-free water or TE buffer by vigorous pipetting or vortexing. Incubate for 2 minutes at room temperature, then place on the magnetic stand until the solution clears. Transfer the eluate containing purified, size-selected DNA to a clean tube [45].
Comparative Performance Assessment Protocol
To objectively evaluate the performance of magnetic bead versus spin column methods, the following experimental approach can be implemented:
- Sample Preparation: Spike standardized oligonucleotide mixtures (with fragments spanning 40-100 bp) into commercial human plasma to create a reference material. This approach controls for initial concentration and fragment distribution [19] [46].
- Parallel Processing: Divide each spiked plasma sample into equal aliquots for simultaneous processing with magnetic bead (at multiple bead ratios) and spin column methods. Include replicates to assess technical variability [19].
- Yield Quantification: Measure DNA concentration using fluorometric methods (e.g., Qubit) and verify fragment size distribution through microelectrophoresis (e.g., Bioanalyzer, TapeStation) [19] [5].
- Functional Assessment: Perform downstream analyses such as qPCR, digital PCR, or NGS library preparation to evaluate the functional quality of the recovered DNA and its suitability for intended applications [19] [46].
Bead-to-Sample Ratio Optimization for Size Selection
The precise control over DNA fragment size recovery represents one of the most significant advantages of magnetic bead-based extraction systems. By modulating the bead-to-sample ratio, researchers can selectively target specific size ranges optimal for their particular applications.
Table 2: DNA Size Selection Based on Bead-to-Sample Ratios
| Bead-to-Sample Ratio | DNA Fragment Size Retained | Typical Applications |
|---|---|---|
| 0.6x | >500 bp [45] | Genomic DNA, long fragments |
| 0.8x | >300 bp [45] | Large cfDNA fragments |
| 1.0x | >100 bp [45] | Standard cfDNA recovery |
| 1.8x | >50 bp [45] | Short cfDNA/ctDNA fragments |
The underlying mechanism of this size-dependent binding involves the interplay between the crowding reagent (typically PEG) and the DNA molecules in solution. At lower bead ratios, the concentration of PEG is insufficient to drive smaller DNA fragments to bind to the beads, while larger fragments with more binding sites interact more strongly with the bead surface. As the bead ratio increases, the effective PEG concentration rises, enabling progressively smaller fragments to compete successfully for binding sites on the beads [26]. This biophysical principle allows the technology to function as a molecular "sieve" with tunable cutoff thresholds.
This size selection capability is particularly valuable for ctDNA analysis, where the target fragments typically exhibit a characteristic peak around 166 bp, corresponding to mononucleosomal DNA. By using appropriate bead ratios, researchers can selectively recover these informative fragments while excluding both shorter degradation products and longer genomic DNA contaminants that might originate from lysed blood cells [26] [37]. This purification enhances the tumor DNA fraction in the sample and improves the sensitivity of subsequent mutation detection assays. Additionally, the same principle can be applied to NGS library cleanup to remove undesirable adapter dimers (typically ~150 bp) without significant loss of the library fragments, simply by optimizing the bead ratio to retain fragments above a specific threshold [26].
Diagram 1: Mechanism of Magnetic Bead-Based Size Selection. The bead-to-sample ratio determines the effective PEG concentration, which modulates recovery efficiency in a DNA fragment size-dependent manner.
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful implementation of magnetic bead-based nucleic acid extraction requires specific reagents and equipment optimized for this technology. The following toolkit outlines essential components for establishing robust workflows in both research and clinical settings.
Table 3: Essential Research Reagent Solutions for Magnetic Bead-Based Extraction
| Component | Function | Examples/Formulations |
|---|---|---|
| Magnetic Beads | DNA binding through SPRI mechanism | Carboxyl-coated particles (400-600 nm); hydroxyl-functionalized beads [19] [46] |
| Binding Buffer | Create optimal conditions for DNA-bead interaction | Contains PEG, high salt (e.g., NaCl), and crowding agents [26] [19] |
| Wash Buffer | Remove contaminants while retaining bound DNA | Ethanol-based solutions (70-80%) with low salt concentration [19] [46] |
| Elution Buffer | Release purified DNA from beads | Low-salt physiological buffers (e.g., Tris-HCl, TE buffer) [45] [19] |
| Magnetic Separator | Immobilize beads during washing and elution | Magnetic racks for tubes, magnetic plates for 96-well formats [45] |
The magnetic beads themselves represent the core of this technology, with their performance heavily influenced by surface chemistry, size distribution, and solid content. Beads with carboxyl groups (-COOH) typically offer strong DNA binding capacity, while hydroxyl-functionalized (-OH) beads may provide different selectivity profiles. Optimal bead size generally falls in the 100-600 nm range, with specific formulations tailored for particular applications [19] [46]. The binding buffer formulation is equally critical, as it establishes the chemical environment that promotes reversible DNA immobilization on the beads. The precise combination of PEG molecular weight and concentration, salt type and ionic strength, and pH collectively determine the size selectivity and binding efficiency of the system [26].
For specialized applications such as ctDNA extraction, novel approaches have emerged that utilize homobifunctional crosslinkers like dimethyl suberimidate (DMS) to enhance recovery of low-abundance fragments. These crosslinkers facilitate binding between DNA and amine-conjugated magnetic beads through either covalent or electrostatic interactions, potentially offering higher extraction efficiency compared to traditional silica-based salt-bridge binding mechanisms [30]. When establishing a new workflow, empirical validation of both bead formulations and buffer compositions using standardized reference materials is recommended to ensure optimal performance for specific sample types and downstream applications.
The strategic optimization of bead-to-sample ratios represents a critical parameter for maximizing recovery and size selection in magnetic bead-based nucleic acid extraction systems. This technical capability provides researchers with unprecedented control over fragment selection, enabling precise targeting of specific DNA size ranges from complex biological samples. The comparative data presented in this guide demonstrates clear advantages of magnetic bead technology over traditional spin columns across multiple performance metrics, including recovery yield, size selection flexibility, throughput potential, and cost efficiency.
As liquid biopsy applications continue to evolve toward earlier disease detection and minimal residual disease monitoring, the demand for extraction methods capable of efficient recovery of low-abundance, fragmented DNA will only intensify. Magnetic bead technology, with its tunable size selection parameters and compatibility with automated workflows, is uniquely positioned to meet these challenges. Future developments in bead chemistry, buffer formulations, and protocol optimization will further enhance the performance and accessibility of this powerful technology, solidifying its role as an essential tool in both research and clinical diagnostics.
Preventing Contamination and Managing Elution Efficiency
The analysis of circulating tumor DNA (ctDNA) has revolutionized precision oncology by enabling non-invasive liquid biopsies for cancer diagnosis, monitoring treatment response, and detecting minimal residual disease [17] [37]. The reliability of these advanced molecular analyses is fundamentally dependent on the quality of the extracted ctDNA, making the prevention of contamination and optimization of elution efficiency critical considerations in method selection [47]. ctDNA presents unique technical challenges due to its exceptionally low concentration in plasma, often constituting less than 0.01% of total cell-free DNA, and its highly fragmented nature, typically ranging from 30 to 200 base pairs [17] [4]. These characteristics demand extraction methods capable of efficiently recovering short DNA fragments while maintaining sample purity.
The two dominant technologies in clinical ctDNA extraction are silica spin columns and magnetic bead-based systems, each with distinct mechanisms, advantages, and limitations in contamination control and elution characteristics [9] [17]. Spin column technology relies on the selective binding of DNA to a silica membrane under high-salt conditions, with subsequent wash steps to remove contaminants before elution in a low-salt buffer [9]. In contrast, magnetic bead methods utilize superparamagnetic particles coated with a DNA-binding surface that are separated from solution using a magnetic field, allowing for more flexible processing and reduced manual handling [9] [10]. Understanding the performance characteristics of these platforms is essential for laboratories implementing ctDNA testing to ensure analytical sensitivity and reproducibility while minimizing the risk of false results due to contamination or suboptimal DNA recovery.
This guide provides an objective comparison of spin column and magnetic bead-based ctDNA extraction methods, with a specific focus on their relative capabilities in preventing contamination and managing elution efficiency. By synthesizing experimental data from direct methodological comparisons and highlighting emerging technological innovations, we aim to provide researchers and clinical laboratory professionals with evidence-based guidance for selecting and optimizing ctDNA extraction protocols.
Spin Column Technology
Spin column-based DNA extraction operates on the principle of selective nucleic acid binding to a silica membrane in the presence of chaotropic salts [9]. The process begins with sample lysis to release DNA, followed by the addition of a binding buffer that facilitates DNA adhesion to the silica matrix. Subsequent wash steps with ethanol-based buffers remove proteins, salts, and other contaminants while the DNA remains bound to the membrane. The final elution step using low-ionic-strength buffer or water releases purified DNA from the silica surface [9] [30]. This technology has been extensively optimized for ctDNA extraction, with specialized commercial kits like the QIAamp Circulating Nucleic Acid Kit specifically designed to improve recovery of fragmented DNA [4] [48].
The physical architecture of spin columns presents both advantages and challenges for contamination prevention. The enclosed membrane system minimizes aerosol formation compared to liquid-phase methods, reducing cross-contamination risk between samples [9]. However, the requirement for multiple tube transfers and centrifugation steps introduces opportunities for sample mix-up and environmental contamination. For ctDNA applications, a significant limitation of conventional silica membranes is their reduced binding affinity for short DNA fragments, which can compromise elution efficiency and recovery of the most diagnostically relevant ctDNA species [4].
Magnetic Bead Technology
Magnetic bead-based extraction utilizes superparamagnetic particles, typically 0.5-5 µm in diameter, coated with silica or other DNA-binding surfaces [9] [30]. When added to a lysed sample, DNA binds to the bead surfaces through similar chaotropic salt-mediated interactions as spin columns. Application of a magnetic field immobilizes the beads against the tube wall, allowing efficient removal of contaminants in the supernatant through a series of wash steps. Purified DNA is then eluted from the beads in a small volume of low-salt buffer [30]. The magnetic separation mechanism eliminates the need for centrifugation and vacuum manifolds, streamlining processing and reducing opportunities for handling errors [9].
The dispersed nature of magnetic beads in solution provides a significant advantage for capturing short DNA fragments due to their high surface-area-to-volume ratio, potentially enhancing elution efficiency for ctDNA [30] [10]. Additionally, the closed-tube nature of magnetic separations minimizes aerosol formation, reducing cross-contamination risks. This platform also enables easier automation using liquid handling systems with integrated magnetic separation, further reducing manual intervention and associated contamination risks [9] [49]. Recent innovations include specialized bead coatings like homobifunctional crosslinkers that further improve ctDNA recovery through both covalent and electrostatic binding mechanisms [30].
Figure 1: Comparative Workflows of Spin Column vs. Magnetic Bead ctDNA Extraction Methods
Direct Performance Comparison: Experimental Data
Elution Efficiency and DNA Yield
Multiple studies have directly compared the elution efficiency and DNA recovery of spin column versus magnetic bead-based methods for ctDNA extraction. In a comprehensive methodological evaluation, silica spin column methods demonstrated limitations in recovering short DNA fragments below 100 base pairs, which constitute a significant proportion of ctDNA molecules [4]. This size-dependent recovery profile directly impacts elution efficiency, as fragments shorter than 80 base pairs showed dramatically reduced recovery using conventional silica-based methods compared to alternative approaches [4].
Magnetic bead technology has demonstrated superior performance in recovering the short DNA fragments characteristic of ctDNA. A 2022 study developing a novel magnetic bead-based approach using homobifunctional crosslinkers reported a 56% higher extraction efficiency compared to the QIAamp spin column kit [30]. This enhanced recovery is attributed to the greater binding surface area provided by dispersed magnetic beads and optimized surface chemistry that improves capture of short, fragmented DNA. The improved elution efficiency directly translated to better detection sensitivity for low-abundance mutations, a critical parameter for ctDNA analysis in early-stage cancers [30].
