Digital PCR in Oncology: Achieving Absolute Quantification for Precision Medicine and Liquid Biopsy Applications
Digital PCR (dPCR) represents a third-generation PCR technology that enables absolute, calibration-free quantification of nucleic acids, revolutionizing molecular analysis in oncology.
Digital PCR in Oncology: Achieving Absolute Quantification for Precision Medicine and Liquid Biopsy Applications
Abstract
Digital PCR (dPCR) represents a third-generation PCR technology that enables absolute, calibration-free quantification of nucleic acids, revolutionizing molecular analysis in oncology. This article explores the foundational principles of dPCR, its core methodology, and its transformative clinical applications, particularly in liquid biopsy for circulating tumor DNA (ctDNA) analysis and minimal residual disease (MRD) monitoring. We detail practical troubleshooting and optimization strategies and present robust validation data comparing dPCR performance against established techniques like quantitative PCR (qPCR) and next-generation sequencing (NGS). Aimed at researchers, scientists, and drug development professionals, this review synthesizes evidence demonstrating how dPCR's superior sensitivity, precision, and accuracy are advancing personalized cancer diagnostics, therapeutic monitoring, and early relapse detection.
The Principles of Digital PCR: From Single-Molecule Detection to Absolute Quantification in Cancer Genomics
Digital PCR (dPCR) represents the third generation of Polymerase Chain Reaction technology, following conventional PCR and real-time quantitative PCR (qPCR). This method is founded on the partitioning of a PCR mixture containing the sample into a large number of parallel reactions, so that each partition contains either zero, one, or a few nucleic acid targets according to a Poisson distribution [1]. Following PCR amplification, the fraction of positive partitions is measured via endpoint detection, enabling absolute quantification of the target concentration through Poisson statistics without the need for external calibration curves [1]. This calibration-free technology presents powerful advantages including high sensitivity, absolute quantification, high accuracy and reproducibility, and rapid turnaround time, making it particularly valuable for oncology research where precise nucleic acid quantification is critical [1].
The historical development of dPCR began with precursor work in 1989 using limiting dilution PCR to detect single copies of HIV provirus [1]. The foundations were formally established in 1992 when Morley and Sykes combined limiting dilution PCR with Poisson statistics to isolate, detect, and quantify single nucleic acid molecules [1]. The term "digital PCR" was coined in 1999 by Bert Vogelstein and colleagues, who developed a workflow involving limiting dilution distributed on 96-well plates combined with fluorescence readout to detect mutations of the RAS oncogene in patients with colorectal cancer [1].
Principle and Workflow of Digital PCR
Core Principle
The fundamental principle underlying dPCR is sample partitioning, which enables the transformation of a quantitative measurement problem into a simple binary counting exercise. By dividing the reaction into thousands to millions of nanoliter-scale partitions, the technique effectively dilutes the target molecules to a concentration where most partitions contain either zero or one molecule [1]. This partitioning occurs via random distribution, following Poisson statistics. After endpoint PCR amplification, each partition is analyzed as positive (fluorescence detected) or negative (no fluorescence), allowing precise calculation of the initial target concentration based on the proportion of positive partitions [1].
Experimental Workflow
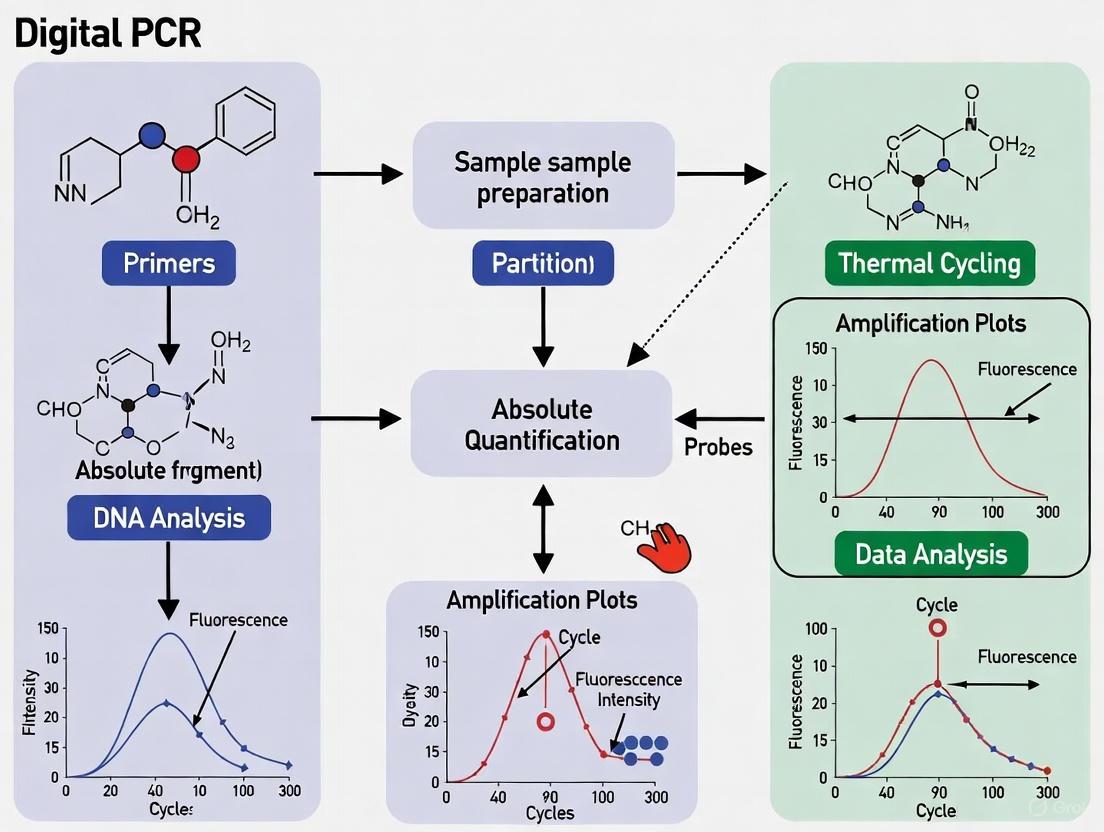
The dPCR workflow consists of four key steps: partitioning the PCR mixture, amplifying individual target-containing partitions, performing endpoint fluorescence analysis, and computing target concentration using Poisson statistics [1]. The following diagram illustrates this workflow and its application in oncology research:
Comparison with Other PCR Technologies
Table 1: Key Characteristics of PCR Technologies
| Parameter | Conventional PCR | Quantitative PCR (qPCR) | Digital PCR (dPCR) |
|---|---|---|---|
| Quantification Method | Semi-quantitative (gel electrophoresis) | Relative quantification (requires standard curve) | Absolute quantification (Poisson statistics) |
| Sensitivity | Moderate | High | Very high (can detect single molecules) |
| Precision | Low | Moderate | High |
| Dynamic Range | Limited | Wide | Wide |
| Tolerance to Inhibitors | Low | Moderate | High [2] |
| Multiplexing Capability | Limited | Moderate | High [3] |
| Primary Application | Target detection | Gene expression, quantification | Rare variant detection, absolute quantification [1] |
Applications in Oncology Research
Liquid Biopsy and Circulating Biomarkers
dPCR has revolutionized liquid biopsy applications in oncology by enabling sensitive detection and quantification of circulating tumor DNA (ctDNA) and other biomarkers. The technology's exceptional sensitivity allows researchers to detect rare genetic mutations within a background of wild-type genes, which is crucial for monitoring tumor heterogeneity and treatment response [1]. In metastatic melanoma, for example, duplex dPCR assays have been developed for simultaneous detection of miR-4488 and miR-579-3p in serum samples, creating a miRatio biomarker that predicts response to MAPK inhibitor therapy [3]. This approach demonstrated superior sensitivity compared to qRT-PCR, particularly for detecting low-abundance miRNAs, highlighting dPCR's utility in longitudinal monitoring of therapeutic response [3].
Rare Mutation Detection and Minimal Residual Disease
The ability to detect rare mutations with high precision makes dPCR particularly valuable for monitoring minimal residual disease (MRD) in hemato-oncology. dPCR is 100-times more sensitive than conventional methods for detecting rare mutations, with the ability to pool samples for even higher sensitivity [4]. Applications include reliable quantification of fusion transcripts like BCR-ABL and NPM1 mutations down to 0.001% frequency, enabling early detection of treatment resistance and disease recurrence [2]. The compartmentalization of the reaction renders dPCR less sensitive to PCR inhibitors, mismatched assay efficiencies, and inter-assay competition, providing more robust quantification of mutant allele frequencies than other molecular techniques [2].
Copy Number Variation Analysis
dPCR provides accurate and precise measurement of DNA copy number variations (CNVs), which is crucial for oncogene amplification studies and cancer genomics research. A 2025 study demonstrated that ddPCR shows 95% concordance with pulsed-field gel electrophoresis (considered a gold standard for CNV identification) for copy number typing of the DEFA1A3 gene, compared to only 60% concordance for qPCR [5]. This high accuracy at both low and high copy numbers, combined with its cost-effectiveness and throughput, makes dPCR an ideal methodology for CNV analysis in clinical cancer research [5].
Experimental Protocols
Protocol 1: Duplex dPCR for Circulating miRNA Ratio Quantification in Metastatic Melanoma
Purpose: Simultaneous detection of miR-4488 and miR-579-3p in serum samples from patients with BRAF-mutant metastatic melanoma to calculate miRatio as a predictive biomarker for MAPK inhibitor therapy response [3].
Materials and Reagents:
- Serum samples from patients and healthy donors
- miRNeasy Mini Kit (Qiagen, 217204) for RNA extraction
- TaqMan Advanced miRNA cDNA Synthesis Kit (ThermoFisher Scientific, A28007)
- TaqMan MicroRNA Assay probes for miR-4488 and miR-579-3p
- dPCR master mix compatible with chosen platform
- QIAcuity Nanoplate 26k (for QIAcuity system) or equivalent partitioning device
Procedure:
- RNA Extraction:
- Extract total RNA from 200 μL serum using miRNeasy Mini Kit according to manufacturer's instructions.
- Elute RNA in 20 μL nuclease-free water.
- Note: Serum RNA concentrations are often below detection thresholds; use fixed input volume of 2 μL for reverse transcription.
Reverse Transcription and Preamplification:
- Perform reverse transcription using TaqMan Advanced miRNA cDNA Synthesis Kit.
- Include the preamplification step to enrich target sequences prior to detection.
- Use universal reverse transcription for all miRNAs to ensure consistent cDNA quality.
dPCR Reaction Setup:
- Prepare duplex reaction mix containing:
- 5 μL 2× dPCR master mix
- 0.5 μL 20× TaqMan MicroRNA Assay probe for miR-4488 (FAM-labeled)
- 0.5 μL 20× TaqMan MicroRNA Assay probe for miR-579-3p (VIC-labeled)
- 1 μL cDNA template
- Nuclease-free water to 10 μL final volume
- Load 9 μL reaction mix into nanoplate wells.
- Prepare duplex reaction mix containing:
Partitioning and Amplification:
- Perform partitioning according to instrument specifications (26,000 partitions for QIAcuity Nanoplate 26k).
- Run thermal cycling with the following conditions:
- Enzyme activation: 95°C for 2 minutes
- 40 cycles of:
- Denaturation: 95°C for 15 seconds
- Annealing/Extension: 60°C for 60 seconds
Data Analysis:
- Image partitions and analyze fluorescence signals for both channels.
- Calculate absolute copies/μL for each miRNA using instrument software.
- Compute miRatio as: miR-4488 copies / miR-579-3p copies
- Perform ROC analysis to determine prognostic value of miRatio.
Technical Notes:
- The duplex assay maintains analytical performance comparable to singleplex reactions while reducing sample and reagent consumption [3].
- dPCR shows superior sensitivity to qRT-PCR, particularly for low-abundance miRNAs like miR-4488 [3].
- miRatio effectively predicts disease outcome when measured at baseline prior to therapy and exhibits dynamic changes during treatment [3].
Protocol 2: Copy Number Variation Analysis Using ddPCR
Purpose: Accurate determination of gene copy number variations in cancer research, using the DEFA1A3 gene as a model locus [5].
Materials and Reagents:
- Genomic DNA samples (40-100 ng/μL recommended)
- ddPCR Supermix for Probes (no dUTP)
- FAM-labeled TaqMan assay for target gene (DEFA1A3)
- HEX-labeled TaqMan assay for reference gene (RNase P or other 2-copy gene)
- DG8 Cartridges for droplet generation
- Droplet Generation Oil
- 96-well PCR plates
Procedure:
- Reaction Setup:
- Prepare 20 μL reaction mix containing:
- 10 μL 2× ddPCR Supermix
- 1 μL FAM-labeled target assay (20×)
- 1 μL HEX-labeled reference assay (20×)
- 50-100 ng genomic DNA
- Nuclease-free water to 20 μL
- Include negative controls without template.