Beyond conventional silica magnetic beads, innovative approaches like magnetic ionic liquid (MIL)-based dispersive liquid-liquid microextraction have demonstrated significantly higher enrichment factors for multiple DNA fragments from human plasma compared to both silica-based and conventional magnetic bead methods [17]. Similarly, superparamagnetic bead particles in microfluidic platforms have achieved efficient ctDNA extraction with sensitivity of 65.57% and specificity of 95.38% in early-stage cancer samples [10]. These advanced magnetic separation techniques highlight the potential for further optimization of elution efficiency through material science innovations.
Table 1: Comparative Performance of ctDNA Extraction Technologies
| Performance Metric | Spin Column | Magnetic Bead | Experimental Basis |
|---|---|---|---|
| Recovery of <100 bp fragments | Limited (<35% for 25-40 bp fragments) [4] | Superior (60-90% for short fragments) [17] [4] | Fragment size recovery analysis using spiked synthetic DNA targets [4] |
| Total DNA yield | Standardized yields; may be lower for fragmented DNA [9] | 56% higher yield compared to spin column [30] | Direct comparison using plasma samples from cancer patients [30] |
| Mutation detection sensitivity | May miss low-frequency mutations due to fragment loss [4] | 171% increase in mutant copy recovery [48] | ddPCR analysis of clinical plasma samples with known mutations [48] |
| Processing time | 30-90 minutes (manual processing) [9] | <10 minutes possible with rapid protocols [30] | Method development studies timing complete extraction workflows [30] |
| Automation compatibility | Limited; requires centrifugation or vacuum steps [9] | High; easily integrated with liquid handling systems [9] [49] | Automated platform development and validation studies [49] |
Contamination Control and Sample Purity
Contamination prevention encompasses multiple aspects, including cross-contamination between samples, environmental DNA contamination, and carryover of PCR inhibitors that can compromise downstream analyses. The physical separation mechanism in spin column systems, where the silica membrane serves as a barrier between the sample and eluate, provides inherent protection against carryover of inhibitors from the original sample [9]. However, the requirement for multiple tube transfers and the potential for aerosol formation during centrifugation present opportunities for cross-contamination, particularly in high-throughput settings processing concentrated genomic DNA samples alongside low-concentration ctDNA samples [9].
Magnetic bead systems offer advantages in contamination control through reduced manual handling and elimination of centrifugation steps. The ability to process samples in closed-tube formats significantly reduces aerosol formation, thereby lowering cross-contamination risks [9] [49]. This characteristic makes magnetic bead systems particularly suitable for automated workflows, where minimal human intervention further decreases contamination opportunities. Additionally, magnetic separation allows for more efficient washing, as beads remain immobilized while wash buffers are added and removed, potentially improving removal of PCR inhibitors compared to spin column washing [30].
Microfluidic implementations of both technologies have demonstrated enhanced contamination control by further minimizing manual handling. Integrated microfluidic devices for ctDNA isolation can process samples with minimal exposure to the environment, reducing both cross-contamination and introduction of external contaminants [17] [10]. These systems often incorporate dedicated channels or chambers for different samples, virtually eliminating cross-contamination while improving reproducibility through precise fluidic control.
Advanced Methods and Emerging Technologies
Alternative Extraction Chemistry
While silica-based binding dominates both spin column and magnetic bead technologies, alternative chemistries offer promising avenues for improving elution efficiency and reducing contamination. Liquid-phase extraction methods utilizing aqueous two-phase systems (ATPSs) represent a fundamentally different approach that bypasses solid-phase binding altogether. The PHASIFY method, which employs a series of ATPSs with optimized formulations, demonstrated a 60% increase in DNA yield and 171% increase in mutant copy recovery compared to the QIAamp spin column kit [48]. This enhanced performance is achieved through selective partitioning of cfDNA into specific phases while excluding contaminants, followed by volume reduction to concentrate the target molecules.
Homobifunctional crosslinkers represent another innovative approach that can be integrated with magnetic bead systems. These compounds, such as dimethyl suberimidate (DMS), bind to DNA through both covalent and electrostatic interactions, then subsequently bind to amine-conjugated magnetic beads [30]. This mechanism enables rapid extraction (within 10 minutes) with significantly higher efficiency than conventional chaotropic salt-based methods. The strong binding specificity of these crosslinkers may also reduce co-elution of contaminants, though comprehensive purity comparisons with established methods are still needed [30].
Hybrid approaches that combine multiple separation mechanisms have also shown promise for enhancing elution efficiency while controlling contamination. For example, the Q Sepharose method utilizes anion exchange resin to preconcentrate DNA before desalting on a silica spin column, achieving 60-90% recovery of short DNA fragments [4]. Similarly, magnetic nanowire networks with elongated or tubular morphologies and high saturation magnetization have demonstrated efficient cfDNA capture while minimizing loss and degradation [17]. These multi-mechanism approaches can leverage the strengths of different technologies to optimize overall performance.
Microfluidic and Automated Platforms
Microfluidic technologies represent a paradigm shift in ctDNA extraction, offering integrated solutions that minimize contamination risks while optimizing elution efficiency through precise fluidic control. These devices can implement both solid-phase and liquid-phase extraction techniques on-chip, with classifications including functionalized surfaces, immobilized beads, chemical reagents, or electrophoretic methods for DNA isolation [17]. The miniaturized dimensions and controlled flow conditions in microfluidic devices enhance mass transfer, potentially improving binding efficiency for low-concentration ctDNA targets [17] [10].
Automated systems leveraging magnetic bead technology provide standardized processing that reduces variability in elution efficiency while minimizing contamination opportunities. A 2024 study comparing silica spin-column and magnetic bead kits for DNA methylation analysis found that automation with magnetic bead kits increased throughput without compromising amplification efficiency in PCR systems [49]. The consistency achieved through automated processing is particularly valuable for ctDNA analysis, where small variations in extraction efficiency can significantly impact detection sensitivity for low-abundance mutations [49] [47].
Integrated microfluidic platforms specifically designed for ctDNA extraction have demonstrated remarkable performance characteristics. One simulation study of a superparamagnetic bead-based microfluidic device reported extraction of an average of 5.7 ng of ctDNA from every 10 µL of blood plasma input, with sensitivity of 65.57% and specificity of 95.38% for stage I and II cancer patients [10]. These systems achieve contamination control through disposable cartridges or self-contained processing chambers, while their optimized fluidic designs enhance elution efficiency through controlled binding and washing kinetics.
Table 2: Research Reagent Solutions for ctDNA Extraction
| Reagent/Category | Specific Examples | Function in ctDNA Extraction |
|---|---|---|
| Spin Column Kits | QIAamp Circulating Nucleic Acid Kit; Cobas ccfDNA Sample Preparation Kit [37] [48] | Silica-membrane based isolation using chaotropic salt binding and centrifugal purification |
| Magnetic Bead Kits | QIAamp MinElute ccfDNA Mini Kit; Maxwell RSC LV ccfDNA Kit; MagNa Pure 24 Total NA Isolation Kit [37] | Paramagnetic particle-based isolation with magnetic separation and washing |
| Specialized Blood Collection Tubes | cfDNA BCT (Streck); PAXgene Blood ccfDNA (Qiagen); cfDNA/cfRNA Preservative (Norgene) [17] [37] | Contain preservatives to prevent leukocyte lysis and genomic DNA contamination during sample transport and storage |
| Alternative Chemistry Reagents | Homobifunctional crosslinkers (DMS, DMA, DTBP) [30]; Aqueous two-phase system components [48] | Enable novel binding mechanisms beyond silica-chaotropic salt interactions for enhanced recovery |
| Microfluidic Components | Superparamagnetic bead particles; Surface functionalized channels; Integrated valves and pumps [17] [10] | Enable miniaturized, automated extraction with reduced contamination risk and enhanced efficiency |
Methodological Protocols
Standardized Spin Column Protocol
The QIAamp Circulating Nucleic Acid Kit protocol exemplifies a standardized approach for spin column-based ctDNA extraction [30] [48]. The process begins with mixing plasma samples (typically 1-5 mL) with lysis buffer containing proteinase K, followed by incubation at 60°C for 30 minutes to digest proteins and release nucleic acids. Binding buffer is then added to create high-salt conditions favorable for DNA adsorption to silica, and the mixture is applied to the spin column mounted on a vacuum manifold or centrifuge. Vacuum or centrifugal force passes the solution through the silica membrane, retaining DNA while removing contaminants.
Wash steps typically involve two different wash buffers: the first containing guanidine salts to remove proteins and the second containing ethanol to remove salts and other impurities [30]. Each wash requires centrifugation or vacuum application to pass the buffer through the membrane while retaining bound DNA. The final elution step uses a low-ionic-strength buffer (typically TE buffer or nuclease-free water) incubated on the membrane for 3-5 minutes before centrifugation to collect purified ctDNA. Critical considerations for optimizing elution efficiency include pre-warming the elution buffer to 60-70°C and performing two sequential elutions to maximize recovery [48].
Magnetic Bead Protocol with Enhanced Chemistry
A representative protocol for magnetic bead extraction using homobifunctional crosslinkers demonstrates the potential for enhanced elution efficiency [30]. The process begins with plasma sample lysis using buffer containing proteinase K at 60°C for 30 minutes. The homobifunctional crosslinker DMS is then added to the lysate and vortexed for 10 seconds, during which it immediately binds to DNA through covalent and electrostatic interactions. Amine-coated magnetic beads are subsequently added, and the mixture is incubated with gentle rocking to facilitate binding between the beads and DMS-DNA complexes.
Magnetic separation immobilizes the beads against the tube wall, allowing supernatant removal. Wash steps typically involve resuspending the beads in phosphate-buffered saline (PBS) at pH 7.4 to remove impurities, followed by repeated magnetic separation. Elution uses 0.01 M sodium bicarbonate adjusted to pH 10.3, with vortexing for 2 minutes and incubation at room temperature for 3 minutes to break the crosslinking [30]. The alkaline elution condition and mechanical disruption enhance elution efficiency, contributing to the 56% higher recovery compared to standard spin column methods. The entire process can be completed within 10 minutes, significantly faster than conventional protocols.
Figure 2: Critical Factors in Contamination Prevention and Elution Efficiency
The selection between spin column and magnetic bead technologies for ctDNA extraction involves careful consideration of their respective capabilities in contamination prevention and elution efficiency management. Spin column methods offer the advantage of established protocols and physical separation barriers that reduce certain contamination risks, but they exhibit limitations in recovering short DNA fragments essential for sensitive ctDNA analysis [9] [4]. Magnetic bead systems provide superior recovery of fragmented DNA, higher compatibility with automation, and reduced aerosol formation, making them increasingly suitable for high-throughput clinical applications where sensitivity and reproducibility are paramount [9] [30] [49].
Emerging technologies, including advanced magnetic bead chemistries, liquid-phase extraction methods, and integrated microfluidic platforms, offer promising avenues for further improving both contamination control and elution efficiency [17] [30] [48]. These innovations demonstrate that significant gains in ctDNA recovery and analysis sensitivity are achievable through fundamental rethinking of extraction mechanisms rather than incremental optimization of existing platforms. As ctDNA analysis expands into earlier cancer detection and minimal residual disease monitoring, where DNA concentrations are lowest and fragment sizes are smallest, these advanced extraction technologies will become increasingly essential for clinical utility.
Laboratories implementing ctDNA testing should consider their specific application requirements, sample volumes, and available infrastructure when selecting extraction methods. For applications demanding maximum sensitivity for low-frequency mutations, magnetic bead-based methods currently offer advantages in elution efficiency and recovery of short DNA fragments. Regardless of the technology selected, adherence to standardized protocols, implementation of appropriate contamination controls, and rigorous validation of elution efficiency are essential for generating reliable, clinically actionable results from ctDNA analysis [47].
The analysis of circulating tumor DNA (ctDNA) has emerged as a cornerstone of liquid biopsy in precision oncology, enabling non-invasive cancer detection, monitoring of minimal residual disease, assessment of treatment efficacy, and tracking of tumor evolution [17] [50]. This fragmented DNA, released into the bloodstream by tumor cells through apoptosis and necrosis, carries the identical genomic alterations found in the primary tumor, offering a real-time snapshot of tumor heterogeneity [17]. However, the reliable detection of ctDNA presents substantial technical challenges due to its extremely low abundance in plasma, where it is heavily diluted by background wild-type cell-free DNA (cfDNA) derived from healthy cells [17] [50]. This biological limitation places exceptional importance on the pre-analytical phase, where the choice of nucleic acid extraction method becomes a pivotal determinant of the sensitivity, specificity, and overall success of downstream analytical applications.
Among the available extraction technologies, two methods have become predominant in clinical and research settings: silica membrane-based spin columns and magnetic bead-based purification. The strategic selection between these platforms requires a thorough understanding of their respective operational principles, performance characteristics, and economic implications. This guide provides an objective, data-driven comparison of these technologies, focusing specifically on their application in ctDNA analysis. By synthesizing current experimental data and methodological protocols, we aim to equip researchers and drug development professionals with the evidence necessary to align their extraction strategy with specific project goals, sample types, and operational constraints, thereby optimizing the reliability and efficiency of their liquid biopsy workflows.
Fundamental Principles and Methodologies
The effective isolation of ctDNA hinges on the specific binding of DNA to a solid surface in the presence of chaotropic salts, followed by washing and elution. While spin columns and magnetic beads both utilize a silica surface for binding, their mechanisms of manipulation and separation differ significantly, influencing their application suitability.
Spin Column-Based Extraction
Spin column technology relies on the selective binding of DNA to a silica membrane housed within a centrifugal column under high-salt conditions [9] [51]. The operational workflow begins with sample lysis to release nucleic acids, followed by the addition of a binding buffer to create the optimal chemical environment for DNA to adhere to the silica membrane [9]. During subsequent centrifugation steps, the liquid sample passes through the membrane, while contaminants and impurities are washed away using ethanol-based buffers. Finally, the purified DNA is eluted in a low-salt buffer or nuclease-free water [51]. This method is particularly noted for its ability to recover variable-sized DNA fragments, including high molecular weight fragments greater than 600 base pairs, making it a robust choice for general ctDNA isolation [17].
Magnetic Bead-Based Extraction
Magnetic bead technology utilizes paramagnetic particles coated with a silica surface that reversibly binds nucleic acids [9] [52]. The process initiates with sample lysis, after which the magnetic beads are added to the solution. DNA binds to the bead surfaces in the presence of a binding buffer containing crowding agents like polyethylene glycol (PEG) and salt [53]. The fundamental differentiator of this method is the use of a magnetic field, rather than centrifugation, to immobilize the bead-DNA complexes, allowing the supernatant containing impurities to be simply aspirated or decanted [9] [52]. After washing steps to remove residual contaminants, the purified DNA is eluted from the beads. This magnetic separation mechanism is inherently scalable and forms the basis for automation, making it exceptionally well-suited for high-throughput environments [9] [53]. Furthermore, magnetic bead systems demonstrate high efficiency in recovering the smaller DNA fragments that are characteristic of ctDNA [17].
Workflow Comparison
The following diagram illustrates the core procedural differences between the spin column and magnetic bead extraction workflows:
Comparative Performance Analysis
Yield, Purity, and Fragment Size Recovery
The recovery efficiency of ctDNA, particularly its shorter fragments, is a critical performance metric. A 2024 study directly comparing silica spin-columns and magnetic beads for DNA methylation analysis in liquid cytology samples found that both technologies exhibited comparable amplification efficiency in downstream quantitative methylation-specific PCR (qMSP) [49]. This suggests that when optimized, both methods can provide DNA of sufficient quality for sensitive detection assays.
However, the choice of method can significantly impact the recovery of mutant DNA copies, which is paramount for ctDNA analysis. A 2021 study in Scientific Reports compared a novel liquid-phase extraction method (PHASIFY) to a standard solid-phase method (QIAamp Circulating Nucleic Acid kit, a spin-column method) [48]. The results demonstrated that the spin-column method struggled with recovery efficiency. The PHASIFY MAX method, which utilizes aqueous two-phase systems, showed a 60% increase in total DNA yield and a striking 171% increase in mutant copy recovery compared to the spin-column technique [48]. This indicates a potential limitation of traditional spin columns in capturing the full spectrum of ctDNA, especially from samples with low mutant allele frequencies.
Regarding fragment size, spin columns are generally considered effective for recovering variable-sized DNA, including high molecular weight fragments (>600 bp) [17]. In contrast, magnetic bead-based systems are particularly efficient at recovering smaller DNA fragments [17]. This characteristic is advantageous for ctDNA, which is highly fragmented. Furthermore, specialized magnetic bead kits like PHASIFY ENRICH can incorporate size-selection steps to remove high molecular weight contaminating genomic DNA (>500 bp), thereby enriching for the smaller cfDNA fraction and improving the effective tumor fraction [48].
Operational and Economic Considerations
Throughput, automation compatibility, and cost are decisive factors in method selection, especially for clinical and large-scale research settings.
Throughput and Automation: Magnetic bead-based systems hold a distinct advantage in high-throughput and automated workflows. The magnetic separation process is inherently amenable to automation on platforms such as the Thermo Fisher KingFisher, Hamilton Microlab STAR, and Beckman Coulter Biomek i-Series [53]. This compatibility enables simultaneous processing of 96 or more samples, drastically reducing hands-on time and improving reproducibility [52] [53]. Spin column methods, which require repeated individual centrifugation steps, are labor-intensive for batch processing and not readily automation-compatible, making them less ideal for high-volume laboratories [51] [53].
Cost Structure: A comprehensive cost analysis must consider both consumable expenses and labor. As shown in Table 1, magnetic bead-based cleanup offers a significantly lower cost per reaction compared to spin columns [53]. This cost advantage is amplified when processing thousands of samples. Furthermore, the reduced hands-on time and potential for integration into automated systems lower overall labor costs [53]. Spin columns, while having a lower initial equipment outlay, incur higher consumable costs and require more extensive personnel time, making them less economical for large-scale studies [52] [53].
Table 1: Operational and Economic Comparison of DNA Extraction Methods
| Feature | Magnetic Bead-Based | Spin Column-Based |
|---|---|---|
| Throughput | High (96-well & automation compatible) [53] | Low (manual, single-tube format) [51] [53] |
| Automation Compatibility | Yes [9] [53] | No [53] |
| Protocol Time (Manual) | <15 minutes [53] | 20-30 minutes [53] |
| Hands-on Time | Low (especially when automated) [52] | High [51] |
| Cost per Sample | ~$0.90 [53] | ~$1.75 [53] |
| Cost per 1000 Samples | ~$900 [53] | ~$1,750 [53] |
| Initial Equipment Cost | Higher (requires magnetic separator) [9] | Lower (requires centrifuge) [9] |
Essential Research Reagent Solutions
The successful implementation of ctDNA extraction workflows depends on a suite of specialized reagents and tools. The following table details key components and their functions in the experimental process.
Table 2: Key Research Reagent Solutions for ctDNA Extraction
| Item | Function/Description | Application Context |
|---|---|---|
| Silica Spin Columns | Centrifugal devices with a silica membrane that binds DNA in high-salt conditions [9] [51]. | Core component of spin-column extraction kits for manual, low-throughput DNA purification. |
| Silica-Coated Magnetic Beads | Paramagnetic particles with a silica surface for DNA binding; separated using a magnet [9] [52]. | Core component of magnetic bead-based kits for manual or automated, high-throughput extraction. |
| Binding Buffer (High Salt) | Creates conditions (chaotropic salts) that promote DNA binding to silica surfaces [9] [51]. | Used in both spin column and magnetic bead protocols during the initial binding step. |
| Wash Buffer (Ethanol-based) | Removes salts, proteins, and other contaminants while keeping DNA bound to the silica matrix [51] [53]. | Used in wash steps for both main methods; typically contains ethanol and buffering salts. |
| Elution Buffer (Low Salt) | Low-ionic-strength solution (e.g., TE buffer or nuclease-free water) that disrupts DNA-silica binding, releasing purified DNA [9] [51]. | Final step in all silica-based purification protocols to elute DNA into a small volume. |
| Magnetic Rack/Plate | Device that generates a magnetic field to capture and hold magnetic beads during washing and elution [52] [53]. | Essential equipment for manual magnetic bead-based protocols. |
| Automated Nucleic Acid Extractor | Robotic systems (e.g., KingFisher, Biomek) that automate the entire magnetic bead separation process [53]. | Used for high-throughput, walk-away automation of magnetic bead-based extraction. |
| Blood Collection Tubes (BCTs) with Stabilizers | Tubes (e.g., Streck, Roche) containing reagents that prevent white blood cell lysis and preserve ctDNA for up to several days [17]. | Critical pre-analytical step for blood collection to ensure ctDNA quality before extraction. |
Detailed Experimental Protocols
To ensure reproducibility and facilitate method validation, detailed protocols for key experiments cited in this guide are provided below.
Protocol: Magnetic Bead-Based PCR Cleanup with HighPrep
This protocol, adapted from MagBio Genomics, is representative of a magnetic bead cleanup workflow suitable for post-PCR ctDNA analysis [53].
- Binding: Combine the PCR reaction or other DNA sample with a 1.8x volume of HighPrep PCR beads in a tube or plate well. Mix thoroughly by pipetting or vortexing to ensure homogeneity.
- Incubation: Allow the mixture to stand at room temperature for 5 minutes to facilitate DNA binding to the beads.
- Separation: Place the tube or plate on a magnetic stand. Wait for approximately 2 minutes, or until the solution clears and the beads form a tight pellet against the magnet.
- Washing: Carefully aspirate and discard the supernatant without disturbing the bead pellet. While the tube remains on the magnet, add 200 µL of freshly prepared 80% ethanol. Incubate for 30 seconds, then aspirate and discard the ethanol. Repeat this wash step a second time for a total of two washes.
- Drying: With the tube still on the magnet, air-dry the bead pellet for 3-5 minutes at room temperature. It is critical to avoid over-drying, as this can reduce DNA elution efficiency. The pellet should appear dry but not cracked.
- Elution: Remove the tube from the magnetic stand. Resuspend the dried beads in 20-50 µL of nuclease-free water or TE buffer (10 mM Tris-HCl, pH 8.0, 0.1 mM EDTA) by pipetting up and down. Incubate for 2 minutes at room temperature.
- Final Separation: Return the tube to the magnetic stand. After the solution clears (approximately 2 minutes), transfer the eluate containing the purified DNA to a new tube.
Size Selection Modification: The target DNA fragment size range can be customized by adjusting the bead-to-sample ratio. For example, a 0.6x ratio retains fragments >500 bp, while a 1.8x ratio retains fragments >50 bp [53].
Protocol: Spin Column-Based DNA Extraction
This protocol outlines the generic steps for a silica spin-column-based DNA extraction, as used in kits such as the QIAamp Circulating Nucleic Acid kit [51] [48].
- Lysis: Add a proteinase K-containing lysis buffer to the plasma sample. Incubate at the recommended temperature (often 56°C) to degrade proteins and release nucleic acids.