- Prepare 20 μL reaction mix containing:
Droplet Generation:
- Transfer 20 μL reaction mix to DG8 Cartridge wells.
- Add 70 μL Droplet Generation Oil to each well.
- Place DG8 Gasket on cartridge.
- Generate droplets using QX200 Droplet Generator.
PCR Amplification:
- Carefully transfer 40 μL emulsified reaction to 96-well PCR plate.
- Seal plate with foil heat seal.
- Perform amplification with the following thermal profile:
- Enzyme activation: 95°C for 10 minutes
- 40 cycles of:
- Denaturation: 94°C for 30 seconds
- Annealing/Extension: 60°C for 60 seconds
- Enzyme deactivation: 98°C for 10 minutes
- Hold at 4°C
Droplet Reading and Analysis:
- Load plate into QX200 Droplet Reader.
- Analyze each well sequentially for FAM and HEX fluorescence.
- Use QuantaSoft software to identify positive and negative droplets for both targets.
- Calculate copy number using the formula:
- Copy Number = 2 × (Concentration of Target / Concentration of Reference)
Technical Notes:
- Optimal DNA input is 50-100 ng; too much DNA can lead to poor partitioning and inaccurate results [5].
- Reference gene should be a confirmed 2-copy gene per diploid genome.
- Results show 95% concordance with PFGE-determined copy numbers, significantly outperforming qPCR (60% concordance) [5].
- The method is accurate across a wide range of copy numbers (2-16 copies) [5].
Research Reagent Solutions
Table 2: Essential Reagents and Materials for dPCR Experiments
| Reagent/Material | Function | Application Examples | Considerations |
|---|---|---|---|
| TaqMan Assays | Sequence-specific detection with fluorescent probes | Rare mutation detection, copy number variation [4] | Enable multiplexing with different fluorophores; extensive validated portfolio available |
| dPCR Master Mix | Optimized reaction mixture for partitioning and amplification | All dPCR applications | Formulations vary by platform; may include different inhibitor-resistant polymerases |
| Partitioning Plates/Cartridges | Create nanoliter-scale reactions | Microchamber-based dPCR systems (QIAcuity) [6] | Fixed partition numbers (e.g., 26,000 partitions for QIAcuity Nanoplate 26k) |
| Droplet Generation Oil | Create water-in-oil emulsion for droplet-based systems | Droplet digital PCR (QX200) [6] | Requires specific surfactants for stability during thermal cycling |
| RNA Extraction Kits | Nucleic acid purification from complex samples | Liquid biopsy applications (serum, plasma) [3] | Efficiency critical for low-abundance targets; miRNeasy Mini Kit recommended for serum miRNAs |
| cDNA Synthesis Kits | Reverse transcription of RNA targets | miRNA analysis, gene expression [3] | TaqMan Advanced miRNA cDNA Synthesis Kit includes preamplification for low-abundance targets |
Technical Considerations and Data Analysis
Platform Selection
Table 3: Comparison of Digital PCR Platforms
| Platform | Partitioning Technology | Number of Partitions | Throughput | Key Features |
|---|---|---|---|---|
| QX200 Droplet Digital PCR (Bio-Rad) | Water-in-oil droplets | ~20,000 droplets per reaction | 96-well plate format | Established workflow, proven sensitivity [6] |
| QIAcuity (Qiagen) | Microfluidic nanoplates | 26,000-100,000 depending on plate | Integrated partitioning, thermocycling, and imaging [6] | Simplified workflow, reduced risk of contamination |
| QuantStudio Absolute Q (Thermo Fisher) | Microfluidic array plate (MAP) | ~20,000 wells per panel [4] | Modular automation for high-throughput | Microfluidic array plate technology with <5% dead volume [4] |
| Digital LightCycler (Roche) | Microchamber arrays | Varies by chip | Rapid cycling | Glass chip technology with surface chemistry |
Uncertainty Estimation and Data Analysis
Accurate variance estimation remains challenging in dPCR due to violations of standard statistical assumptions. Recent advances include two flexible methods (NonPVar and BinomVar) for calculating variance in dPCR data that improve standard error and confidence interval estimation [7]. These methods are particularly valuable for complex functions of partition counts like copy number variation, fractional abundance, and DNA integrity. Free computational tools (R Shiny app) are available to facilitate method selection and implementation [7].
For gene expression studies in oncology, proper reference gene validation is essential. Research on cervical precancer samples found that while GAPDH and ACTB were the most stable genes, they were expressed at very high levels, making them less suitable for normalizing lower-expression biomarkers [8]. Instead, GUSB and HMBS were recommended as a stable reference gene pair for dPCR gene expression analysis in liquid-based cytology samples [8].
Digital PCR represents a significant advancement in nucleic acid quantification technology, offering absolute quantification without standard curves, exceptional sensitivity for rare variant detection, and superior tolerance to inhibitors compared to qPCR. In oncology research, these characteristics make dPCR particularly valuable for liquid biopsy applications, minimal residual disease monitoring, copy number variation analysis, and precision medicine approaches requiring exact quantification of biomarkers. As the technology continues to evolve with improvements in multiplexing capabilities, workflow simplification, and data analysis methods, dPCR is poised to play an increasingly critical role in cancer research and clinical translation.
Digital PCR (dPCR) represents the third generation of polymerase chain reaction technology, enabling the absolute quantification of nucleic acids without the need for a standard curve. Its core principle relies on the partitioning of a sample into a multitude of individual reactions, followed by end-point detection and application of Poisson statistics to determine target concentration [1]. This calibration-free technology provides powerful advantages including high sensitivity, absolute quantification, high accuracy and reproducibility, and rapid turnaround time, making it particularly valuable for oncology research where precise measurement of rare mutations is critical [1]. The ability of dPCR to detect rare genetic mutations within a background of wild-type genes has paved the way for tumor heterogeneity analysis and liquid biopsy applications, revolutionizing how clinicians monitor treatment response in cancer patients [1].
Core Technical Principles
The Partitioning Process
Partitioning constitutes the foundational step of digital PCR, wherein the PCR mixture containing the sample is distributed across thousands to millions of discrete compartments. This process achieves a random distribution of nucleic acid molecules across the partitions such that each compartment contains either zero, one, or a few target molecules according to a Poisson distribution [1]. Two major partitioning methodologies have emerged: water-in-oil droplet emulsification (droplet digital PCR or ddPCR) and microchamber-based systems (chamber digital PCR or cdPCR) [1].
Droplet digital PCR creates partitions by dispersing the sample into nanoliter-sized droplets within an immiscible oil phase, typically using microfluidic chips that generate monodisperse droplets at high speeds (1-100 kHz) [1]. A key technical consideration is droplet stabilization with appropriate surfactants to prevent coalescence during thermal cycling [1]. Microchamber-based dPCR utilizes arrays of microscopic wells or chambers embedded in a solid chip, offering higher reproducibility and ease of automation at the cost of fixed partition numbers and typically higher expenses [1]. A hybrid approach known as Crystal Digital PCR combines aspects of both technologies by creating 2D monolayer arrays of monodisperse droplets, called "droplet crystals," within a microfluidic chip [9].
Table 1: Comparison of Digital PCR Partitioning Methods
| Partitioning Method | Partition Characteristics | Key Advantages | Common Platforms/Examples |
|---|---|---|---|
| Droplet Digital PCR (ddPCR) | Aqueous droplets in oil (pL-nL volume) | High scalability, cost-effectiveness | Bio-Rad QX200, Naica System (Sapphire chip) |
| Chamber Digital PCR (cdPCR) | Solid microchambers/wells | High reproducibility, ease of automation | QuantStudio 3D, QIAcuity |
| Crystal Digital PCR | 2D monolayer droplet arrays (0.43 nL mean volume) | Combines benefits of droplet and chamber approaches | Naica System (Stilla Technologies) |
End-Point Fluorescence Analysis
Following PCR amplification through thermal cycling, dPCR employs end-point fluorescence analysis to detect amplification in each partition [1]. This represents a fundamental distinction from quantitative PCR (qPCR), which monitors amplification in real-time. Partitions are classified as "positive" (containing amplified target) or "negative" (lacking target) based on fluorescence intensity exceeding a predetermined threshold [1].
Two primary readout methodologies exist for this analysis. Planar imaging utilizes fluorescence microscopy or scanners to capture a static image of microchamber arrays or deposited droplets, enabling simultaneous analysis of all partitions [1]. The Naica Prism3 system, for instance, employs a three-color fluorescence detection system with excitation bands at 415-480 nm (blue), 530-550 nm (green), and 615-645 nm (red), compatible with common fluorophores including FAM, VIC, HEX, and Cy5 [9]. Alternatively, in-line detection flows droplets sequentially through a detection channel where fluorescence is measured one-by-one using a light source coupled to detectors, analogous to flow cytometry [1].
Poisson Statistics for Absolute Quantification
The mathematical foundation of digital PCR relies on Poisson statistics to calculate target concentration from the fraction of positive partitions [1]. The model operates on the principle that nucleic acid molecules are randomly distributed across partitions, with many partitions containing zero molecules, some containing one molecule, and fewer containing multiple molecules.
The standard Poisson model estimates the average number of molecules per partition (λ) using the equation: λ = -ln(1 - p) where p represents the proportion of positive partitions [1]. The target concentration is then calculated by dividing λ by the partition volume. This approach assumes that all partitions have identical volumes, an assumption that can be violated in practical applications, particularly at higher concentrations [10].
To address partition volume variability, the Poisson-Plus model was developed, which accounts for effective load volume variation across partitions [10]. This model characterizes the distribution of partition volumes and incorporates this information into the concentration calculation, significantly improving quantification accuracy, especially when partition size variation is substantial [10]. The Poisson-Plus model expresses the probability of a partition being negative as: P(neg) = e^(½σ²C² - Cv₀) where C is the concentration, v₀ is the mean partition volume, and σ is the standard deviation of partition volumes [10].
Table 2: Key Parameters in Digital PCR Quantification
| Parameter | Description | Impact on Quantification |
|---|---|---|
| Number of Partitions | Total partitions analyzed | Higher numbers improve precision and dynamic range |
| Partition Volume | Volume of individual partitions | Critical for converting λ to concentration; inaccuracy introduces systematic error |
| Positive Partitions | Partitions showing amplification signal | Used with total partitions to calculate λ |
| Negative Partitions | Partitions without amplification signal | Proportion used in Poisson equation to calculate λ |
| λ (lambda) | Average number of molecules per partition | Fundamental parameter calculated from negative partition fraction |
| Volume Variation (σ/v₀) | Ratio of standard deviation to mean partition volume | Significant variation requires Poisson-Plus correction for accurate results |
Advanced Applications in Oncology Research
Multiplexing Strategies for Complex Biomarker Analysis
Digital PCR platforms support sophisticated multiplexing approaches that enable simultaneous detection of multiple targets within a single reaction, a critical capability for comprehensive oncology biomarker analysis. The standard "one color - one target" approach, while reliable, is inherently limited by the number of available fluorescence detection channels on the instrument [11]. To overcome this limitation, advanced strategies have been developed.
Amplitude-based multiplexing manipulates probe and primer concentrations to generate populations with different fluorescence amplitudes within a single detection channel [9]. While this approach increases multiplexing capacity, it may be affected by sample inhibitors or poor nucleic acid quality, potentially causing population smearing or displacement [9]. Color-combination multiplexing represents a more robust approach where targets are encoded by unique combinations of multiple fluorophores, dramatically expanding multiplexing capacity [11]. This method simplifies analysis by categorizing partitions into two groups: "all negative" partitions with low fluorescence across all channels, and partitions displaying high fluorescence for the specific fluorophore combination encoding a target sequence [11].
Protocol: Duplex dPCR for Circulating miRNA Quantification in Metastatic Melanoma
Background: Circulating miRNAs (cmiRNAs) have emerged as valuable non-invasive biomarkers for monitoring therapeutic response in metastatic melanoma. A duplex dPCR assay was developed for simultaneous detection of miR-4488 (oncogenic) and miR-579-3p (tumor-suppressive) to calculate their expression ratio (miRatio) as a predictive biomarker for MAPK inhibitor response [3] [12].
Methods:
Sample Preparation: Collect serum samples from BRAF-mutated metastatic melanoma patients prior to and during MAPK inhibitor therapy. Isolate total RNA from 200 μL serum using miRNeasy Mini Kit [3] [12].