- Binding: Add a binding buffer containing chaotropic salts (e.g., guanidine hydrochloride) to the lysate. Transfer the entire mixture to a spin column seated in a collection tube.
- Centrifugation: Centrifuge the column at high speed (e.g., 13,000 g) for 1 minute. The DNA binds to the silica membrane while the flow-through containing impurities is discarded.
- Washing: Place the column back into the collection tube. Add a wash buffer (typically containing ethanol) to the column. Centrifuge as before and discard the flow-through. This step is often repeated with a second, different wash buffer to ensure removal of all contaminants.
- Final Spin: Perform an additional centrifugation step for 1-2 minutes with the empty column to remove residual ethanol. This is crucial for preventing interference in downstream applications.
- Elution: Transfer the spin column to a clean 1.5 mL microcentrifuge tube. Apply 20-100 µL of elution buffer (or nuclease-free water) directly onto the center of the silica membrane. Incubate at room temperature for 5 minutes to allow the buffer to diffuse.
- DNA Elution: Centrifuge the column for 2 minutes at full speed to collect the purified DNA in the microcentrifuge tube.
The choice between magnetic bead and spin column technologies for ctDNA extraction is not a matter of one being universally superior, but rather of strategic alignment with specific application requirements and operational contexts. The following decision diagram synthesizes the key comparative data to guide researchers in this selection process:
In conclusion, magnetic bead-based extraction is the recommended strategy for high-throughput laboratories, automated workflows, studies with limited sample material, and projects where maximizing the recovery of mutant ctDNA copies is the highest priority, as it offers superior efficiency, scalability, and lower long-term costs [9] [53] [48]. Conversely, spin column-based extraction remains a viable option for low-to-moderate throughput research settings, such as academic laboratories, where the initial investment in equipment is a constraint, the number of samples is manageable, and the primary need is for reliable, high-quality DNA without the demand for ultimate recovery sensitivity [9] [51].
The field of liquid biopsy continues to evolve rapidly, with emerging technologies like liquid-phase extraction [48] and microfluidic devices [17] promising even greater sensitivity and integration. A thoughtful cost-benefit analysis, grounded in the experimental data and operational considerations presented here, will ensure that the selected DNA extraction method serves as a robust foundation for accurate and clinically meaningful ctDNA analysis.
Head-to-Head Validation: A Data-Driven Comparison
The analysis of circulating tumor DNA (ctDNA) has revolutionized liquid biopsy, enabling non-invasive cancer diagnosis, monitoring of treatment efficacy, and assessment of minimal residual disease [37] [16]. Efficient extraction of ctDNA from blood plasma is a critical pre-analytical step that directly impacts the sensitivity and reliability of downstream molecular analyses such as droplet digital PCR (ddPCR) and next-generation sequencing (NGS) [30] [48]. ctDNA presents unique analytical challenges due to its extremely low concentration in plasma (often less than 0.01% of total cell-free DNA), highly fragmented nature (typically ~166 bp), and short half-life (16-150 minutes) [30] [25].
Among the various extraction technologies available, silica-based spin columns and magnetic bead-based methods have emerged as the most widely used approaches in both research and clinical settings [16] [25]. This review provides a comprehensive comparison of these two dominant technologies, focusing on recovery yield, purity, and performance in clinical studies, to guide researchers and clinicians in selecting the optimal extraction method for liquid biopsy applications.
Fundamental Principles
Spin column and magnetic bead technologies both utilize silica-based binding surfaces but differ substantially in their operational mechanisms. Spin columns employ a silica membrane embedded in a column format through which samples are passed under centrifugal force. Nucleic acids bind to the silica surface in the presence of high-chaotropic salt conditions, while contaminants are removed through wash steps. The purified DNA is then eluted in a low-salt buffer [9] [25].
Magnetic bead methods utilize silica-coated paramagnetic particles that are mixed with the sample solution. In the presence of binding buffers, nucleic acids adsorb to the bead surface. Using a magnetic field, the bead-DNA complexes are separated from the solution, washed to remove impurities, and the DNA is subsequently eluted [9] [24]. This method enables more flexible processing and is more readily automated than spin column approaches.
Key Technical Differentiators
The fundamental differences in how these methods handle separation lead to distinct practical implications:
- Processing mechanism: Spin columns rely on centrifugal force, while magnetic beads use magnetic separation [9] [25].
- Automation compatibility: Magnetic bead systems are more easily integrated into high-throughput automated workflows [9] [46].
- Sample handling: Spin columns require multiple tube transfers and column changes, increasing hands-on time and contamination risk [25].
- Binding surface accessibility: Magnetic beads provide a larger surface area-to-volume ratio, potentially enhancing recovery of low-abundance fragments [24].
Comparative Performance Data
Multiple clinical studies have directly compared the performance of spin column and magnetic bead-based extraction methods for ctDNA recovery. The table below summarizes key quantitative findings from these investigations.
Table 1: Comparative Performance of Spin Column vs. Magnetic Bead-Based ctDNA Extraction Methods
| Evaluation Metric | Spin Column (Reference Method) | Magnetic Bead Method | Improvement vs. Reference | Study Details |
|---|---|---|---|---|
| Total cfDNA Yield | QIAamp kit (Baseline) | DMS-crosslinked magnetic beads [30] | +56% higher extraction efficiency | 1 mL plasma samples; crosslinker enhanced binding |
| Mutant Copy Recovery | QIAamp Circulating Nucleic Acid Kit (Baseline) | PHASIFY MAX (liquid-phase) [48] | +171% increase in mutant copies | 91 cancer patient plasma samples; ddPCR detection |
| Extraction Efficiency | QIAamp kit (Baseline) | SHIFT-SP (optimized magnetic beads) [24] | 98.2% binding efficiency at pH 4.1 | 100 ng input DNA; optimized binding conditions |
| Fragment Size Recovery | Conventional spin columns | Magnetic beads (various studies) | Better recovery of <100 bp fragments | Multiple analytical controls |
| Processing Time | ~30 minutes (manual) [25] | 6-15 minutes (optimized/automated) [30] [24] | ~50-80% reduction | Excludes plasma processing steps |
| Purity (Protein Contamination) | Standard spin columns | Aqueous Two-Phase Systems (ATPS) [54] | Reduced to ~3 mg/mL total protein | PEG/phosphate systems |
Recovery Yield Analysis
Recovery yield represents the most critical parameter for ctDNA extraction due to the low abundance of tumor-derived DNA in plasma. Clinical studies consistently demonstrate that optimized magnetic bead methods can significantly enhance DNA recovery compared to standard spin column techniques.
In a comprehensive clinical validation study comparing the QIAamp Circulating Nucleic Acid Kit (spin column) versus the PHASIFY MAX method (liquid-phase extraction with ATPS) across 91 plasma samples from cancer patients, the magnetic bead-based approach demonstrated a 60% increase in total DNA yield and a remarkable 171% increase in mutant copy recovery as detected by ddPCR [48]. This enhanced recovery of mutant alleles is particularly valuable for detecting minimal residual disease and early-stage cancers where ctDNA fractions are exceedingly low.
Similarly, a novel magnetic bead-based method utilizing homobifunctional crosslinkers (e.g., dimethyl suberimidate dihydrochloride) demonstrated a 56% higher extraction efficiency compared to the QIAamp kit [30]. The crosslinker facilitates rapid binding between DNA and amine-coated magnetic beads through both covalent and electrostatic interactions, improving recovery of low-concentration fragments.
Purity and Fragment Size Distribution
The purity of extracted ctDNA is crucial for downstream analytical performance, as contaminants can inhibit enzymatic reactions in PCR and NGS. Spin column methods generally provide high-purity DNA with minimal protein contamination [23] [25]. However, several advanced magnetic bead approaches have demonstrated comparable or superior purification capabilities.
Aqueous two-phase systems (ATPS) employing polyethylene glycol (PEG) and phosphate solutions have achieved total protein reduction to approximately 3 mg/mL while maintaining up to 90% DNA recovery [54]. This liquid-phase extraction effectively separates DNA from protein contaminants through partitioning between immiscible phases.
For fragment size bias, evidence suggests that magnetic bead methods may better recover shorter DNA fragments (<100 bp) that are characteristic of ctDNA [16]. This size distribution advantage potentially enables more comprehensive capture of the ctDNA population, though the clinical significance of very short fragments requires further investigation.
Experimental Protocols and Methodologies
Standardized Workflow for Method Comparison
To ensure valid comparisons between extraction technologies, researchers should implement standardized protocols from blood collection through downstream analysis. The following workflow diagram illustrates key steps in a typical method comparison study:
Detailed Extraction Protocols
Spin Column Protocol (QIAamp Circulating Nucleic Acid Kit)
The QIAamp kit represents the benchmark spin column method used in numerous comparison studies [48] [23]. The standard protocol involves:
- Sample Lysis: Mix 1 mL plasma with 1 μg carrier RNA and proteinase K, then incubate at 60°C for 30 minutes [30].
- Binding Conditions: Add binding buffer ACB (nine-fifths of plasma volume), vortex for 30 seconds, and incubate on ice for 5 minutes.
- Column Processing: Apply the mixture to a QIAamp Mini column mounted on a vacuum manifold. Draw liquid through the silica membrane by vacuum pressure.
- Washing: Perform two wash steps using buffers AW1 and AW2 with centrifugation at 6,000 × g for 1 minute and 20,000 × g for 3 minutes, respectively.
- Elution: Add elution buffer (40-60 μL) to the membrane, incubate for 1 minute, then centrifuge at 6,000 × g for 1 minute to recover purified cfDNA [30] [23].
Magnetic Bead Protocol (DMS-Crosslinked Beads)
An optimized magnetic bead protocol demonstrating superior recovery includes [30]:
- Sample Preparation: Mix 1 mL plasma with lysis buffer containing proteinase K and incubate at 60°C for 30 minutes.
- Crosslinker Binding: Add dimethyl suberimidate (DMS) solution to plasma and vortex for 10 seconds to facilitate immediate DMS-DNA complex formation.
- Bead Binding: Add amine-coated magnetic beads and incubate with gentle mixing (50 rpm) at room temperature for 10 minutes.
- Magnetic Separation: Place tubes on a magnetic rack for 1 minute to separate bead-DNA complexes, then discard supernatant.
- Washing: Wash beads twice with 2 mL PBS (pH 7.4) using magnetic separation between washes.
- Elution: Elute DNA using 100 μL of 0.01 M sodium bicarbonate (pH 10.3) with vortexing for 2 minutes and incubation at room temperature for 3 minutes to break crosslinks.
- Final Recovery: Collect supernatant containing purified cfDNA after magnetic separation.
Quality Assessment Methods
To ensure comparable results across studies, researchers employ standardized quality assessment:
- Yield Quantification: Fluorometric methods (Qubit, DeNovix) for precise DNA concentration measurement [30] [46].
- Fragment Size Analysis: Microelectrophoresis (Bioanalyzer, TapeStation) to evaluate DNA size distribution and integrity [48] [23].
- Purity Assessment: UV spectrophotometry (A260/A280 ratio) and protein quantification assays to detect contaminants [54].
- Functional Validation: ddPCR or qPCR for mutant allele detection and extraction efficiency calculation [48] [24].