Reverse Transcription: Perform reverse transcription using TaqMan Advanced miRNA cDNA Synthesis Kit with 2 μL input of total RNA (for serum samples) or 10 ng total RNA (for cellular controls) [3] [12].
dPCR Reaction Setup:
- Prepare duplex reaction mix containing:
- Primers and probes for both miR-4488 and miR-579-3p
- Fluorescently labelled probes (e.g., FAM and HEX/VIC)
- dPCR master mix
- cDNA template
- Load samples into appropriate partitioning device (e.g., Sapphire chip for Crystal Digital PCR) [9]
- Prepare duplex reaction mix containing:
Partitioning and Amplification:
- Partition samples using Naica Geode instrument or equivalent system
- Perform thermal cycling with appropriate amplification protocol
- Maintain overpressure (1 bar) throughout partitioning and amplification to preserve droplet integrity [9]
Fluorescence Detection and Analysis:
- Acquire three-color fluorescence images using Naica Prism3 or equivalent system
- Analyze images to classify partitions as double-positive, single-positive for either target, or double-negative
- Calculate absolute copies/μL for each miRNA using Poisson statistics
- Compute miRatio as expression ratio between miR-4488 and miR-579-3p [3] [12]
Results Interpretation: The duplex dPCR assay demonstrated superior sensitivity compared to qRT-PCR, particularly for low-abundance miR-4488. miRatio effectively predicted treatment outcome when measured at baseline and showed dynamic changes during therapy, supporting its utility as a longitudinal monitoring biomarker in metastatic melanoma [3] [12].
Protocol: Three-Color Crystal Digital PCR for EGFR Mutation Detection
Background: Detection of EGFR mutations (L858R, L861Q, T790M) in non-small cell lung cancer requires highly sensitive multiplexing capability to identify resistance mutations and guide targeted therapy decisions [9].
Methods:
Chip Preparation:
- Obtain primed Sapphire chips containing emulsion oil with stabilizing surfactants
- Remove inlet port caps and pipette 20 μL PCR mix into each of 4 inlet ports
- Seal with pressure-permeable caps [9]
Partitioning and Amplification:
- Place prepared chips into Naica Geode instrument
- Initiate program to increase pressure to 1 bar, driving PCR mix through 33 droplet production nozzles
- Monitor formation of monodisperse droplets (94 μm diameter, 0.43 nL volume) that self-arrange into hexagonal 2D monolayer (droplet crystal)
- Execute user-defined thermal cycling program while maintaining overpressure [9]
Three-Color Endpoint Detection:
- Transfer chips to Naica Prism3 automated fluorescence microscope
- Acquire high-resolution images using three independent LED excitation sources:
- Blue channel: 415-480 nm (ex. FAM, AlexaFluor488)
- Green channel: 530-550 nm (ex. VIC, HEX, Yakima Yellow)
- Red channel: 615-645 nm (ex. Cy5, Quasar705)
- Use tri-band emission filter (495-520 nm//560-610 nm//655-720 nm) [9]
Data Analysis with Spillover Compensation:
- Apply fluorescence spillover compensation to account for spectral overlap between channels
- Classify partitions based on three-color fluorescence patterns
- Quantify mutant allele frequencies using Poisson statistics
- Implement population identification algorithms to distinguish different mutation profiles [9]
Essential Research Reagent Solutions
Table 3: Key Research Reagent Solutions for Digital PCR in Oncology
| Reagent/Consumable | Function | Application Example |
|---|---|---|
| Sapphire Chip (Stilla Technologies) | Microfluidic chip for partitioning samples into 2D droplet arrays | Crystal Digital PCR workflow for EGFR mutation detection [9] |
| TaqMan Advanced miRNA cDNA Synthesis Kit | Reverse transcription and preamplification of miRNA targets | Preparation of cDNA from circulating miRNAs in metastatic melanoma serum samples [3] [12] |
| miRNeasy Mini Kit | RNA extraction from serum/plasma | Isolation of circulating miRNAs from liquid biopsy samples [3] [12] |
| Absolute Q Multiplex Oncology Assays | Pre-designed, pre-tested assay panels | Detection of cancer-related mutations without requiring assay optimization [13] |
| Droplet Stabilization Surfactants | Prevent droplet coalescence during thermal cycling | Maintain partition integrity in droplet-based dPCR systems [1] |
| Fluorophore-Labeled Probes (FAM, VIC, HEX, Cy5) | Target-specific detection in multiple channels | Multiplex detection of oncogenic mutations in three-color dPCR [9] |
The core mechanism of digital PCR—partitioning, end-point analysis, and Poisson statistics—provides a robust foundation for absolute quantification of nucleic acids in oncology research. The partitioning process enables single-molecule resolution, while end-point fluorescence detection offers binary readout of amplification events. Poisson statistics transforms this qualitative information into precise quantitative data, with advanced models like Poisson-Plus correcting for technical variations. These fundamental principles support increasingly sophisticated applications in oncology, from circulating miRNA profiling in metastatic melanoma to multiplexed mutation detection in lung cancer, positioning dPCR as an indispensable technology for precision medicine research.
The journey to absolute quantification of nucleic acids represents a cornerstone of modern molecular biology, particularly in the field of oncology research where precise measurement of rare mutations can dictate therapeutic decisions. This evolution began with technically demanding, low-throughput methods and has progressed to fully automated, digital platforms capable of single-molecule detection. The transition from limiting dilution techniques to today's commercial digital PCR (dPCR) systems has fundamentally transformed the capabilities of researchers and clinicians in detecting and quantifying genetic markers with unprecedented sensitivity and precision [1]. This application note details this technological progression, provides validated protocols for current dPCR applications in oncology, and outlines the essential tools required to implement these methodologies in a research setting.
The Path to Digital PCR: A Historical Timeline
The conceptual foundation of dPCR was laid in 1992 when Morley and Sykes combined limiting dilution PCR with Poisson statistics to isolate, detect, and quantify single nucleic acid molecules [1]. This method involved performing PCR on a series of sample dilutions and analyzing them by gel electrophoresis to count target molecules based on the fraction of negative reactions, enabling the detection of mutated sequences amidst a vast background of wild-type genes [1].
The term "digital PCR" was officially coined in 1999 by Bert Vogelstein and his team. They developed a workflow using limiting dilution distributed across 96-well plates combined with fluorescence readout to detect RAS oncogene mutations in the stools of patients with colorectal cancer [1]. A significant breakthrough came in 2003 with the development of the BEAMing technology, which simplified compartmentalization by using water-in-oil droplets to parallelize PCR reactions [1]. The subsequent commercialization of dPCR platforms, driven by advances in microfabrication and microfluidics, has provided the robust, high-throughput systems in use today [1].
The following diagram illustrates the key milestones in this developmental journey:
Quantitative Comparison of Modern dPCR Platforms
The current dPCR landscape is characterized by two main partitioning technologies: droplet-based systems and nanoplate-based systems. The table below summarizes the performance characteristics of two leading platforms as demonstrated in a cross-platform evaluation study.
Table 1: Performance Comparison of dPCR Platforms for Gene Copy Number Analysis
| Parameter | QIAGEN QIAcuity One (ndPCR) | Bio-Rad QX200 (ddPCR) |
|---|---|---|
| Partitioning Technology | Nanoplate-based (microchambers) [1] | Droplet-based (water-in-oil) [1] |
| Partition Volume | Nanoliter-scale chambers [1] | Picoliter-scale droplets [1] |
| Limit of Detection (LOD) | 0.39 copies/µL input [14] | 0.17 copies/µL input [14] |
| Limit of Quantification (LOQ) | 1.35 copies/µL input [14] | 4.26 copies/µL input [14] |
| Precision (CV range) | 7% - 11% (with synthetic oligos) [14] | 6% - 13% (with synthetic oligos) [14] |
| Restriction Enzyme Impact | Lower impact on precision [14] | Higher precision with HaeIII vs. EcoRI [14] |
| Best Precision Range | ~31 - 534 copies/µL input [14] | ~270 copies/µL input [14] |
This empirical comparison highlights that while the QX200 ddPCR system offers a marginally superior LOD, the QIAcuity One ndPCR system demonstrates a better LOQ and is less affected by the choice of restriction enzyme, a crucial factor when analyzing complex genomic DNA with potential tandem repeats [14].
Application Note: Detection of a Rare Oncogenic Mutation
Background and Principle
The ability to detect and quantify a rare mutant allele (e.g., a KRAS G12D mutation) within a background of wild-type DNA is critical for cancer diagnosis, monitoring minimal residual disease, and tracking therapy resistance [13]. dPCR is ideally suited for this application because it partitions the sample, effectively enriching the rare target to a detectable concentration in a subset of partitions, allowing for absolute quantification without a standard curve [15] [13].
Protocol: Rare Mutation Detection via Probe-Based ddPCR
Methodology: This protocol uses allele-specific TaqMan probes to differentially detect wild-type and mutant KRAS sequences in a duplex reaction [13].
Workflow Overview:
Step-by-Step Procedure:
Reaction Setup:
- Prepare a 20-22 µL reaction mix on ice containing:
- 10 µL of 2x ddPCR Supermix for Probes (no dUTP).
- 1 µL of 20x KRAS G12D Wild-Type Assay (FAM-labeled probe).
- 1 µL of 20x KRAS G12D Mutation Assay (HEX/VIC-labeled probe).
- 50-100 ng of template DNA (from patient plasma, FFPE tissue, or cell lines).
- Nuclease-free water to the final volume.
- Vortex gently and centrifuge to collect the mixture at the bottom of the tube.
- Prepare a 20-22 µL reaction mix on ice containing:
Droplet Generation:
- Transfer the 20 µL reaction mix to the sample well of a DG8 cartridge.
- Pipette 70 µL of Droplet Generation Oil into the oil well.
- Place the cartridge into the Droplet Generator. The QX200 system will automatically generate ~20,000 nanodroplets.
- Carefully transfer the emulsified sample (~40 µL) to a 96-well PCR plate. Seal the plate with a foil heat seal.
PCR Amplification:
- Place the sealed plate in a thermal cycler and run the following protocol:
- Step 1: Enzyme activation at 95°C for 10 minutes.
- Step 2: 40 cycles of:
- Denaturation: 94°C for 30 seconds.
- Annealing/Extension: 55-60°C (assay-specific) for 60 seconds.
- Step 3: Enzyme deactivation at 98°C for 10 minutes.
- Step 4: Hold at 4°C. (Ramp rate: 2°C/second).
- Place the sealed plate in a thermal cycler and run the following protocol:
Droplet Reading and Analysis:
- Transfer the PCR plate to the QX200 Droplet Reader.
- The reader will aspirate each sample and stream the droplets single-file past a two-color optical detection system.
- Using the associated analysis software, set thresholds to distinguish four populations: 1) mutant-positive (FAM), 2) wild-type-positive (HEX), 3) double-positive, and 4) double-negative droplets.
Quantification and Quality Control:
- The software will apply Poisson statistics to the fraction of positive droplets to calculate the absolute concentration (copies/µL) of both wild-type and mutant targets in the original sample.
- Calculate the mutant allele frequency (MAF) as: (Mutant concentration / (Mutant + Wild-type concentration)) * 100.
- Ensure the total number of accepted droplets is >10,000 for reliable quantification.
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful implementation of dPCR requires a suite of reliable reagents and tools. The following table details key materials for setting up a dPCR workflow in an oncology research laboratory.
Table 2: Essential Reagents and Materials for dPCR Oncology Research
| Item | Function / Description | Example Application in Oncology |
|---|---|---|
| dPCR Supermix | A chemical formulation containing DNA polymerase, dNTPs, buffer, and stabilizers optimized for partition-based PCR. | Core component of all dPCR reactions for target amplification [13]. |
| TaqMan Assays | Hydrolysis probes (FAM, HEX, VIC, etc.) and primer sets designed for specific mutation detection. | Detection of oncogenic mutations (e.g., EGFR, BRAF, KRAS) and reference genes in a duplex/multiplex format [13]. |
| Restriction Enzymes | Enzymes that digest DNA at specific recognition sites. | Used to fragment long genomic DNA to ensure efficient encapsulation of target sequences and access to tandemly repeated genes (e.g., HaeIII, EcoRI) [14]. |
| Droplet Generation Oil & Cartridges | Consumables for generating a stable water-in-oil emulsion. | Essential for creating the thousands of partitions in droplet-based systems like the QX200 [1]. |
| Optical Seals & Plates | Seals and plates compatible with thermal cycling and fluorescence reading. | Prevent evaporation and cross-contamination during the PCR process. |
| Reference Genomic DNA | DNA from well-characterized cell lines with known mutation status. | Critical for assay validation, as a positive control, and for determining the limit of detection [14]. |
| Synthetic Oligonucleotides | Custom-designed DNA sequences mimicking wild-type and mutant targets. | Used for absolute standard curve generation and optimizing assay conditions without source DNA [14]. |
The evolution from manual limiting dilution to automated commercial dPCR platforms has provided oncology researchers with a powerful tool for absolute quantification. The high sensitivity, precision, and robustness of these systems enable applications that were previously challenging or impossible, such as monitoring low-level resistance mutations and validating biomarkers from liquid biopsies. By leveraging the protocols and tools outlined in this document, researchers can reliably implement these advanced techniques to accelerate discoveries in cancer biology and therapeutic development.
Digital PCR (dPCR) represents a transformative approach in molecular biology, enabling the precise detection and absolute quantification of nucleic acids. As a third-generation PCR technology, it operates on a fundamentally different principle than quantitative real-time PCR (qPCR). Unlike qPCR, which relies on the kinetics of amplification during the exponential phase and requires a standard curve for relative quantification, dPCR achieves absolute quantification by partitioning a sample into thousands of nanoliter-scale reactions, performing endpoint PCR, and applying Poisson statistics to count individual molecules [16] [17]. This direct counting method eliminates the need for external calibrators and provides unmatched precision for applications demanding high sensitivity, particularly in the field of oncology research where detecting rare mutations or subtle genetic variations can directly impact therapeutic decisions [13] [18].
The core principle of dPCR involves dividing a PCR reaction into a large number of partitions such that each contains either zero or a limited number of target molecules [19]. Following amplification, each partition is analyzed for fluorescence. Partitions containing the target sequence (positive) are counted against those without (negative). The absolute concentration of the target nucleic acid in the original sample is then calculated using Poisson distribution statistics to account for the random distribution of molecules, providing a direct count in copies per microliter without reference to standards [16] [20] [17]. This white paper details the key advantages of this methodology and provides specific application protocols for oncology research.
Core Analytical Advantages
Absolute Quantification Without Standard Curves
The independence from standard curves constitutes one of the most significant advantages of dPCR. In traditional qPCR, quantification is indirect. The cycle threshold (CT) value of an unknown sample is compared to a standard curve generated from samples of known concentration, introducing potential errors from pipetting inaccuracies during standard dilution, differences in amplification efficiency between the standard and the target, and inter-assay variability [16] [21] [22]. dPCR circumvents these issues entirely by providing a direct digital count of target molecules [20].
Mechanism of Absolute Quantification: The partitioning step is crucial. When a sample is divided into thousands of partitions (e.g., 20,000 droplets or micro-wells), most partitions contain either zero or one target molecule. After endpoint PCR amplification, the ratio of positive to negative partitions is used in the Poisson equation (copies/μL = -ln(1-p) / V, where p is the fraction of positive partitions and V is the partition volume) to calculate the absolute target concentration [16] [17] [22]. This process converts an analog, relative measurement into a digital, absolute count.
Impact on Reproducibility: This direct counting method dramatically improves reproducibility between laboratories and instruments. Studies have shown that dPCR exhibits significantly lower coefficients of variation (CV) compared to qPCR, especially at low target concentrations. For instance, in viral load testing, dPCR demonstrated an average CV of 11.7%, compared to 25.8% for qPCR [22]. This enhanced precision is critical for longitudinal monitoring of disease biomarkers in oncology, such as tracking minimal residual disease (MRD) through circulating tumor DNA (ctDNA) [16].
Superior Sensitivity and Precision
dPCR excels in detecting and quantifying rare targets within complex backgrounds, a common challenge in cancer genomics.
Rare Allele Detection: The physical separation of target molecules in dPCR effectively enriches rare sequences and eliminates competition for reagents from the more abundant, non-target DNA. This allows for the detection of mutant alleles present at frequencies as low as 0.01% in a wild-type background [13] [18]. This sensitivity is paramount for liquid biopsy applications, where ctDNA fragments carrying oncogenic mutations (e.g., in EGFR, BRAF, or KRAS) can constitute less than 0.1% of the total cell-free DNA in a patient's blood [16].
Robustness to Inhibitors: dPCR is notably more tolerant to common PCR inhibitors found in clinical samples (e.g., hemoglobin, heparin, bile salts). Inhibitors are diluted across the many partitions, minimizing their effective concentration in any single reaction. Furthermore, since dPCR is an endpoint measurement, it does not rely on amplification kinetics. A delayed amplification due to a minor inhibitor will still result in a positive signal as long as the reaction reaches its endpoint, whereas the same delay would significantly alter the CT value and calculated concentration in qPCR [16] [17]. This robustness simplifies sample preparation and increases the reliability of results from complex matrices like plasma, stool, or FFPE tissues [16].
Table 1: Key Performance Advantages of dPCR over qPCR
| Analytical Parameter | Digital PCR (dPCR) | Quantitative PCR (qPCR) |
|---|---|---|
| Quantification Method | Absolute, via direct counting [20] [17] | Relative, requires a standard curve [20] |
| Precision (at low concentration) | High (CVs ~10-15%) [21] [22] | Lower (CVs can be 20-30% or higher) [21] [22] |
| Sensitivity for Rare Alleles | Very High (can detect <0.1%) [16] [13] | Limited (typically >1%) |
| Effect of PCR Inhibitors | High tolerance [16] [17] | Sensitive [16] |
| Dynamic Range | Linear over a wide concentration range [16] | Limited by the standard curve [20] |
Applications in Oncology Research
The unique analytical strengths of dPCR make it an indispensable tool for addressing critical questions in cancer research and drug development.
Liquid Biopsy and Rare Mutation Detection
Liquid biopsy, the analysis of tumor-derived material like ctDNA from blood plasma, offers a non-invasive method for cancer diagnosis, prognosis, and monitoring therapy response. dPCR is the gold-standard technology for validating and quantifying low-frequency mutations in ctDNA due to its superior sensitivity and precision [16] [13] [18]. It is routinely used to monitor the emergence of therapy-resistant clones by tracking specific somatic mutations over time, enabling earlier treatment adjustments than would be possible with radiographic imaging.
Copy Number Variation (CNV) Analysis
Gene amplifications and deletions are key drivers in many cancers (e.g., HER2 amplification in breast cancer). dPCR provides exceptional resolution for CNV analysis by simultaneously and absolutely quantifying the target gene and a reference gene in a single multiplex reaction. The high precision of dPCR allows for the confident discrimination of small, sub-fold changes in copy number, which is challenging with qPCR's standard curve-based approach [16] [17] [18].
Validation of Next-Generation Sequencing (NGS) Findings
dPCR serves as a powerful orthogonal method for validating genetic alterations identified by NGS. It provides an independent, highly quantitative, and cost-effective means to confirm the presence and frequency of specific mutations, fusions, or copy number alterations in a subset of samples, thereby increasing the confidence in NGS data [18].
Table 2: Key Oncology Research Applications for dPCR
| Application Area | Primary Benefit of dPCR | Specific Examples |
|---|---|---|
| Liquid Biopsy / Rare Mutation Detection | Superior sensitivity for low-frequency alleles [16] [13] | Quantifying ctDNA; monitoring EGFR T790M mutations in NSCLC; detecting KRAS mutations [16] [18] |
| Copy Number Variation (CNV) Analysis | High precision for determining gene copy numbers without a standard curve [16] [17] | Determining HER2 amplification status; assessing MYC amplifications [13] [18] |
| Treatment Monitoring & MRD | Accurate, reproducible quantification for tracking minimal disease [16] | Detecting molecular relapse post-treatment; monitoring MRD [16] |
| NGS Validation | Absolute quantification for orthogonal confirmation [18] | Validating SNP, fusion, and CNV calls from NGS panels [18] |
Experimental Protocols
Protocol 1: Detection of Rare Mutations in Cell-Free DNA for Liquid Biopsy
Objective: To absolutely quantify a low-frequency somatic mutation (e.g., EGFR p.T790M) in plasma-derived cell-free DNA (cfDNA).
Principle: A duplex dPCR assay is designed with two probe-based assays: one specific for the mutant allele (e.g., VIC-labeled) and one for the wild-type sequence (e.g., FAM-labeled). The sample is partitioned, and the number of mutant-positive partitions is counted to determine the absolute mutant allele frequency [16].
Materials:
- QX200 Droplet Digital PCR System (Bio-Rad) or equivalent [21]
- ddPCR EvaGreen Supermix or TaqMan ddPCR Supermix for Probes
- FAM-labeled TaqMan assay for wild-type EGFR
- VIC-labeled TaqMan assay for EGFR p.T790M mutation
- DG8 Cartridges and Droplet Generation Oil
- Thermal Sealer
- 96-well PCR plates
Workflow Diagram:
Diagram Title: dPCR Liquid Biopsy Workflow
Step-by-Step Procedure:
- Reaction Setup: On ice, prepare a 20 µL reaction mix containing:
- 10 µL of 2x ddPCR Supermix for Probes.
- 1 µL of EGFR Wild-Type Assay (FAM).
- 1 µL of EGFR T790M Assay (VIC).
- 8 µL of extracted cfDNA (typically 5-50 ng).
- Mix thoroughly by pipetting; do not vortex.
Droplet Generation: Transfer 20 µL of the reaction mix to a DG8 cartridge. Carefully add 70 µL of Droplet Generation Oil to the oil well. Place the cartridge in the QX200 Droplet Generator. Following droplet generation, carefully transfer the emulsified sample (~40 µL) to a 96-well PCR plate. Seal the plate with a foil heat seal.
PCR Amplification: Place the sealed plate in a thermal cycler and run the following protocol:
- Enzyme activation: 95°C for 10 minutes.
- 40-45 cycles of:
- Denaturation: 94°C for 30 seconds.
- Annealing/Extension: 55-60°C for 1 minute (optimize based on assay).
- Enzyme deactivation: 98°C for 10 minutes.
- Hold at 12°C.
Droplet Reading: Transfer the PCR plate to the QX200 Droplet Reader. The reader will aspirate each sample, measuring the fluorescence in each droplet (FAM and VIC channels).
Data Analysis: Use the instrument's analysis software (e.g., QuantaSoft). Set amplitude thresholds to clearly distinguish positive and negative droplet populations for both channels. The software will automatically apply Poisson statistics to calculate the absolute concentration (copies/µL) of wild-type and mutant DNA in the original sample. Calculate the mutant allele frequency: (Mutant concentration / (Mutant + Wild-type concentration)) * 100.
Protocol 2: Absolute Copy Number Variation (CNV) Analysis
Objective: To determine the absolute copy number of a target gene (e.g., HER2) relative to a reference gene (e.g., RNase P) in genomic DNA.
Principle: A duplex dPCR reaction simultaneously quantifies the target and reference genes. The absolute copy number of the target per genome equivalent is calculated based on the known diploid copy number (2) of the reference gene [17] [18].
Materials:
- Absolute Q Digital PCR System (Thermo Fisher) or equivalent [13] [17]
- dPCR Master Mix (e.g., Absolute Q Digital PCR Master Mix)
- FAM-labeled TaqMan assay for the target gene (HER2)
- VIC-labeled TaqMan assay for the reference gene (RNase P)
- Nanoplate (e.g., QuantStudio Absolute Q Digital PCR Plate)
- Plate Sealer
Workflow Diagram:
Diagram Title: dPCR CNV Analysis Workflow
Step-by-Step Procedure:
- Reaction Setup: Prepare the dPCR reaction mix according to the manufacturer's instructions for a final volume of 25-40 µL (varies by platform). The mix should contain:
- 1x dPCR Master Mix.
- Optimized concentrations of FAM-labeled HER2 assay and VIC-labeled RNase P assay.
- 10-50 ng of restriction enzyme-digested genomic DNA (digestion is recommended to disrupt DNA aggregates and ensure accurate partitioning).
Loading and Partitioning: Pipette the entire reaction mix into the designated well of the nanoplate. Seal the plate. Place the sealed plate into the dPCR instrument. The instrument will automatically perform the partitioning of the sample into thousands of nanoscale chambers.
PCR Amplification and Imaging: The instrument runs a standardized endpoint PCR protocol. Upon completion, it automatically scans each chamber to capture fluorescence data for both FAM and VIC channels.
Data Analysis: The instrument's software automatically identifies positive and negative partitions for both target and reference assays and calculates their absolute concentrations in copies/µL.
Copy Number Calculation:
- Calculate the concentration ratio: R = [Target] / [Reference].
- The expected ratio for a diploid gene is 1.0 (2 copies / 2 copies).
- The estimated copy number of the target gene is: R * 2.