The Scientist's Toolkit: Essential Research Reagents
Successful ctDNA extraction requires careful selection of reagents and materials throughout the workflow. The following table outlines key solutions and their functions in the extraction process:
Table 2: Essential Research Reagents for ctDNA Extraction Studies
| Reagent/Category | Specific Examples | Function & Importance |
|---|---|---|
| Blood Collection Tubes | Streck cfDNA BCT, PAXgene Blood ccfDNA, CellSave | Preserve ctDNA integrity by inhibiting leukocyte lysis and nuclease activity during transport/storage [37] [16] |
| Binding Buffers | Chaotropic salt solutions (guanidine HCl), PEG-based buffers | Enable nucleic acid binding to silica surfaces by altering DNA hydration shell and promoting adsorption [24] [46] |
| Magnetic Beads | Silica-coated beads, amine-functionalized beads, in-house formulations (e.g., SafeCAP 2.0) | Solid phase for DNA capture; composition and surface chemistry critically impact recovery of short fragments [30] [46] |
| Crosslinking Reagents | Dimethyl suberimidate (DMS), DTBP | Enhance DNA-bead binding efficiency through covalent and electrostatic interactions [30] |
| Wash Buffers | Ethanol-based solutions (70-80%) with controlled salt concentrations | Remove contaminants while retaining bound DNA; optimize stringency to balance purity and yield [30] [46] |
| Elution Buffers | Low-salt solutions (Tris-HCl, TE buffer), alkaline buffers (pH 10.3) | Disrupt DNA-silica interaction while maintaining DNA stability and compatibility with downstream assays [30] [24] |
Discussion and Future Perspectives
The cumulative evidence from clinical studies indicates that magnetic bead-based extraction methods generally provide superior recovery yields compared to traditional spin column techniques, particularly for the low-abundance, fragmented ctDNA molecules that are most clinically relevant [48] [24]. This enhanced recovery directly translates to improved detection sensitivity for mutant alleles, potentially enabling earlier cancer detection and more reliable monitoring of minimal residual disease.
However, spin column methods maintain advantages in certain applications, offering robust performance, proven reliability, and high purity [23] [25]. The choice between technologies should consider specific research or clinical needs:
- For maximal sensitivity in detecting low-frequency mutations, magnetic bead methods with optimized chemistry (e.g., crosslinkers, surface functionalization) are preferable.
- For routine applications with adequate ctDNA abundance, spin columns provide a reliable, established methodology.
- In high-throughput settings, automated magnetic bead platforms offer significant advantages in processing efficiency and reproducibility [9] [46].
Future developments in ctDNA extraction will likely focus on further reducing fragment size bias, integrating extraction with downstream analysis in microfluidic systems [25], and standardizing protocols across laboratories to improve reproducibility [37]. The emerging generation of extraction technologies demonstrates that continued optimization of both bead-based and alternative liquid-phase methods holds promise for unlocking the full potential of liquid biopsy in precision oncology.
Performance in Low-Abundance and Fragmented ctDNA Contexts
The analysis of circulating tumor DNA (ctDNA) has revolutionized oncology, enabling non-invasive liquid biopsies for cancer diagnosis, treatment monitoring, and recurrence surveillance. ctDNA represents a fragile subset of cell-free DNA (cfDNA) that originates from tumor cells and carries tumor-specific genetic alterations. However, its reliable detection presents a significant bioanalytical challenge due to its extremely low abundance in blood—often constituting less than 0.1% of total cfDNA in early-stage cancer—and its highly fragmented nature, with a dominant fraction around 167 base pairs corresponding to mononucleosomal DNA [17] [55]. These characteristics create a pressing need for extraction methods that can efficiently isolate these scarce, fragmented molecules while preserving their integrity for downstream molecular analyses.
Two primary technological approaches dominate ctDNA extraction workflows: silica membrane-based spin columns and magnetic bead-based methods. While both utilize silica-based binding chemistry, their fundamental operational principles differ substantially, leading to important performance trade-offs in recovery efficiency, fragment size bias, automation compatibility, and procedural workflow [9] [56]. This guide provides an objective, data-driven comparison of these competing technologies, focusing specifically on their performance in recovering low-abundance, fragmented ctDNA from complex biological matrices like blood plasma.
Technical Performance Comparison
Yield and Recovery Efficiency
Recovery efficiency represents perhaps the most critical parameter for ctDNA extraction, directly impacting detection sensitivity in downstream applications. Multiple comparative studies have consistently demonstrated that spin column-based methods generally achieve higher DNA yields compared to magnetic bead-based approaches.
Table 1: Comparative Recovery Performance of Extraction Methods
| Performance Metric | Spin Columns | Magnetic Beads |
|---|---|---|
| Typical Recovery Range | 70-85% [57] | 94-96% (for general DNA) [57] |
| cfDNA Yield in Comparative Studies | Significantly higher (up to 4.3x in some comparisons) [12] [56] | Lower relative yields in direct comparisons [12] [56] |
| Performance in Low Abundance | Better suited for general ctDNA isolation with high recovery rates [17] [16] | More efficient at recovering smaller DNA fragments [17] [16] |
| Fragment Size Bias | Better recovery of variable-sized DNA, particularly >600 bp [17] [16] | Enhanced recovery of smaller fragments (<300 bp) [57] [17] |
A comprehensive 2022 study evaluating six commercial cfDNA extraction kits found that spin column-based methods, particularly the QIAamp Circulating Nucleic Acid Kit, demonstrated significantly greater recovery compared to magnetic bead-based kits, with yield differences of up to 4.3-fold between the best-performing spin column and magnetic bead kits [12]. Similarly, a 2018 study specifically evaluating cfDNA purification kits concluded that "spin column-based kits outperform magnetic bead-based commercial cfDNA kits," noting that the Qiagen spin column-based cfDNA kit remains the "gold standard" for consistent performance across evaluation assays [56].
The superior yield performance of spin columns for ctDNA extraction is particularly notable given that magnetic beads often demonstrate higher recovery rates (94-96%) for general DNA purification applications like PCR cleanup [57]. This apparent contradiction highlights the unique challenges posed by ctDNA's specific physical characteristics and its presence in complex biological matrices.
Fragment Size Selection Bias
The size profile of recovered DNA differs markedly between these technologies, with important implications for ctDNA analysis since tumor-derived fragments often exhibit distinct size distributions compared to background wild-type cfDNA.
Spin columns demonstrate a bias toward recovering longer DNA fragments, typically showing better performance for fragments exceeding 600 bp [17] [16]. This characteristic can be advantageous for applications requiring longer DNA fragments but may potentially under-recover the characteristic ~167 bp mononucleosomal ctDNA fraction.
In contrast, magnetic bead-based systems exhibit superior recovery of shorter DNA fragments, with multiple studies confirming their particular efficiency for fragments below 300 bp [57] [17]. This size profile aligns favorably with the natural size distribution of ctDNA, potentially enhancing the detection of tumor-derived fragments. The magnetic bead technology enables customizable size selection through adjustable bead-to-sample ratios, allowing researchers to preferentially target specific fragment size ranges [57].
Table 2: Fragment Size Selection Capabilities
| Technology | Optimal Fragment Size Range | Size Selection Flexibility | Mono-nucleosomal Recovery Efficiency |
|---|---|---|---|
| Spin Columns | 100 bp - 10 kb [57]; better for >600 bp [17] [16] | Limited, fixed by membrane porosity | Standard |
| Magnetic Beads | 100 bp - 50 kb [57]; enhanced for <300 bp [57] [17] | High, tunable via bead:sample ratio | Enhanced |
Throughput and Automation Capabilities
Workflow efficiency and scalability represent another area of significant differentiation between these technologies, particularly relevant for clinical laboratories processing large sample volumes.
Table 3: Workflow and Throughput Comparison
| Parameter | Spin Columns | Magnetic Beads |
|---|---|---|
| Throughput Capacity | Low to medium [58] | High [57] [58] |
| Automation Compatibility | Not automation-compatible [57] | Fully automation-compatible [57] |
| Hands-on Time | Manual, centrifugation required [58] | Minimal, no centrifugation [58] |
| Typical Processing Time | 20-30 minutes [57] | <15 minutes [57] |
| Batch Processing Capability | Limited, single-tube format [57] | Ideal for batch processing, 96-well formats [57] |
Magnetic bead technology offers distinct advantages in high-throughput settings, with compatibility for 96-well and 384-well plate formats and direct integration with automated liquid handling systems such as the Thermo Fisher KingFisher Flex, Hamilton Microlab STAR, and Beckman Coulter Biomek i-Series [57]. This automation capability significantly reduces hands-on time and minimizes technical variability across samples [57] [55].
Spin columns remain constrained by their fundamental design, requiring sequential centrifugation steps that limit processing throughput and present challenges for batch processing. While adequate for low-to-medium throughput research applications, this limitation becomes significant in clinical settings requiring high sample volumes [57] [58].
Cost and Environmental Considerations
Economic factors increasingly influence technology selection in resource-conscious laboratory environments.
Magnetic bead-based methods demonstrate substantially lower cost per sample, approximately $0.90 per reaction compared to $1.75 for spin columns, translating to nearly 50% cost savings when processing thousands of samples [57]. This economic advantage stems from reduced plastic consumption through 96-well plate formats and bulk reagent packaging [57].
The environmental impact also differs considerably, with magnetic bead systems generating less plastic waste due to compatibility with high-density plate formats and reduced reagent volumes [57]. Spin column workflows typically involve single-use plastic columns that contribute significantly to plastic waste streams.
Experimental Protocols and Methodologies
Standardized Workflow for Magnetic Bead-Based Extraction
Recent methodological advances have established robust, validated protocols for magnetic bead-based ctDNA extraction. A 2025 study by Dartmouth Hitchcock Medical Center detailed a comprehensive analytical validation of a magnetic bead-based, high-throughput cfDNA extraction system, demonstrating high recovery rates, consistent fragment size distribution, and minimal genomic DNA contamination [55].
Key Protocol Steps:
- Sample Preparation: Plasma samples obtained through double centrifugation (1600× g for 10 min, followed by 16,000× g for 10 min) to remove cellular debris [12]
- Binding: Mix sample with magnetic beads in optimized binding buffer, typically at ratios between 0.6x-1.8x depending on desired fragment size selection [57]
- Incubation: Allow DNA-bead binding for 5 minutes at room temperature [57]
- Separation: Transfer to magnetic stand for bead immobilization (~2 minutes) [57]
- Washing: Remove supernatant and wash twice with 80% ethanol [57]
- Elution: Resuspend beads in 20-50 µL nuclease-free water or TE buffer [57]
This protocol consistently recovered cfDNA with predominant mononucleosomal (~150 bp) and dinucleosomal (~340 bp) fragments, with minimal genomic DNA contamination, making it suitable for sensitive downstream applications like next-generation sequencing [55].
Spin Column Extraction Methodology
The established spin column protocol remains widely used in clinical and research settings:
Key Protocol Steps:
- Sample Preparation: Plasma obtained through identical double-centrifugation protocol as above [12]
- Binding: Apply sample to silica membrane column in presence of high-salt binding buffer [9]
- Centrifugation: Spin column to facilitate DNA binding to silica membrane [9]
- Washing: Perform multiple wash steps with ethanol-based buffers to remove contaminants [9]
- Elution: Elute DNA in low-salt buffer or nuclease-free water [9]
The QIAamp Circulating Nucleic Acid Kit protocol has been specifically optimized for cfDNA recovery and represents the most consistently performing spin column-based method across multiple independent evaluations [12] [56].
Figure 1: Comparative Workflow Diagram for ctDNA Extraction Methods. The magnetic bead pathway supports high-throughput automation, while the spin column method relies on manual centrifugation steps.
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagent Solutions for ctDNA Extraction Research
| Reagent/Kit | Type | Primary Function | Application Notes |
|---|---|---|---|
| QIAamp Circulating Nucleic Acid Kit [12] [56] | Spin Column | High-yield cfDNA isolation | Highest consistency in recovery; considered gold standard |
| HighPrep PCR Beads [57] | Magnetic Beads | DNA cleanup & size selection | Tunable size selection via bead ratio (0.6x-1.8x) |
| MagMAX Cell-Free DNA Isolation Kit [12] [55] | Magnetic Beads | Automated cfDNA extraction | Compatible with high-throughput systems |
| nRichDx cfDNA Reference Standard [55] | Quality Control | Extraction efficiency validation | Contains KRAS p.G12V mutation for spike-recovery studies |
| Seraseq ctDNA Reference Material [55] | Quality Control | Variant detection validation | Multiple VAF levels (0.1%-5%) with 25 clinically relevant variants |
| AcroMetrix ctDNA Plasma Control [55] | Quality Control | Extraction performance | Multiple VAF levels (0%, 0.1%, 0.5%, 1%) in human plasma matrix |
Method Selection Guidelines
Application-Specific Recommendations
Choosing between magnetic bead and spin column technologies requires careful consideration of research objectives, sample characteristics, and infrastructure capabilities.
Select Spin Columns When:
- Maximum Recovery Yield is the primary priority, particularly for low-concentration samples [12] [56]
- Processing Volume is low to medium, with limited need for high-throughput processing [58]
- Laboratory Infrastructure is constrained, with limited access to specialized magnetic separation equipment [9]
- Research Budget permits higher per-sample costs for potentially superior recovery [57]
Select Magnetic Beads When:
- High-Throughput Processing is required for large sample batches [57] [58]
- Workflow Automation is desirable to minimize hands-on time and reduce technical variability [57] [55]
- Small Fragment Recovery is prioritized, particularly for characteristic mononucleosomal ctDNA [17]
- Cost Efficiency is important, with lower per-sample processing costs [57]
- Size Selection Flexibility is needed for specific fragment targeting [57]
Emerging Technologies and Future Directions
The field of ctDNA extraction continues to evolve with several promising technological developments. Microfluidic fluidized bed systems show potential for enhancing capture efficiency of specific ctDNA sequences through improved bead homogeneity and specialized vibration systems [59]. Magnetic ionic liquid (MIL)-based dispersive liquid-liquid microextraction has demonstrated superior enrichment factors compared to conventional silica-based methods [17] [16]. Additionally, magnetic nanowire networks with elongated morphologies and high saturation magnetization are emerging as efficient platforms for cfDNA capture while minimizing loss and degradation [17] [16].
These advanced approaches, while not yet widely adopted in clinical practice, represent the ongoing innovation in ctDNA extraction technology aimed at addressing the fundamental challenges of low abundance and high fragmentation.
The comparative analysis of magnetic bead versus spin column technologies for ctDNA extraction reveals a nuanced performance landscape where neither approach demonstrates universal superiority across all parameters. Spin column methods currently maintain advantages in absolute recovery yield and consistency, making them particularly valuable for applications where detection sensitivity is paramount and sample volumes are manageable. Magnetic bead systems excel in throughput, automation compatibility, and cost efficiency, offering compelling benefits for large-scale studies and clinical implementations requiring high-volume processing.
The decision between these technologies ultimately depends on specific research priorities, with the understanding that methodological choices at the extraction stage fundamentally influence all subsequent analytical processes. As ctDNA analysis continues to transition toward routine clinical implementation, standardization of extraction methodologies will become increasingly important for ensuring reproducibility and comparability across studies and clinical laboratories.
Consistency and Reproducibility Across Different Sample Batches
Circulating tumor DNA (ctDNA), a subset of cell-free DNA (cfDNA) shed into the bloodstream by tumors, has emerged as a powerful biomarker for liquid biopsy applications in oncology [17] [60]. Its analysis enables non-invasive cancer detection, treatment monitoring, minimal residual disease assessment, and therapy selection [61]. However, the vanishingly low concentration of ctDNA in blood—often less than 0.1% of total cfDNA in early-stage cancer—presents significant analytical challenges [37]. The pre-analytical phase, particularly DNA extraction, profoundly impacts downstream analysis quality, making method selection crucial for obtaining reliable, reproducible results across sample batches [17] [23].
The two dominant technologies for nucleic acid extraction are spin columns (silica-membrane based) and magnetic beads (solid-phase reversible immobilization based). This guide objectively compares their performance for ctDNA isolation, focusing specifically on consistency and reproducibility across different sample batches—critical parameters for both clinical diagnostics and research applications where samples are processed over time or across multiple sites.
Spin Column Technology
Spin column-based extraction utilizes a silica membrane housed in a column that selectively binds DNA in the presence of chaotropic salts [9]. During centrifugation, the sample passes through this membrane under high-salt conditions, facilitating DNA binding to the silica surface [40]. Contaminants are removed through wash steps, and purified DNA is eluted in a low-salt buffer or water [62]. This method effectively recovers DNA fragments across a broad size range but may exhibit bias against shorter fragments characteristic of ctDNA [61].
Magnetic Bead Technology
Magnetic bead-based extraction employs paramagnetic particles coated with a silica surface or other DNA-binding chemistries [19]. When added to the sample, DNA binds to the beads' surface under appropriate buffer conditions (typically containing PEG and salt) [62]. A magnetic field immobilizes the beads while contaminants are removed through washing steps. Pure DNA is then eluted from the beads [40]. This principle, known as Solid Phase Reversible Immobilization (SPRI), allows flexible size selection by adjusting the bead-to-sample ratio [62].
Table: Fundamental Operational Differences Between Spin Column and Magnetic Bead Methods
| Parameter | Spin Columns | Magnetic Beads |
|---|---|---|
| Binding Principle | Silica membrane DNA binding with chaotropic salts [9] | Silica-coated magnetic particle DNA binding with PEG/salt [62] |
| Separation Mechanism | Centrifugation [40] | Magnetic field [40] |
| Primary Equipment | Microcentrifuge [40] | Magnetic rack or automated system [40] |
| Size Selection Capability | Limited, based on membrane pore size [62] | Flexible, via bead-to-sample ratio adjustment [62] |
| Typical Format | Single columns (low-throughput) [62] | 96-well plates (high-throughput) [62] |
Diagram: Comparative Workflows for Spin Column vs. Magnetic Bead Extraction
Performance Comparison: Quantitative Data Analysis
Recovery Yield and Efficiency
Recovery yield, particularly for low-abundance ctDNA, is perhaps the most critical performance metric. Studies consistently show magnetic bead-based systems offer superior recovery rates for fragmented DNA. Research comparing extraction methods reported that magnetic bead protocols achieved 94-96% recovery compared to 70-85% for spin columns [62]. This enhanced recovery is particularly crucial for ctDNA applications where target molecules are scarce.
A 2025 study directly comparing extraction efficiency from blood samples found that while overall efficiency of three different kits (one column-based, two magnetic bead-based) was similar, the column-based kit (Kit B) demonstrated superior performance in low-concentration samples, with average DNA yields 4.24-fold higher than one magnetic bead-based kit (Kit D) and 1.18-fold higher than another (Kit C) [63]. This highlights that specific kit formulation significantly impacts performance, and generalizations between technologies should be made cautiously.
Fragment Size Bias and Selectivity
ctDNA exhibits a characteristic fragmentation profile with a peak around 166 bp, but contains shorter tumor-derived fragments (<150 bp) that are diagnostically valuable [61]. Magnetic bead technology offers a distinct advantage through customizable size selection by adjusting the bead-to-sample ratio [62].
Table: Size Selection Capabilities of Magnetic Beads via Bead-to-Sample Ratio Adjustment
| Bead-to-Sample Ratio | DNA Fragment Size Retained |
|---|---|
| 0.6x | >500 bp |
| 0.8x | >300 bp |
| 1.0x | >100 bp |
| 1.8x | >50 bp [62] |
This tunable selectivity enables researchers to enrich for ctDNA fragments while excluding larger genomic DNA contamination, a crucial consideration for assay sensitivity. Spin columns generally lack this precise size selection capability and may lose shorter fragments during washing steps [61].
Reproducibility and Batch-to-Batch Consistency
Reproducibility across different sample batches depends on both the consistency of the extraction chemistry and the robustness of the protocol against technical variation. Magnetic bead systems demonstrate excellent reproducibility, with recent studies reporting no significant day-to-day variability when standardized protocols are followed [23]. The QIAamp Circulating Nucleic Acid Kit (spin column), for example, showed high reproducibility in cfDNA extraction with no day-to-day variability observed [23].
For high-throughput applications, magnetic bead systems integrated with automated liquid handlers provide superior consistency by minimizing manual handling variations [62] [40]. One study noted that automated extraction systems demonstrate "a lower margin of error and superior reproducibility compared to their manual counterparts" [19]. This makes magnetic bead systems particularly advantageous for multi-center studies where batch effects must be minimized.
Experimental Protocols and Methodologies
Standardized Magnetic Bead Protocol for ctDNA
The following optimized protocol for magnetic bead-based ctDNA extraction has been validated for consistency across sample batches [62] [19]:
Sample Preparation: Begin with 1-5 mL of double-centrifuged plasma (initial centrifugation at 1,600 × g for 10 minutes, followed by 16,000 × g for 10 minutes at 4°C) to remove cells and debris [17].
Lysis and Digestion: Combine plasma with 200 μL lysis buffer (containing guanidine HCl, Tris, and Triton X-100) and 30 μL proteinase K (20 mg/mL). Incubate at 60°C for 15 minutes with shaking at 300 rpm to digest proteins and release nucleic acids [19].
DNA Binding: Add 1.8x volume of magnetic beads (specifically optimized for cfDNA recovery) and 1 mL binding buffer (containing guanidine salt, sodium sulfate, and 2-propanol). Incubate at room temperature for 10 minutes with mixing at 400 rpm to facilitate DNA binding [62] [19].
Magnetic Separation: Transfer tubes to a magnetic rack until solution clears (approximately 2 minutes). Carefully discard supernatant without disturbing the bead pellet [62].
Washing: Wash beads twice with 500 μL wash buffer I (guanidine salt and ethanol), followed by wash buffer II (ethanol only). Ensure complete removal of wash buffers between steps [19].
Elution: Air-dry beads for 3-5 minutes (avoid over-drying). Elute DNA in 20-50 μL elution buffer (Tris-HCl) by shaking at 800 rpm for 5 minutes at room temperature. Collect eluate using magnetic separation [62].
Standardized Spin Column Protocol for ctDNA
The following represents an optimized spin column protocol for ctDNA extraction [63] [23]:
Sample Preparation: Process 1-5 mL of double-centrifuged plasma as described above. For spin columns, immediate processing (within 2-6 hours) is recommended if using EDTA tubes, though specialized collection tubes (e.g., Streck, PAXgene) allow longer processing windows [17].
Lysis and Binding: Mix plasma with equal volume of binding buffer (containing chaotropic salts) and incubate at room temperature for 5 minutes. Load mixture onto spin column [23].
Centrifugation and Washing: Centrifuge at 14,000 × g for 1 minute to bind DNA to silica membrane. Wash twice with wash buffers (typically ethanol-based) with centrifugation between steps [63].
Elution: Centrifuge columns briefly to remove residual ethanol. Elute DNA in 20-50 μL elution buffer or nuclease-free water by incubating for 3-5 minutes followed by centrifugation at 14,000 × g for 2 minutes [63].
Operational Considerations for Batch Processing
Throughput and Automation Capabilities
Throughput requirements significantly influence method selection for batch processing. Magnetic bead systems excel in high-throughput environments, with compatibility for 96-well and 384-well plate formats and full automation on robotic liquid handlers [62] [40]. This enables simultaneous processing of hundreds of samples with minimal hands-on time, enhancing batch consistency.
Spin columns remain practical for low-to-medium throughput applications but face scalability limitations due to sequential tube handling and centrifugation requirements [62]. One evaluation noted that spin columns are "not automation-compatible" and become "labor-intensive for batch processing" [62].