- For example, a ratio of 1.5 would indicate an estimated HER2 copy number of 3.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Reagents and Materials for dPCR Experiments
| Item | Function/Description | Example Products/Brands |
|---|---|---|
| dPCR Instrument | Partitions the sample, performs thermal cycling, and reads fluorescence endpoint. | QX200 Droplet Digital PCR System (Bio-Rad), QuantStudio Absolute Q (Thermo Fisher), QIAcuity (QIAGEN) [21] [17] [14] |
| dPCR Master Mix | Optimized buffer containing DNA polymerase, dNTPs, and salts for efficient dPCR amplification. | ddPCR Supermix (Bio-Rad), Absolute Q Master Mix (Thermo Fisher), QIAcuity Probe PCR Kit (QIAGEN) |
| TaqMan Assays | Sequence-specific primers and fluorescently labeled probes for target detection. | Pre-designed assays for oncology targets (e.g., EGFR, KRAS, BRAF, HER2) [13] [18] |
| Partitioning Consumables | Cartridges, oil, and plates required for creating nanoscale reactions. | DG8 Cartridges & Droplet Generation Oil (Bio-Rad), Absolute Q Digital PCR Plates (Thermo Fisher) [21] [17] |
| Nucleic Acid Template | The sample of interest (DNA, cDNA, cfDNA). | Purified from cells, tissues, or plasma. Use high-quality extraction kits. |
| Restriction Enzymes | Used to digest genomic DNA to prevent aggregation and ensure random distribution during partitioning. | EcoRI, HaeIII (selection depends on amplicon sequence) [14] |
dPCR in Action: Methodological Workflows and Translational Applications in Oncology
Digital PCR (dPCR) represents a third-generation PCR technology that enables absolute quantification of nucleic acids without the need for a standard curve [1]. By partitioning a sample into thousands of individual reactions, dPCR allows for the precise counting of target molecules using Poisson statistics, offering superior sensitivity, accuracy, and resistance to inhibitors compared to quantitative PCR (qPCR) [6] [23] [18]. This Application Note details the standard dPCR workflow, with a specific focus on its critical applications in oncology research, including the detection of rare somatic mutations, copy number variations (CNVs), and DNA methylation in circulating tumor DNA (ctDNA) for liquid biopsy [24] [23] [18]. The provided protocols and data are tailored to support researchers and drug development professionals in implementing this powerful technology.
Digital PCR (dPCR) is a method for the absolute quantification of nucleic acid molecules. The core principle involves distributing a PCR reaction mix across a large number of discrete partitions, such that each contains zero, one, or a few target molecules [1]. Following end-point amplification, each partition is analyzed for fluorescence, and the fraction of positive partitions is used to calculate the absolute concentration of the target sequence in the original sample based on Poisson distribution statistics [1] [23]. This compartmentalization allows dPCR to excel in applications requiring high sensitivity and precision, such as detecting rare mutant alleles in a background of wild-type DNA for minimal residual disease (MRD) monitoring in hematologic malignancies [23], identifying CNVs in cancer genomes [5] [18], and quantifying tumor-specific methylation markers in liquid biopsies [24].
Standard dPCR Workflow
The standard dPCR workflow consists of four main stages: sample and assay preparation, partitioning and amplification, fluorescence reading, and data analysis. The following diagram and sections detail each step.
Detailed Experimental Protocols
Protocol 1: Basic dPCR Setup for Absolute Quantification This protocol is adapted for the detection of genetic alterations, such as single-nucleotide variants (SNVs) or CNVs, from genomic DNA [23] [5].
Sample Preparation:
- Extract genomic DNA from patient samples (e.g., blood, bone marrow, or tumor tissue) using a validated method (e.g., Maxwell RSC Instrument with PureFood GMO kit or similar) [6].
- Quantify DNA concentration using a fluorescence-based method. For CNV analysis, ensure DNA integrity by assessing the ratio of long to short amplicons (e.g., 250 bp vs. 65 bp amplicons of the EMC7 gene) [24].
- Dilute DNA to a working concentration in nuclease-free water. The optimal final amount per reaction is typically 1-100 ng, depending on the application and the expected target copy number [6] [5].
Master Mix Assembly:
- Prepare a reaction mix on ice. A typical 20-22 µL reaction for a droplet-based system might include:
- 10 µL of 2x ddPCR Supermix (or platform-specific master mix).
- 1.8 µL of each primer (final concentration 900 nM each).
- 0.5 µL of each probe (final concentration 250 nM each). For duplex assays, use probes labeled with different fluorophores (e.g., FAM and HEX/VIC).
- X µL of DNA template (e.g., 5 µL of 10 ng/µL DNA).
- Nuclease-free water to the final volume.
- Mix thoroughly by pipetting. Avoid vortexing after adding the master mix to prevent foam formation.
- Prepare a reaction mix on ice. A typical 20-22 µL reaction for a droplet-based system might include:
Partitioning:
- Droplet-based systems (e.g., Bio-Rad QX200): Transfer the entire reaction mix to the sample well of a DG8 cartridge. Pipette 70 µL of droplet generation oil into the oil well. Place the cartridge and a rubber gasket into the droplet generator. After generation, carefully transfer the resulting ~40 µL of droplets to a 96-well PCR plate [6].
- Chip-based systems (e.g., Qiagen QIAcuity): Load the reaction mix directly into the designated wells of a nanoplate (e.g., 26k partitions per well). The instrument performs automated partitioning and sealing [6].
PCR Amplification:
- Seal the plate with a foil heat seal (for droplet systems) or use the integrated seal (for chip systems).
- Place the plate in a thermal cycler and run a standard PCR protocol. An example for a TaqMan-based assay:
- Enzyme activation: 95°C for 10 minutes.
- 40-45 cycles of:
- Denaturation: 94°C for 30 seconds.
- Annealing/Extension: 55-60°C for 60 seconds (optimize temperature based on assay design).
- Enzyme deactivation: 98°C for 10 minutes.
- Hold at 4°C or 12°C.
Post-Amplification Handling:
- For droplet systems, ensure the plate is sealed correctly before reading to prevent droplet evaporation.
Protocol 2: Methylation-Specific ddPCR for Lung Cancer ctDNA Analysis This protocol details the application of dPCR for detecting DNA methylation biomarkers in plasma-derived cell-free DNA (cfDNA) [24].
Plasma Collection and cfDNA Extraction:
- Collect whole blood in EDTA tubes and centrifuge at 2,000 g for 10 minutes within 4 hours of venepuncture to isolate plasma.
- Perform a second centrifugation of the plasma at 10,000 g for 10 minutes to remove residual cells.
- Extract cfDNA from 4 mL of plasma using a dedicated kit (e.g., DSP Circulating DNA Kit on QIAsymphony SP). Elute DNA in 60 µL of elution buffer [24].
Bisulfite Conversion:
- Concentrate the extracted cfDNA to 20 µL using a centrifugal filter unit (e.g., Amicon Ultra-0.5).
- Treat the DNA with bisulfite using a commercial kit (e.g., EZ DNA Methylation-Lightning Kit) to convert unmethylated cytosines to uracils. Elute the converted DNA in 15 µL of elution buffer [24].
Methylation-Specific ddPCR Assay:
- Design primers and probes that specifically target the methylated (bisulfite-converted) sequence of interest (e.g., HOXA9 and other lung cancer-specific markers) [24].
- Assemble the ddPCR master mix as in Protocol 1, using the bisulfite-converted DNA as template.
- Perform partitioning, amplification, and reading as described in Protocol 1.
Data Analysis and Interpretation
After the run, the dPCR software analyzes each partition and generates a plot (1D or 2D) to distinguish positive from negative populations.
Analysis Workflow
- Threshold Setting: The software automatically applies a fluorescence threshold to classify partitions as positive or negative. For duplex assays, 2D amplitude plots are used to distinguish four populations: double-negative, FAM-positive, HEX/VIC-positive, and double-positive [6].
- Concentration Calculation: The software uses the fraction of positive partitions (λ) and the total partition volume to calculate the absolute concentration in copies per microliter of the input reaction using the Poisson distribution formula: Concentration = -ln(1 - λ) / Partition Volume [1] [23].
- Interpretation:
- For rare mutation detection (e.g., JAK2V617F), the result is often expressed as a mutant allele frequency (MAF) or fractional abundance: MAF = (Concentration of Mutant Allele) / (Concentration of Reference Allele + Concentration of Mutant Allele). dPCR can reliably detect MAFs as low as 0.01% to 0.001% [23].
- For CNV analysis, the copy number is calculated by taking the ratio of the target gene concentration to the concentration of a reference gene (assumed to be two copies per diploid genome). For example: CN = 2 × (Concentration of Target Gene / Concentration of Reference Gene) [5] [18].
Performance Data and Validation
Robust validation is crucial for implementing dPCR in a research or regulated environment. The following tables summarize key performance metrics from recent studies.
Table 1: Validation Parameters for dPCR in GMO and CNV Analysis [6] [5]
| Parameter | Description | Result (Bio-Rad QX200) | Result (Qiagen QIAcuity) |
|---|---|---|---|
| Dynamic Range | Linear range of quantification | 0.1% to 10% GMO [6] | 0.1% to 10% GMO [6] |
| Linearity | R² value of measured vs. expected concentration | > 0.998 [6] | > 0.998 [6] |
| Precision | Repeatability (Coefficient of Variation, CV) | < 5% for %GMO [6] | < 10% for %GMO [6] |
| Limit of Blank (LoB) | Highest result from a blank sample | Not Detected [6] | Not Detected [6] |
| Limit of Detection (LoD) | Lowest concentration reliably detected | 0.05% GMO [6] | 0.05% GMO [6] |
| Concordance with Gold Standard | Comparison with PFGE for CNV | 95% [5] | Not Reported |
Table 2: Performance of Methylation-Specific ddPCR in Lung Cancer Detection [24]
| Performance Metric | Non-Metastatic (Stage I-III) Disease | Metastatic (Stage IV) Disease |
|---|---|---|
| Sensitivity (Positive Rate) | 38.7% - 46.8%* | 70.2% - 83.0%* |
| Specificity | > 99% (in healthy controls) | > 99% (in healthy controls) |
| Markers Analyzed | Five-gene multiplex (e.g., HOXA9) | Five-gene multiplex (e.g., HOXA9) |
| Sample Type | Plasma | Plasma |
| Application | Early detection, MRD | Treatment monitoring, prognosis |
*Sensitivity varied based on the statistical cut-off method used to determine ctDNA positivity [24].
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents and Materials for dPCR Workflows
| Item | Function | Example Products / Notes |
|---|---|---|
| dPCR Master Mix | Provides DNA polymerase, dNTPs, and optimized buffers for amplification. | ddPCR Supermix for Probes (Bio-Rad), QIAcuity Probe PCR Master Mix (Qiagen). Must be compatible with the partitioning system. |
| Primers & Probes | Target-specific oligonucleotides for amplification and detection. | Hydrolysis probes (TaqMan) are standard. Must be designed for high specificity and efficiency. Methylation-specific primers are used for epigenetic analysis [24]. |
| DNA Extraction Kits | Isolate high-quality genomic DNA or cfDNA from various sample types. | Maxwell RSC PureFood GMO Kit (tissue), DSP Circulating DNA Kit (plasma cfDNA) [24] [6]. |
| Bisulfite Conversion Kit | Chemically converts unmethylated cytosine to uracil for methylation analysis. | EZ DNA Methylation-Lightning Kit [24]. Efficiency of conversion is critical for assay performance. |
| Droplet Generation Oil | Creates a water-in-oil emulsion for partitioning in droplet-based systems. | DG32 Droplet Generation Oil for Probes (Bio-Rad). Requires specific surfactants for stability during thermocycling [1]. |
| Partitioning Plates/Cartridges | The physical consumable used to generate the nanoscale reactions. | DG8 Cartridges & Gaskets (Bio-Rad QX200), QIAcuity Nanoplate 26k (Qiagen) [6]. |
The standard dPCR workflow provides a robust and precise method for the absolute quantification of nucleic acids. Its unparalleled sensitivity and accuracy make it an indispensable tool in oncology research, particularly for liquid biopsy applications, rare mutation detection, and copy number variation analysis. By following the detailed protocols and validation frameworks outlined in this Application Note, researchers can reliably implement dPCR to accelerate biomarker discovery, therapy monitoring, and ultimately, the development of personalized cancer treatments.
Digital PCR (dPCR) represents the third generation of PCR technology, enabling the absolute quantification of nucleic acids without the need for a standard curve. This calibration-free technology provides powerful advantages for liquid biopsy analysis, including high sensitivity, absolute quantification, high accuracy and reproducibility, as well as a rapid turnaround time [1]. In liquid biopsy applications, dPCR plays a crucial role in detecting and quantifying circulating tumor DNA (ctDNA) and other cancer-derived biomarkers from minimal sample volumes.
The fundamental principle of dPCR involves partitioning a PCR mixture supplemented with the sample into a large number of parallel reactions so that each partition contains either 0, 1, or a few nucleic acid targets, following a Poisson distribution. Following PCR amplification, the fraction of positive partitions is extracted from an end-point measurement, allowing computation of the absolute target concentration [1]. This single-molecule detection capability makes dPCR particularly valuable for identifying rare genetic mutations within a background of wild-type genes, which is essential for monitoring tumor dynamics through liquid biopsies [1].
Liquid biopsy itself involves the extraction of tumor-derived components such as circulating tumor cells (CTCs), ctDNA, and tumor extracellular vesicles (EVs) from the bodily fluids of cancer patients [25]. These components provide vital longitudinal information to enhance diagnostic precision for both primary and metastatic malignancies. The minimally invasive nature of liquid biopsy allows for serial sampling, facilitating longitudinal disease progression and treatment response monitoring [25]. Compared to traditional tissue biopsies, liquid biopsies offer significant advantages including minimal invasiveness, accessibility for serial sampling, assessment of tumor heterogeneity, potential for early detection, and lower cost [25].
Application Notes
Rare Mutation Detection in Hemato-Oncology
Digital PCR serves as a valuable tool complementing other molecular techniques in hemato-oncology research. The technology enables reliable quantification of low-abundance variants even in a high-wild-type background, making it ideal for detecting fusion genes like BCR-ABL and mutations such as the NPM1 Type A insertion with sensitivity down to 0.001% frequency [26]. This high sensitivity is critical for monitoring minimal residual disease (MRD) and guiding treatment decisions in leukemia patients.
The compartmentalization of the dPCR reaction renders it more tolerant to PCR inhibitors than qPCR, which is particularly advantageous when working with blood and bone marrow samples where inhibitors often co-purify with nucleic acids [26]. This tolerance ensures more robust quantification of mutant allele frequencies, even in challenging sample types. Furthermore, dPCR provides greater flexibility for multiplexing variants and wild-type sequences in the same reaction, enabling simultaneous assessment of multiple biomarkers [26].
Table 1: Performance Characteristics of dPCR in Hemato-Oncology Applications
| Parameter | Performance | Significance |
|---|---|---|
| Sensitivity | Detection down to 0.001% mutant allele frequency [26] | Enables MRD monitoring and early relapse detection |
| Precision | High intra-batch and inter-batch reproducibility [27] | Ensures reliable longitudinal monitoring |
| Dynamic Range | Wide linear range (e.g., 13.45-129,693 copies/μL for miRNA assays) [27] | Allows quantification across varying tumor burdens |
| Multiplexing Capacity | Simultaneous detection of 4 mutations plus wild-type with 2 fluorophores [28] | Comprehensive biomarker profiling from limited samples |
Ultrasensitive miRNA Detection in Hepatocellular Carcinoma
A recent application of droplet digital PCR (ddPCR) with locked nucleic acid (LNA)-modified probes has demonstrated an ultrasensitive and standardized method for miRNA quantification in liquid biopsies. Researchers developed a robust assay for miR-192-5p, a liver-enriched miRNA downregulated in hepatocellular carcinoma (HCC) [27]. The LNA probe technology improved positive droplet counts by 32%, enhancing the assay's sensitivity for detecting low-abundance plasma miRNAs [27].
The validated assay showed excellent precision with intra-batch CV of 2.31-21.63% and inter-batch CV of 17.54%, with sensitivity thresholds of LoB=1.75, LoD=3.33, and LoQ=13.45 copies/μL across a linear range of 13.45-129,693 copies/μL (R²=0.9965) [27]. When applied to clinical samples, HCC patients showed significantly lower miR-192-5p levels (444.2 vs. 753.5 copies/μL, p<0.001) with an AUC of 0.70 for distinguishing HCC from controls [27]. The development of such standardized workflows resolves significant barriers in miRNA liquid biopsy analysis and enables precise quantification of cancer-specific miRNAs.
Table 2: Analytical Validation of ddPCR Assay for miR-192-5p in HCC [27]
| Validation Parameter | Result | Acceptance Criteria |
|---|---|---|
| Trueness (vs. RT-qPCR) | R=0.92 | High correlation with reference method |
| Intra-batch Precision | CV 2.31-21.63% | Good repeatability |
| Inter-batch Precision | CV 17.54% | Acceptable reproducibility |
| Limit of Blank (LoB) | 1.75 copies/μL | Appropriate background signal |
| Limit of Detection (LoD) | 3.33 copies/μL | High sensitivity |
| Limit of Quantification (LoQ) | 13.45 copies/μL | Reliable quantification threshold |
| Linear Range | 13.45-129,693 copies/μL (R²=0.9965) | Wide dynamic range |
Clinical Applications in Cancer Monitoring
The clinical utility of dPCR in liquid biopsy analysis spans the entire cancer care continuum, from early detection to monitoring treatment response. Recent research presented at the AACR Annual Meeting 2025 highlighted several key applications. In colorectal cancer, the VICTORI study demonstrated that ctDNA analysis using dPCR-based methods could detect 94.3% ctDNA positivity in treatment-naive patients and 72.4% in patients with radiologically evident disease who received neoadjuvant therapy [29]. Crucially, 87% of recurrences were preceded by ctDNA positivity, while no ctDNA-negative patient relapsed, highlighting the prognostic value of dPCR-based liquid biopsy monitoring [29].
In bladder cancer, the TOMBOLA trial compared ddPCR and whole-genome sequencing (WGS) for ctDNA detection in 1,282 paired plasma samples, revealing an 82.9% concordance between the methods [29]. ddPCR showed higher sensitivity in low tumor fraction samples, with 12.9% of samples positive only by ddPCR, though both methods demonstrated comparable predictive power for recurrence-free survival [29]. This underscores the particular advantage of dPCR in cases with limited ctDNA shedding.
The ROME trial provided compelling evidence for combining tissue and liquid biopsy approaches, demonstrating that despite only 49% concordance between tissue and liquid biopsies in detecting actionable alterations, combining both modalities significantly increased overall detection of actionable alterations and led to improved survival outcomes in patients receiving tailored therapy [29]. This highlights the complementary nature of dPCR-based liquid biopsy to traditional tissue-based analysis.
Experimental Protocols
Sample Preparation and Nucleic Acid Extraction
Principle: Proper sample handling and nucleic acid extraction are critical for reliable dPCR analysis of liquid biopsies. Blood samples must be processed promptly to prevent degradation of analytes and ensure accurate quantification of rare targets.
Materials:
- Blood collection tubes (EDTA, Streck, or specialized cfDNA tubes)
- Centrifuge capable of 1600-2500 × g
- Plasma separation filters (optional)
- Nucleic acid extraction kits specific for ctDNA or miRNA
- DNase/RNase-free reagents and consumables
- Spectrophotometer or fluorometer for nucleic acid quantification
Procedure:
- Blood Collection and Processing:
- Collect peripheral blood in appropriate collection tubes (typically 10-20 mL).
- Process samples within 2-4 hours of collection to prevent nucleic acid degradation.
- Centrifuge at 1600-2500 × g for 10-20 minutes at 4°C to separate plasma from cellular components.
- Transfer the plasma supernatant to a fresh tube without disturbing the buffy coat.
- Perform a second centrifugation at 16,000 × g for 10 minutes to remove remaining cells and debris.
- Aliquot and store plasma at -80°C if not extracting immediately.
- Nucleic Acid Extraction:
- Extract ctDNA using silica membrane-based columns or magnetic beads optimized for small fragments.
- For miRNA analysis, use specialized kits that preserve small RNA species.
- Elute in an appropriate low-EDTA or EDTA-free buffer to prevent interference with downstream PCR.
- Quantify extract using fluorometric methods specific for dsDNA or RNA.
- Store purified nucleic acids at -80°C for long-term preservation.
Technical Notes:
- Avoid repeated freeze-thaw cycles of both plasma and extracted nucleic acids.
- Include control samples from healthy donors to establish baseline values.
- For ctDNA analysis, assess DNA integrity via fragment analysis if possible.
- Record extraction yield and quality metrics for normalization purposes.
ddPCR Assay for miRNA Quantification in HCC
Principle: This protocol describes the robust quantification of miR-192-5p in plasma samples from hepatocellular carcinoma patients using LNA-enhanced ddPCR, based on the method developed by Liu et al. [27].
Materials:
- ddPCR supermix for probes (no dUTP)
- LNA-enhanced miRNA-specific primers and probes
- Droplet generation oil and cartridges
- DG8 cartridges and gaskets
- Thermal cycler with gradient capability
- Droplet reader compatible with your ddPCR system
- Reverse transcription reagents
Procedure:
- Reverse Transcription:
- Dilute RNA extracts to appropriate concentration in nuclease-free water.
- Prepare reverse transcription reaction with 5 μL template RNA, 2 μL reverse transcriptase, 2.5 μL buffer, 1 μL dNTPs (10 mM), and 1 μL miRNA-specific stem-loop RT primer.
- Incubate at 16°C for 30 min, 42°C for 30 min, 85°C for 5 min, then hold at 4°C.
- Dilute cDNA 5-10 fold with nuclease-free water before ddPCR.
ddPCR Reaction Setup:
- Prepare reaction mix containing 10 μL ddPCR supermix, 1 μM forward primer, 1 μM reverse primer, 300 nM LNA-modified probe, and 5 μL template cDNA.
- Adjust total volume to 20 μL with nuclease-free water.
- Gently mix and briefly centrifuge.
Droplet Generation:
- Transfer 20 μL reaction mix to DG8 cartridge well.
- Add 70 μL droplet generation oil to the appropriate well.
- Place gasket on cartridge and generate droplets in droplet generator.
- Carefully transfer generated droplets to a 96-well PCR plate.
- Seal the plate with a foil heat seal.
PCR Amplification:
- Perform thermal cycling with the following conditions:
- 95°C for 10 min (enzyme activation)
- 45 cycles of:
- 94°C for 30 s (denaturation)
- 55°C for 60 s (annealing/extension)
- 98°C for 10 min (enzyme deactivation)
- 4°C hold
- Use a ramp rate of 2°C/s for all steps.
- Perform thermal cycling with the following conditions:
Droplet Reading and Analysis:
- Place plate in droplet reader for individual droplet analysis.
- Set appropriate fluorescence detection thresholds for positive/negative droplet classification.
- Analyze data using companion software to calculate absolute copies/μL based on Poisson statistics.
Validation Parameters:
- Determine limit of blank (LoB), limit of detection (LoD), and limit of quantification (LoQ) using serial dilutions.
- Assess precision through intra-assay and inter-assay replication.
- Evaluate linearity across expected concentration range (13.45-129,693 copies/μL for miR-192-5p) [27].
- Test interference from common contaminants (hemoglobin, bilirubin, triglycerides).
Multiplex dPCR for Rare Mutation Detection
Principle: This protocol enables simultaneous detection of multiple mutations plus wild-type sequences using intensity-based multiplexing and color combination, based on the RainDrop dPCR system capable of generating 1-10 million picoliter-sized droplets [28].
Materials:
- dPCR master mix suitable for multiplexing
- Mutation-specific primers and probes with different fluorophores
- Wild-type sequence-specific probes
- Microfluidic chip or droplet generator
- Thermal cycler
- Fluorescence imaging system or droplet reader
Procedure:
- Assay Design and Optimization:
- Design primers and probes to target specific mutations (e.g., KRAS, EGFR, BRAF).
- Utilize different fluorophore combinations (FAM, VIC, etc.) for each target.
- Adjust probe concentrations to achieve balanced fluorescence intensities.
- Validate specificity using control templates with known mutation status.
Reaction Setup:
- Prepare master mix containing dPCR supermix, optimized primer and probe concentrations, and template DNA.
- Use 50-100 ng of input DNA per reaction, depending on application.
- Include no-template controls and wild-type-only controls in each run.
Partitioning:
- Load reaction mixture into microfluidic device or droplet generator.
- For RainDrop system, generate 1-10 million picoliter-sized droplets [28].
- Ensure uniform droplet size and distribution for accurate quantification.
Amplification:
- Perform endpoint PCR with optimized thermal cycling conditions.
- Use touchdown protocol if necessary to improve specificity for mutation detection.
- Include a final hold at 4°C until analysis.
Signal Detection and Analysis:
- Analyze droplets using multi-channel fluorescence detection.
- Set thresholds to distinguish positive and negative droplets for each target.
- Apply Poisson correction to calculate absolute copy numbers of each target.
- For rare mutation detection, establish cutoff values based on wild-type background.
Sensitivity Optimization:
- Dilute mutant DNA into wild-type background to establish limit of detection.
- For RainDrop system, demonstrate linear response (R² = 0.998) down to 1 mutant among 250,000 wild-type molecules, with LOD of 1 in 1,000,000 [28].
- Validate with clinical samples of known mutation status.