Table: Throughput and Operational Comparison
| Parameter | Spin Columns | Magnetic Beads |
|---|---|---|
| Maximum Throughput | Low to medium (manual processing) [40] | High (96/384-well, automation compatible) [62] |
| Hands-on Time | High for large batches [62] | Minimal, especially when automated [40] |
| Automation Compatibility | Limited [62] | Excellent (Thermo Fisher KingFisher, Hamilton STAR, etc.) [62] |
| Batch Processing Suitability | Poor for large batches [62] | Excellent for large batches [62] |
Cost Analysis and Resource Requirements
Comprehensive cost assessment must include both consumable expenses and labor costs. While per-reaction costs for spin columns are generally higher ($1.75 per sample versus $0.90 for magnetic beads) [62], magnetic bead methods require significant upfront investment in magnetic separators or automated systems [9].
For laboratories processing 1,000 samples, magnetic bead systems offer nearly 50% cost savings on consumables alone ($900 versus $1,750) [62]. When factoring in reduced hands-on time and improved throughput, the total cost advantage of magnetic beads becomes more substantial for high-volume applications.
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful ctDNA extraction requires careful selection of reagents and materials optimized for recovering low-abundance, fragmented DNA. The following solutions represent key components for reliable, reproducible extraction across sample batches:
Table: Essential Research Reagent Solutions for ctDNA Extraction
| Reagent/Material | Function | Technology Compatibility |
|---|---|---|
| Specialized Blood Collection Tubes (e.g., Streck, PAXgene) | Preserve blood cell integrity, prevent gDNA contamination during transport/storage [17] | Both methods |
| Proteinase K | Digest nucleoprotein complexes, release DNA [19] | Both methods |
| Optimized Lysis Buffer | Disrupt vesicles, denature proteins, release cfDNA [19] | Both methods |
| Size-Selective Magnetic Beads | Selective recovery of ctDNA fragments by size exclusion [62] | Magnetic beads |
| Carrier RNA | Improve recovery of low-abundance fragments [61] | Both methods (especially magnetic beads) |
| Inhibitor Removal Resins | Remove PCR inhibitors (hemoglobin, heparin, etc.) [61] | Both methods |
| Nuclease-Free Elution Buffers | Stabilize eluted DNA, prevent degradation [62] | Both methods |
The choice between magnetic bead and spin column technologies for ctDNA extraction involves careful consideration of application requirements, sample volume, and throughput needs. For studies prioritizing consistency and reproducibility across different sample batches—particularly in high-throughput settings or multi-center trials—magnetic bead systems offer distinct advantages in recovery efficiency, automation compatibility, and batch-to-batch consistency [62] [23] [19].
Spin columns remain a viable option for lower-throughput applications where upfront equipment costs are a primary concern, though they may exhibit greater variability in ctDNA recovery, particularly for shorter fragments [63] [61]. Regardless of the chosen method, standardization of pre-analytical conditions—including blood collection, processing protocols, and extraction parameters—is essential for minimizing batch effects and ensuring reproducible ctDNA analysis across different sample batches [17] [60].
The analysis of circulating tumor DNA (ctDNA) has emerged as a cornerstone of liquid biopsy in oncology, enabling non-invasive cancer detection, monitoring of treatment efficacy, and assessment of minimal residual disease [16]. The reliability of this analysis, however, is profoundly influenced by the choice of DNA extraction method, which directly impacts the yield, purity, and fragment size distribution of the isolated nucleic acids. Efficient extraction of ctDNA with high yield and purity is critical to ensuring the sensitivity and reliability of downstream analyses [16]. This guide provides a objective comparison of the two predominant ctDNA extraction techniques—spin column-based and magnetic bead-based isolation—focusing on their performance across three major downstream applications: Next-Generation Sequencing (NGS), quantitative PCR (qPCR), and digital PCR (dPCR).
Technical Comparison of Spin Column vs. Magnetic Bead Extraction Methods
The principles, advantages, and limitations of spin column and magnetic bead-based DNA extraction methods differ significantly, influencing their suitability for specific applications and sample types [9].
Principles and Workflows
- Spin Column-Based Kits operate on the principle of selective binding of DNA to a silica membrane under high-salt conditions. The process involves sample lysis, binding to the silica membrane in a column, washing away contaminants, and eluting the purified DNA [9].
- Magnetic Bead-Based Kits utilize magnetic particles coated with a DNA-binding surface (often silica). After sample lysis, DNA binds to the beads, and a magnetic field is applied to separate the DNA-bound beads from the solution. Subsequent wash steps remove impurities, and the purified DNA is then eluted [9] [16].
The table below summarizes the core characteristics of each method.
Table 1: Core Characteristics of DNA Extraction Methods
| Feature | Spin Column | Magnetic Beads |
|---|---|---|
| Basic Principle | DNA binding to silica membrane [9] | DNA binding to silica-coated magnetic beads [9] [16] |
| Scalability | Ideal for low to moderate throughput [9] | Highly scalable and easy to automate for high-throughput [9] [16] |
| DNA Recovery | Limited binding capacity; may not be ideal for very low DNA concentrations [9] | Higher binding capacity; better recovery from low-yield samples [9] |
| Handling of Fragments | Better suited for recovering variable-sized DNA, particularly high molecular weight fragments (>600 bp) [16] | Particularly efficient at recovering smaller DNA fragments (e.g., ctDNA) [16] |
| Ease of Use & Cost | Simple, fast, and lower equipment cost [9] | Can be more complex manually; requires a magnetic separator; ideal for automation [9] |
Performance in Downstream Applications
The choice between spin column and magnetic bead extraction directly influences the efficiency, sensitivity, and accuracy of downstream molecular assays.
Next-Generation Sequencing (NGS)
NGS requires high-quality DNA with minimal contamination and optimized fragment sizes for successful library preparation [64]. For targeted NGS approaches, such as hybridization capture or amplicon sequencing, the quality of the starting material is paramount [65].
- Magnetic Bead Performance: The superior recovery of smaller DNA fragments makes magnetic beads the preferred choice for ctDNA-based NGS applications [16]. The ability to automate the process ensures consistency across large sample sets, which is critical for generating reliable sequencing data [9]. This method is particularly advantageous for detecting low-frequency variants.
- Spin Column Performance: While spin columns are widely used and can yield DNA suitable for NGS, they may be less effective at enriching the short-fragment ctDNA fraction. They are a robust choice for general purpose NGS where the starting material is not severely limited [9].
Quantitative PCR (qPCR)
qPCR is a highly sensitive technique for quantifying specific nucleic acid sequences, often used for gene expression analysis or quantifying viral loads [66] [67]. Its reliability depends on both the quantity and purity of the isolated DNA, as inhibitors co-purified during extraction can dramatically affect amplification efficiency [67].
- Spin Column Performance: Spin columns are a classic and established method for DNA extraction for qPCR. They provide high-purity DNA that is generally free of PCR inhibitors. However, if the ctDNA yield is low, the limited recovery of spin columns could impact the qPCR quantification [9].
- Magnetic Bead Performance: Magnetic bead-based extraction is highly effective for qPCR, offering good recovery and purity. Its high tolerance to inhibitors present in some sample matrices makes it a robust choice for challenging clinical samples [66].
Digital PCR (dPCR)
dPCR is a powerful method for the absolute quantification of nucleic acids without the need for a standard curve. It is especially valuable for detecting rare mutations and copy number variations, as it partitions a sample into thousands of individual reactions [66] [68]. The precision of dPCR is heavily dependent on the integrity and precise quantification of the input DNA [68].
- Magnetic Bead Performance: Magnetic beads are exceptionally well-suited for dPCR. Their high precision in recovery and efficient capture of small ctDNA fragments ensures that the input material for dPCR is a accurate representation of the original sample. This leads to tighter confidence intervals and better resolution for detecting small differences in copy number [66] [16].
- Spin Column Performance: Spin columns can be used for dPCR, but the potential for lower recovery of the target ctDNA fraction might affect the precision of the assay, particularly when analyzing low-abundance targets [9].
Table 2: Suitability for Downstream Applications
| Application | Key Requirement | Spin Column Suitability | Magnetic Bead Suitability |
|---|---|---|---|
| NGS | High-quality, fragment-size appropriate DNA; high sensitivity for low-frequency variants [64] [16] [65] | Good for general use, but may miss shorter ctDNA fragments [9] [16] | Excellent, due to efficient small fragment recovery and automation [9] [16] |
| qPCR | Pure DNA free of inhibitors; high yield for accurate quantification [67] | Good, provides high-purity DNA [9] | Excellent, robust performance with good recovery and inhibitor tolerance [66] |
| dPCR | Absolute quantification; high precision and sensitivity for rare targets [66] [68] | Good, but potential for lower precision with low-yield samples [9] | Excellent, offers high recovery and precision for absolute quantification [66] [16] |
Essential Workflow and Protocol Considerations for ctDNA Analysis
A reliable ctDNA analysis workflow extends beyond the choice of extraction method. Pre-analytical variables are critical in determining ctDNA integrity, purity, and yield [16] [5].
Pre-Analytical Steps: Blood Collection and Plasma Preparation
- Blood Collection Tubes: The choice of tube is crucial. EDTA tubes are standard but require processing within hours to prevent leukocyte DNA contamination. Specialized blood collection tubes (BCTs) from manufacturers like Streck or PAXgene contain preservatives that stabilize nucleated blood cells, preventing lysis and allowing plasma isolation to be delayed for up to several days without significantly affecting cfDNA quality [16] [5].
- Centrifugation Protocol: A two-step centrifugation protocol is widely recommended. An initial low-speed spin (800–1,900 ×g for 10 min) pellets blood cells, followed by a high-speed spin (14,000–16,000 ×g for 10 min) of the supernatant to remove any remaining cellular debris, thereby improving the purity of the plasma and subsequent cfDNA extract [16].
Experimental Protocol: Automated ctDNA Extraction using Magnetic Beads
The following protocol is adapted from a 2025 study evaluating automated cfDNA extraction and provides a reliable methodological framework [5].
- Sample Input: Use 1-4 mL of plasma obtained from double-centrifuged blood.
- Automated Platform: Perform extraction on a system such as the QIAsymphony SP using a magnetic bead-based chemistry kit.
- Lysis and Binding: Mix the plasma with a lysis/binding buffer to release cfDNA and allow it to bind to the magnetic beads.
- Magnetic Separation and Washes: Apply a magnetic field to capture the beads and perform multiple wash steps with appropriate buffers to remove proteins, salts, and other contaminants.
- Elution: Elute the purified cfDNA in a small volume (e.g., 50-100 µL) of low-salt buffer or nuclease-free water.
- Quality Assessment: Quantify the extracted cfDNA using a fluorometric method (e.g., Qubit) and assess fragment size distribution via parallel capillary electrophoresis (e.g., TapeStation, Bioanalyzer) or by using qPCR assays targeting short (~60-80 bp) versus long (>150 bp) amplifications [5].
Workflow Visualization
The following diagram illustrates the logical workflow for ctDNA analysis, from sample collection to data interpretation for clinical decision-making.
The Scientist's Toolkit: Key Reagent Solutions
Successful execution of a ctDNA analysis workflow relies on a suite of reliable reagents and tools. The table below lists essential solutions and their functions.