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for dPCR-Based Liquid Biopsy
| Reagent/Kit | Function | Application Notes |
|---|---|---|
| ctDNA Extraction Kits | Isolation of cell-free DNA from plasma | Optimized for short fragment recovery; minimal contamination from genomic DNA |
| miRNA Extraction Kits | Preservation and isolation of small RNAs | Specifically designed to retain miRNAs <25 nt; includes carrier RNA to improve yield |
| LNA-enhanced Probes | Superior hybridization to miRNA targets | Increases melting temperature and specificity; improves positive droplet counts by 32% [27] |
| dPCR Supermix | Reaction buffer for partitioning | Formulated with high polymerase stability; optimized for droplet or chip-based systems |
| Mutation-specific Assays | Detection of oncogenic mutations | Designed for KRAS, EGFR, BRAF, and other common mutations; validated for multiplexing |
| Reference Assays | Detection of wild-type sequences | Serves as internal control for input quantification and normalization |
| Droplet Generation Oil | Creates stable emulsion for ddPCR | Includes surfactants to prevent coalescence during thermal cycling [1] |
| Quality Control Standards | Assessment of assay performance | Contains known mutation fractions for sensitivity determination and protocol validation |
Digital PCR has established itself as an indispensable technology in the liquid biopsy workflow, providing the sensitivity and precision required for absolute quantification of rare tumor-derived biomarkers in bodily fluids. The applications detailed in these application notes and protocols—from rare mutation detection in hemato-oncology to miRNA quantification in hepatocellular carcinoma—demonstrate the versatility of dPCR across cancer types and biomarker classes. The experimental protocols provide robust methodologies that researchers can implement to advance their oncology research programs, with particular emphasis on standardization and validation to ensure reproducible results. As liquid biopsy continues to transform cancer detection and monitoring, dPCR remains at the forefront, enabling researchers to track tumor dynamics non-invasively with unprecedented precision.
Monitoring Minimal Residual Disease (MRD) and Predicting Early Relapse
Minimal Residual Disease (MRD), also termed Measurable Residual Disease, refers to the small population of cancer cells that persist in a patient after treatment, at levels undetectable by conventional morphological methods [30]. In hematological malignancies, such as leukemia, lymphoma, and multiple myeloma, and increasingly in solid tumors like non-small cell lung cancer (NSCLC), these residual cells are the primary source of clinical relapse [30] [31]. The detection of MRD has emerged as a powerful biomarker, moving from a theoretical research concept to a practice-changing tool that offers a highly sensitive measure of treatment response [32]. Achieving MRD negativity has been strongly correlated with significantly improved Progression-Free Survival (PFS) and Overall Survival (OS) across multiple cancer types, making it a compelling surrogate endpoint in clinical trials and a critical goal for therapy [30] [32].
The clinical value of MRD monitoring lies in its ability to transform patient management. It enables risk stratification, allowing clinicians to identify patients at high risk of relapse who may benefit from treatment intensification or novel therapies, while also sparing those with sustained MRD negativity from unnecessary, toxic treatments [30] [33]. Furthermore, serial MRD assessment provides a dynamic picture of treatment efficacy, enabling earlier adaptation of therapy compared to traditional imaging or clinical examination, which can only detect recurrence once a substantial tumor burden (millions of cells) has accumulated [31]. As novel therapies improve complete response rates, the need for more sensitive measures like MRD to evaluate depth of response and guide post-remission strategies becomes paramount [32] [33].
Current MRD Detection Methods: A Comparative Landscape
A variety of techniques are employed for MRD detection, each with distinct principles, sensitivities, and clinical applications. The choice of method depends on the cancer type, available tissue, genetic landscape, and required sensitivity.
Table 1: Comparison of Common MRD Detection Methods
| Method | Applicability | Sensitivity | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Flow Cytometry (FCM) | ~100% (hematological) [30] | (10^{-3}) – (10^{-6}) [30] | Wide applicability, fast turnaround, relatively inexpensive [30] | Lack of standardization, changes in immunophenotype, requires fresh cells [30] |
| Quantitative PCR (qPCR) | ~40-50% (AML) [30] | (10^{-4}) – (10^{-6}) [30] | Highly sensitive and standardized for specific targets [30] | Only one gene assessed per assay; requires a priori knowledge of mutation [30] |
| Next-Generation Sequencing (NGS) | >95% [30] | (10^{-2}) – (10^{-6}) [30] | Broad applicability, can detect novel mutations, multiple genes analyzed at once [30] | High cost, slow turnaround, complex data analysis, not yet fully standardized [30] |
| Digital PCR (dPCR) | Varies by assay design | Up to 0.001% (up to (10^{-5})) [26] | Absolute quantification without standard curves, high precision, tolerant to inhibitors, rapid turnaround [1] [26] | Limited to a predefined set of mutations, lower multiplexing capability than NGS [1] |
While next-generation sequencing offers broad genomic coverage, digital PCR excels in the precise and absolute quantification of low-frequency variants, making it an ideal tool for longitudinal monitoring of known biomarkers after their initial identification [26]. Its robustness against PCR inhibitors also makes it particularly suitable for analyzing challenging sample types like blood and bone marrow [26].
Digital PCR: Principles and Superiority for MRD Quantification
Digital PCR (dPCR) represents the third generation of PCR technology, following conventional PCR and quantitative real-time PCR (qPCR) [1]. Its fundamental principle is based on the partitioning of a single PCR reaction mixture into thousands to millions of discrete nanoliter-scale reactions. This partitioning results in each compartment containing either zero, one, or a few target nucleic acid molecules, following a Poisson distribution [1]. Following end-point PCR amplification, each partition is analyzed for fluorescence. The fraction of positive partitions is then used to calculate the absolute concentration of the target molecule in the original sample using Poisson statistics, eliminating the need for a standard curve [1] [26].
This compartmentalization provides dPCR with key advantages for MRD detection:
- Absolute Quantification: dPCR provides a direct count of target molecules, removing the reliance on external standards and reducing inter-assay variability [26].
- High Sensitivity and Precision: The ability to detect a single molecule within a partition allows dPCR to reliably quantify rare targets at frequencies as low as 0.001% in a high wild-type background, which is crucial for MRD [26].
- High Tolerance to Inhibitors: Partitioning effectively dilutes PCR inhibitors present in the sample, making dPCR more robust than qPCR when analyzing complex biological samples like blood or bone marrow [26].
- Flexibility in Assay Design: The endpoint detection and compartmentalization make dPCR less sensitive to variations in amplification efficiency, facilitating more flexible multiplexing of variant and wild-type sequences in the same reaction [26].
dot code for generating the dPCR Workflow diagram:
Application Note: dPCR for MRD in Hematological Malignancies
Background and Rationale
In hemato-oncology, specific genetic lesions like the BCR::ABL1 fusion gene in leukemia serve as ideal markers for tracking MRD. Monitoring these targets with high sensitivity is critical for assessing treatment response, guiding therapy changes, and predicting relapse [30] [33]. dPCR has demonstrated clinical utility in reliably quantifying such rare genetic targets down to a frequency of 0.001%, providing a robust tool for longitudinal surveillance [26].
Detailed Experimental Protocol
Objective: To absolutely quantify BCR::ABL1 fusion transcripts and NPM1 Type A mutations in patient blood-derived samples for MRD assessment.
Sample Preparation:
- Source: Collect peripheral blood or bone marrow aspirate in EDTA tubes.
- Nucleic Acid Extraction: Isolve total nucleic acids using a column-based or magnetic bead-based extraction kit. Prefer methods that efficiently remove common PCR inhibitors (e.g., heparin, heme).
- Quality Control: Quantify DNA/RNA using spectrophotometry (e.g., Nanodrop) or fluorometry (e.g., Qubit). For RNA, assess integrity (RNA Integrity Number, RIN) if possible. Convert RNA to cDNA using a reverse transcription kit with random hexamers and/or gene-specific primers.
dPCR Assay Setup:
- Reaction Mix:
- 1X dPCR Master Mix (includes DNA polymerase, dNTPs, buffer).
- Primers: Forward and reverse primers (18-25 bp, Tm 55-65°C, 40-60% GC content) at a final concentration of 400-900 nM each [34].
- Probes: Target-specific probe (e.g., FAM-labeled) and reference gene probe (e.g., VIC-labeled). Probe length should be ~15-30 bp, with a Tm 4-10°C higher than the primers. Avoid G at the 5' end. Use modified bases (LNA, MGB) to increase Tm if needed [34].
- Template: 10-100 ng of cDNA or gDNA per reaction.
- Nuclease-free water to the final volume required by the dPCR instrument.
- Partitioning: Load the reaction mix onto a dPCR plate or cartridge. Use an automated droplet generator or chip-based system (e.g., QIAcuity, QuantStudio Absolute Q) to create ~20,000 partitions per sample.
- PCR Amplification: Run the following thermocycling protocol on a conventional or integrated thermal cycler:
- Initial Denaturation: 95°C for 10 minutes.
- 40-45 Cycles:
- Denature: 95°C for 30 seconds.
- Anneal/Extend: 60°C for 60 seconds (optimize temperature based on assay design).
- Final Hold: 4-10°C (optional 98°C for 10 minutes for enzyme deactivation).
Data Acquisition and Analysis:
- Readout: Place the partitioned sample in a droplet reader or chip imager to measure the fluorescence in each partition (FAM and VIC channels).
- Thresholding: Set fluorescence amplitude thresholds to clearly distinguish positive and negative partitions for each channel. Use no-template controls (NTC) and positive controls to guide threshold setting.
- Concentration Calculation: The software automatically calculates the absolute concentration of the target (copies/μL) and the reference gene using Poisson modeling. The key result for MRD is the mutant allele frequency, expressed as: [ \text{Mutant Allele Frequency (\%)} = \frac{[\text{Target Gene}]}{[\text{Reference Gene}]} \times 100]
- Sensitivity Determination: The limit of detection (LOD) is influenced by the total number of partitions analyzed and the amount of wild-type background. For a 0.001% sensitivity, sufficient input material and partition numbers are critical.
Table 2: Key Research Reagent Solutions for dPCR MRD Assays
| Reagent/Material | Function | Example Specifications |
|---|---|---|
| dPCR Master Mix | Provides optimized buffer, enzymes, and dNTPs for amplification within partitions. | Contains hot-start DNA polymerase, dNTPs, MgCl₂; compatible with probe-based detection. |
| Assay-specific Primers | Flank and define the specific genomic target (e.g., BCR::ABL1 fusion junction) for amplification. | 18-25 bp; Tm 55-65°C; avoid dimers/hairpins; HPLC-purified [34]. |
| Fluorescent Probes | Hybridize to the target sequence within the amplicon, providing a fluorescent signal for detection. | TaqMan-style probes (FAM/VIC dyes with quencher); may include MGB or LNA for specificity [34]. |
| dPCR Plates/Cartridges | Microfluidic devices that generate and hold the nanoliter-scale partitions for the reaction. | Disposable chips or cartridges compatible with the specific dPCR instrument platform. |
| Nucleic Acid Extraction Kit | Isolates high-quality, inhibitor-free DNA/RNA from complex clinical samples (blood, bone marrow). | Column- or bead-based; validated for high yield and removal of PCR inhibitors. |
| Reverse Transcription Kit | Converts RNA to complementary DNA (cDNA) for the detection of fusion transcripts or gene expression. | Includes reverse transcriptase, buffers, dNTPs, and random/oligo-dT primers. |
MRD as a Clinical Endpoint and Future Outlook
The prognostic power of MRD is firmly established. In multiple myeloma, MRD negativity has been shown to be a stronger predictor of PFS and OS than complete remission alone [32]. Large meta-analyses, such as the i2TEAMM and EVIDENCE initiatives, have confirmed that MRD negativity correlates with longer remission and survival, supporting its use as an early clinical endpoint to accelerate drug approvals [32]. This has led to its endorsement by the U.S. FDA's Oncologic Drug Advisory Committee for use in accelerated approval pathways for multiple myeloma [32].
The future of MRD monitoring lies in the integration of technologies. A key development is the distinction between tumor-informed and tumor-naïve (agnostic) approaches for ctDNA-based MRD detection in solid tumors [31]. Tumor-informed assays (e.g., Signatera) use prior sequencing of the patient's tumor to create a custom panel of somatic mutations, offering high sensitivity and specificity. Tumor-naïve assays (e.g., Guardant Reveal) use fixed panels of common cancer mutations, offering faster turnaround and broader applicability but potentially lower sensitivity for individual patients [31]. dPCR can be effectively deployed in both strategies, particularly for tracking specific mutations identified via NGS.
dot code for generating the MRD Clinical Decision Pathway diagram:
Challenges remain, including standardizing the definition of "MRD negativity" across platforms, determining the optimal timing for assessment, and understanding the implications of clonal evolution. Furthermore, the concept of "sustained MRD negativity" is gaining traction as a more robust indicator of durable response [32]. As technologies like dPCR continue to evolve, offering even greater sensitivity and multiplexing capabilities, MRD-guided therapy will become an increasingly integral component of personalized cancer care, enabling earlier intervention and improved survival outcomes.
Detecting Rare Mutations and Copy Number Variations in Complex Samples
The accurate detection of rare mutations and copy number variations (CNVs) is a cornerstone of modern precision oncology, enabling everything from early cancer detection to therapeutic monitoring. Digital PCR (dPCR) has emerged as a powerful third-generation PCR technology that addresses critical limitations of traditional methods like quantitative PCR (qPCR), particularly for analyzing complex biological samples [1]. By providing absolute quantification of nucleic acids without the need for standard curves, dPCR offers unparalleled sensitivity and precision for oncology research applications [1] [13].