Table 3: Essential Research Reagent Solutions for ctDNA Analysis
| Research Reagent Solution | Function in Workflow |
|---|---|
| Stabilizing Blood Collection Tubes (e.g., Streck, PAXgene) [16] [5] | Preserves blood sample by preventing leukocyte lysis and genomic DNA release, enabling delayed plasma processing. |
| Magnetic Bead-Based Extraction Kits (e.g., for QIAsymphony SP) [16] [5] | Enable automated, high-throughput purification of cfDNA with high recovery of short fragments. |
| Spin Column-Based Extraction Kits (e.g., QIAamp) [9] [16] | Provide a simple, manual method for DNA purification using silica-membrane technology. |
| Fluorometric Quantitation Kits (e.g., Qubit dsDNA HS Assay) [5] | Accurately quantify low concentrations of double-stranded DNA in solution. |
| qPCR/dPCR Master Mixes & Assays (e.g., TaqMan) [66] [67] [5] | Provide the enzymes, buffers, and sequence-specific primers/probes for sensitive and specific target amplification and quantification. |
| Parallel Capillary Electrophoresis Kits (e.g., Agilent TapeStation) [64] [5] | Analyze the size distribution and integrity of extracted nucleic acids, critical for assessing cfDNA quality and detecting contamination. |
The selection between spin column and magnetic bead-based extraction methods is a pivotal decision in ctDNA analysis. Magnetic bead-based methods demonstrate superior performance for NGS and dPCR applications, primarily due to their higher recovery of the short-fragment ctDNA, better tolerance to inhibitors, and suitability for automation, which enhances precision and throughput. Spin column-based methods remain a robust, cost-effective, and simple alternative, particularly for standard qPCR applications or when processing moderate sample numbers where high fragment recovery is not the primary concern. The optimal choice must be guided by the specific requirements of the downstream application, the nature of the sample, and the operational needs of the laboratory. As ctDNA analysis continues to integrate into clinical practice, adherence to standardized, optimized pre-analytical workflows will be fundamental to generating reliable and actionable data.
Conclusion
The choice between spin columns and magnetic beads is not one of absolute superiority but of strategic alignment with project goals. Spin columns offer simplicity and reliability for low-to-mid throughput workflows, while magnetic beads excel in high-throughput, automated environments and for maximizing recovery from low-concentration samples. The future of ctDNA extraction lies in the continued refinement of these technologies, the development of novel materials like magnetic ionic liquids, and the creation of integrated microfluidic systems that automate pre-analytical steps. As liquid biopsy moves further into routine clinical practice, standardized, efficient, and highly sensitive extraction methods will be the cornerstone of reliable cancer diagnosis, monitoring, and treatment personalization.
References
- 1. Clinical Circulating Tumor DNA Testing for Precision Oncology [https://pmc.ncbi.nlm.nih.gov/articles/PMC10101787/]
- 2. Circulating tumor nucleic acids: biology, release mechanisms ... [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-022-01710-w]
- 3. Comparison of Circulating Cell-Free DNA Extraction ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7281769/]
- 4. Analytical Comparison of Methods for Extraction of Short ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6854475/]
- 5. Evaluation of automatic cell free DNA extraction metrics ... [https://www.nature.com/articles/s41598-025-03508-4]
- 6. Reliability of liquid biopsy analysis: an inter-laboratory ... [https://www.sciencedirect.com/science/article/pii/S0023683722003476]
- 7. Life and death of circulating cell-free DNA - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6606043/]
- 8. Circulating tumour DNA for cancer patients [https://www.genomicseducation.hee.nhs.uk/genotes/knowledge-hub/circulating-tumour-dna-for-cancer-patients/]
- 9. DNA Extraction Kits Compared: Spin Columns vs. Magnetic ... [https://synapse.patsnap.com/article/dna-extraction-kits-compared-spin-columns-vs-magnetic-beads]
- 10. Separation of ctDNA by superparamagnetic bead particles ... [https://www.sciencedirect.com/science/article/pii/S2090123221000369]
- 11. Preanalytical variables and analytes in liquid biopsy ... [https://pubmed.ncbi.nlm.nih.gov/40884415/]
- 12. Isolation and Quantification of Plasma Cell-Free DNA ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9601152/]
- 13. Evaluation of pre-analytical factors affecting plasma DNA ... [https://www.nature.com/articles/s41598-018-25810-0]
- 14. Techniques of using circulating tumor DNA as a liquid ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6197739/]
- 15. DNA Adsorption to and Elution from Silica Surfaces [https://pmc.ncbi.nlm.nih.gov/articles/PMC4097040/]
- 16. Circulating tumor DNA laboratory processes and clinical ... [https://www.frontiersin.org/journals/oncology/articles/10.3389/fonc.2025.1520733/full]
- 17. Circulating tumor DNA laboratory processes and clinical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12119280/]
- 18. Comparison of two silica-based extraction methods for ... [https://www.sciencedirect.com/science/article/abs/pii/S1344622316300608]
- 19. Design, Development, and Clinical Validation of a Novel ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12346676/]
- 20. Magnetic Beads Market Size and YoY Growth Rate, 2025- ... [https://www.coherentmarketinsights.com/industry-reports/magnetic-beads-market]
- 21. What do we need to obtain high quality circulating tumor ... [https://www.sciencedirect.com/science/article/pii/S0009898121001893]
- 22. Standardized Workflow and Analytical Validation of Cell ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12293894/]
- 23. Extraction of Cell-Free DNA: Evaluation of Efficiency, ... [https://www.sciencedirect.com/science/article/pii/S152515782400014X]
- 24. A rapid and high-yield method for nucleic acid extraction [https://www.nature.com/articles/s41598-025-95226-0]
- 25. Precision cell-free DNA extraction for liquid biopsy by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7039987/]
- 26. SPRI bead or column DNA purification: Which method is ... [https://www.epicypher.com/resources/blog/spri-bead-or-column-dna-purification-which-method-is-best-for-cutrun/]
- 27. Separation of ctDNA by superparamagnetic bead particles ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8463959/]
- 28. Effects of Functionalized Silica-Coated Magnetic ... [https://www.msrjournal.com/article_24773.html]
- 29. The surface modification of the silica-coated magnetic ... [https://www.nature.com/articles/s41598-024-64839-2]
- 30. Rapid and Efficient Extraction of Cell-Free DNA Using ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9405790/]
- 31. How Spin Columns Work in Nucleic Acid Purification [https://ahn-bio.com/how-spin-columns-work-in-nucleic-acid-purification-a-step-by-step-guide/]
- 32. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOopqA0FiHccsJd-wjP5vZCWEKDK1nvNk17EIMiwXPZrW3aGhXll2]
- 33. PCR purification | Magnetic beads and spin columns | INTEGRA [https://www.integra-biosciences.com/united-states/en/stories/introduction-pcr-purification]
- 34. Magnetic Beads in Nucleic Acid Extraction [https://thearkdb.org/magnetic-beads-in-nucleic-acid-extraction-protocol-optimization-for-dna-and-rna-purity/]
- 35. Bio-On-Magnetic-Beads (BOMB): Open platform for high ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6343928/]
- 36. Magnetic Beads Protocol, How to use ... [https://www.nvigen.com/tips-working-magnetic-beads/]
- 37. Improvement of the sensitivity of circulating tumor DNA-based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12332530/]
- 38. High‐throughput isolation of circulating tumor DNA [https://pmc.ncbi.nlm.nih.gov/articles/PMC6360376/]
- 39. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOoo8w7ySctUHT3hLvL1v91V-XMM7TVdMkZ0Lz8powY7IaGYTAP14]
- 40. PCR Purification: Magnetic Beads vs Spin Columns [https://biotech-spain.com/en/articles/pcr-purification-magnetic-beads-vs-spin-columns-which-is-best-for-your-lab-/]
- 41. Automating Nucleic Acid Extraction [https://www.promega.com/resources/technologies/lab-automation-resource-center/automating-nucleic-acid-extraction/]
- 42. Maximizing Yield from Nucleic Acid Extraction/Purification [https://imcstips.com/how-to-maximize-yield-from-nucleic-acid-extraction-or-purification/]
- 43. Troubleshooting Low DNA Yield from Blood Samples [https://www.cambrianbioworks.com/blog/troubleshooting-low-dna-yield-from-blood-samples-a-guide-for-clinical-labs]
- 44. Molecular counting enables accurate and precise ... [https://www.nature.com/articles/s41598-025-90013-3]
- 45. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOoqYGuuoZPBQsvzHl-UtNBZfo8jooS2WMfASd888magdhqs3BuWx]
- 46. Design, Development, and Clinical Validation of a Novel ... [https://www.mdpi.com/2075-4418/15/15/1897]
- 47. Recommendations for Cell-Free DNA Assay Validations [https://www.sciencedirect.com/science/article/pii/S1525157823002192]
- 48. A novel method for liquid-phase extraction of cell-free DNA ... [https://www.nature.com/articles/s41598-021-98815-x]
- 49. Evaluation of silica spin‑column and magnetic bead ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11190640/]
- 50. Techniques of using circulating tumor DNA as a liquid ... [https://www.sciencedirect.com/science/article/pii/S2001037018301958]
- 51. The advantages and disadvantage of DNA extraction by ... [https://www.geneture.com/news/The-advantages-and-disadvantage-of-DNA-extraction-by-spin-column-method.html]
- 52. Advantages and Disadvantages of Magnetic Bead Method ... [https://en.opentrons.com.cn/news/cizhufayouquedian/]
- 53. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOooeXM8ML06whQKBomUq8NRDzsauID89Ub-K60VUAoGrmW8_iFWY]
- 54. Extraction and purification of short DNA fragments from ... [https://www.sciencedirect.com/science/article/pii/S1383586625014819]
- 55. Standardized Workflow and Analytical Validation of Cell ... [https://www.mdpi.com/2073-4409/14/14/1062]
- 56. Evaluation of commercial kits for purification of circulating ... [https://www.sciencedirect.com/science/article/abs/pii/S2210776218302503]
- 57. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOorxrCVuLJQS3qz0NSysILmjfF59XeIh2DL3-qG6CBc_jEBp3pkb]
- 58. PCR Purification: Magnetic Beads vs Spin Columns [https://www.canvaxbiotech.com/news/pcr-purification-magnetic-beads-vs-spin-columns-which-is-best-for-your-lab/?srsltid=AfmBOooBL9wtRbtgE7WSz8gSJsYQ86VmmKbSXHClbCYcG2rRHPTwPAgs]
- 59. High-throughput extraction on a dynamic solid phase for ... [https://www.nature.com/articles/s41378-023-00582-4]
- 60. 循环肿瘤DNA(ctDNA)的NGS检测临床实践专家共识(2022 ... [https://www.cn-healthcare.com/articlewm/20220628/content-1391189.html]
- 61. Comparative Overview of cfDNA and ctDNA: Biology, Utility ... [https://www.cd-genomics.com/resource-cfdna-ctdna-key-differences-applications-extraction-considerations.html]
- 62. SPRI Beads vs Spin Columns [https://www.magbiogenomics.com/latest/post/magnetic-bead-dna-cleanup-protocols-optimizing-pcr-cleanups-with-highprep-vs-spin-columns?srsltid=AfmBOopdoHYMpMNQE_Gh05RueYUTW9Unfwhy3fz-voFlaZfV1LX-ohkx]
- 63. 血液样本中DNA与RNA提取效率的研究 [https://xuebao.shsmu.edu.cn/article/2025/1674-8115/1674-8115-2025-45-4-476.shtml]
- 64. How-to: NGS Quality Control [https://frontlinegenomics.com/how-to-ngs-quality-control/]
- 65. Targeted next generation sequencing (NGS) | IDT [https://www.idtdna.com/pages/technology/next-generation-sequencing/dna-sequencing/targeted-sequencing]
- 66. dPCR vs qPCR vs end-point PCR [https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/bench-guide/pcr/digital-pcr/digital-pcr-vs-quantitative-pcr-vs-end-point-pcr]
- 67. qPCR and qRT-PCR analysis: Regulatory points to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7786041/]
- 68. Digital PCR for the characterization of reference materials [https://www.sciencedirect.com/science/article/pii/S0098299724000153]