This application note details how dPCR methodologies enable researchers and drug development professionals to overcome the challenges of detecting rare genetic events in oncology, with specific protocols for mutation detection and CNV analysis in the context of cancer research.
Digital PCR operates through a simple yet powerful principle: partitioning a single PCR reaction into thousands to millions of parallel nanoreactions so that each contains zero, one, or a few nucleic acid targets according to a Poisson distribution [1]. Following end-point amplification, the fraction of positive partitions is counted, and Poisson statistics are applied to calculate the absolute target concentration [1]. This approach provides several key advantages for oncology research:
- Absolute Quantification: Eliminates the need for standard curves, improving accuracy and reproducibility [1] [13]
- Ultra-Sensitive Detection: Capable of detecting rare mutations and targets present at frequencies as low as 0.01% [13]
- High Precision: Provides precise quantification at ultra-low concentrations, enabling confidence in research conclusions [13]
- Inhibitor Tolerance: Partitioning reduces the effect of PCR inhibitors present in complex sample matrices [35]
Two primary partitioning methods have emerged: water-in-oil droplet emulsification (ddPCR) and microchamber-based systems (nanoplate dPCR) [1]. While both offer single-molecule sensitivity, nanoplate-based systems provide a faster, simpler single-step procedure with full automation of partitioning, thermocycling, and imaging [36].
The following workflow diagram illustrates the core dPCR process for detecting rare mutations in a background of wild-type sequences:
Application 1: Rare Mutation Detection in Liquid Biopsies
The detection of rare somatic mutations in circulating tumor DNA (ctDNA) presents significant challenges due to the low abundance of mutant alleles in a high background of wild-type DNA. dPCR's ability to partition samples enables detection of minor alleles to as little as 0.01% and beyond, making it ideal for liquid biopsy applications [13].
Key Experimental Protocol: Rare Mutation Detection
This protocol adapts methodologies from research on detecting RNA editing events and circulating miRNAs for optimal rare mutation detection in oncology samples [37] [3].
Step 1: Nucleic Acid Extraction
- Extract DNA from plasma, tissue, or formalin-fixed paraffin-embedded (FFPE) samples using validated kits
- For blood samples, collect plasma after double-centrifugation to eliminate cellular DNA contamination
- Quantify DNA using fluorometric methods; input requirements typically range from 1-100 ng depending on application
Step 2: Assay Design and Optimization
- Design TaqMan probe-based assays with mutant-specific probes
- Include a reference assay for normalization (e.g., reference gene or wild-type sequence)
- Optimize primer and probe concentrations through checkerboard titrations
- For multiplexing, ensure minimal spectral overlap between fluorophores
Step 3: Partitioning and Amplification
- Prepare dPCR reaction mixture containing:
- dPCR master mix
- Optimized primer-probe sets
- Template DNA
- Restriction enzyme (if using "One-pot" digestion protocol)
- Load mixture onto appropriate partitioning system (droplet generator or nanoplate)
- Perform PCR amplification with optimized cycling conditions:
- Initial denaturation: 95°C for 10 minutes
- 40-55 cycles of:
- Denaturation: 95°C for 30 seconds
- Annealing/Extension: 55-60°C for 60 seconds
- Final hold: 4-10°C
Step 4: Data Analysis
- Analyze partitions using platform-specific software (e.g., QIAcuity Suite, QuantStudio Absolute Q Software)
- Apply Poisson statistics to calculate absolute copy numbers of mutant and wild-type alleles
- Calculate mutant allele frequency: (mutant copies / total copies) × 100
Results and Data Interpretation
dTable 1: Performance Characteristics of dPCR for Rare Mutation Detection
| Parameter | Performance | Comparison to qPCR |
|---|---|---|
| Sensitivity | Detects minor alleles to 0.01% [13] | 10-100× more sensitive [38] |
| Precision | CV < 10% even at low concentrations [27] | Higher variability, especially near LOD |
| Absolute Quantification | Yes, no standard curve required [1] | Requires standard curve for quantification |
| Input Requirement | 1-100 ng DNA | Similar requirements |
| Inhibitor Tolerance | High - inhibitors affect only subset of partitions [35] | Low - inhibitors affect entire reaction |
Application 2: Copy Number Variation Analysis in Cancer Genomics
CNVs play crucial roles in cancer development, progression, and drug response. Accurate CNV quantification is essential for understanding gene dosage effects on oncogene activation and tumor suppressor inactivation. dPCR provides a robust platform for CNV analysis with superior accuracy compared to qPCR, especially at higher copy numbers [5].
Key Experimental Protocol: CNV Determination
This protocol is adapted from validated methods for CYP2D6 CNV determination and DEFA1A3 copy number analysis, optimized for oncology applications such as HER2 amplification testing in breast cancer [39] [5].
Step 1: Sample Preparation and Restriction Digestion
- Extract high-quality genomic DNA from tumor samples (fresh frozen or FFPE)
- Perform restriction enzyme digestion to fragment genomic DNA:
- Traditional Method: Pre-digest DNA before adding to dPCR reaction
- "One-pot" Method: Include restriction enzyme directly in dPCR reaction mixture [39]
- Select restriction enzymes that do not cut within target or reference amplicons
Step 2: Multiplex Assay Design
- Design assays targeting multiple regions of the gene of interest to detect complex structural variants
- Include a reference gene assay (e.g., RNase P, TERT) in a different optical channel
- Use optical multiplexing (different fluorescent dyes) for clearest signal separation
- Validate assay specificity against related pseudogenes or homologous sequences
Step 3: dPCR Reaction Setup
- Prepare reaction mixture containing:
- dPCR master mix
- Multiplexed target and reference assays (e.g., CYP2D6 5'UTR, intron 6, and exon 9 regions) [39]
- Restriction enzyme (for "One-pot" method)
- Template DNA (10-100 ng)
- Partition reaction using preferred platform (ddPCR or nanoplate-based)
- Run amplification with optimized cycling conditions
Step 4: CNV Calculation and Interpretation
- Calculate target concentration (copies/μL) and reference concentration (copies/μL)
- Determine copy number using the formula:
- CN = (Target concentration / Reference concentration) × 2
- Interpret results based on expected copy number ranges:
- Normal: 2 copies
- deletion: 0-1 copies
- Amplification: >2 copies (cancer-dependent)
The following diagram illustrates the strategic approach to multiplexed CNV detection for complex pharmacogenes like CYP2D6, which can be adapted for cancer-related genes:
Results and Data Interpretation
A recent study comparing dPCR to pulsed-field gel electrophoresis (PFGE, considered a gold standard) demonstrated dPCR's exceptional accuracy for CNV determination [5].
dTable 2: Performance Comparison of CNV Detection Methods
| Method | Concordance with PFGE | Correlation with PFGE | Average Difference from PFGE | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| ddPCR | 95% (38/40 samples) [5] | r = 0.90 (p < 0.0001) [5] | 5% [5] | High-throughput, cost-effective, accurate at high copy numbers | Requires optimization of partitioning |
| qPCR | 60% (24/40 samples) [5] | r = 0.57 (p < 0.0001) [5] | 22% [5] | Widely accessible, familiar technology | Poor accuracy at high copy numbers, requires standard curves |
| PFGE | Gold Standard | Gold Standard | Gold Standard | Highly accurate, detects physical fragments | Low-throughput, technically demanding, requires high-quality DNA [5] |
The superior performance of dPCR is particularly evident at higher copy numbers, where qPCR shows significant underestimation and variability [5]. Linear regression analysis of ddPCR versus PFGE demonstrates nearly perfect 1:1 agreement (Y = 0.9953×), while qPCR shows systematic underestimation (Y = 0.8889×) [5].
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful implementation of dPCR in oncology research requires careful selection of reagents and platforms. The following table summarizes key solutions and their applications:
dTable 3: Essential Research Reagent Solutions for dPCR in Oncology
| Reagent/Platform | Function | Application Notes |
|---|---|---|
| TaqMan Assays (Thermo Fisher) | Sequence-specific detection | Over 20 million predesigned assays available; compatible with dPCR with minimal optimization [13] |
| Absolute Q dPCR System (Thermo Fisher) | Microchamber-based dPCR | Uses microfluidic array plates; integrated workflow [39] |
| QIAcuity dPCR System (Qiagen) | Nanoplate-based dPCR | Fully automated partitioning, thermocycling, and imaging; 26,000 partitions per well [38] [36] |
| QX200 ddPCR System (Bio-Rad) | Droplet-based dPCR | Generates ~20,000 droplets per reaction; established workflow for rare mutation detection [5] [39] |
| Locked Nucleic Acid (LNA) Probes | Enhanced hybridization | Increase positive droplet counts by 32%; improve miRNA detection sensitivity [27] |
| One-pot Restriction Digestion | Simplified sample prep | Incorporates restriction enzyme directly in dPCR mix; streamlines CNV workflow [39] |
Digital PCR represents a transformative technology for detecting rare mutations and copy number variations in complex oncology samples. Its ability to provide absolute quantification without standard curves, combined with superior sensitivity and precision, makes it particularly valuable for liquid biopsy applications, tumor heterogeneity studies, and pharmacogenomics research.
The protocols detailed in this application note provide researchers with robust methodologies for implementing dPCR in their oncology research programs. As the technology continues to evolve with improved multiplexing capabilities and streamlined workflows, dPCR is poised to play an increasingly important role in cancer biomarker discovery and validation, ultimately supporting the development of more personalized cancer therapeutics.
For research use only. Not for use in diagnostic procedures.
Multiplexing Strategies for Multi-Target Analysis from Limited Sample Material
The accurate quantification of multiple biomarkers from minimal sample material is a significant challenge in oncology research, particularly with the increasing reliance on liquid biopsies. Multiplex digital PCR (dPCR) has emerged as a powerful solution, enabling the simultaneous, absolute quantification of several nucleic acid targets from a single, limited-volume sample [1]. This approach partitions a PCR reaction into thousands of nanoliter-scale reactions, allowing for the precise counting of individual target molecules without the need for standard curves [1]. The application of multiplex dPCR is transformative for precision medicine, facilitating the detection of complex cancer signatures, copy number variations, and therapy-resistant mutations from samples such as low-input genomic DNA, cell-free DNA (cfDNA), and serum-derived RNA [40] [12] [41]. This application note details validated protocols and strategies for implementing multiplex dPCR assays, framed within the context of absolute quantification for oncology research.
Key Performance Metrics of Multiplex dPCR Assays
The following tables summarize quantitative performance data from recent peer-reviewed studies utilizing multiplex dPCR for oncology applications. These metrics provide benchmarks for sensitivity, precision, and dynamic range.
Table 1: Analytical Performance of Published Multiplex dPCR Assays in Oncology
| Target / Application | Sample Type | Multiplex Level | Key Performance Metrics | Reference |
|---|---|---|---|---|
| Reference Gene Panel (DCK, HBB, PMM1, RPS27A, RPPH1) | gDNA, cfDNA, Synthetic Fragments | Pentaplex (5-plex) | Expanded uncertainty: 9.2-25.2%; Robust linearity & precision; Wide dynamic range [40] | |
| BTK/PLCG2 Resistance Mutations (e.g., C481S, C481F, R665W) | Patient DNA (CLL) | 3 Multiplex Assays | Detected 68 mutations vs. 49 by NGS; Superior sensitivity at low allelic frequencies [41] | |
| Circulating miRNAs (miR-4488 & miR-579-3p) | Patient Serum | Duplex (2-plex) | Enabled miRatio calculation; Strong prognostic value (ROC analysis); Superior sensitivity vs. qRT-PCR [12] | |
| Actionable Variants (EGFR, KRAS, BRAF) | cfDNA (NSCLC) | Up to 37-plex (single well) | Detection as low as 0.025% variant allele frequency [42] | |
| CAR-T Cell Monitoring | Patient Cellular DNA | Triplex (CAR, T-cell, Control) | Absolute quantitation of CAR-T relative to total T cells; Minimal source sample needs [43] |
Table 2: Comparative Analysis of Multiplex dPCR vs. Alternative Methods
| Parameter | Multiplex dPCR | qPCR | Next-Generation Sequencing (NGS) |
|---|---|---|---|
| Quantification | Absolute, without standard curves [44] | Relative, requires standard curve | Relative or absolute (with complex calibration) |
| Sensitivity | High (e.g., <0.1% VAF) [41] [42] | Moderate | Variable; typically 1-5% VAF [41] |
| Multiplexing Capacity | Moderate to High (2-plex to 37-plex demonstrated) [40] [42] | Low to Moderate | Very High |
| Tolerance to Inhibitors | High [44] | Moderate | Low to Moderate |
| Turnaround Time | Rapid (hours post-extraction) [1] | Rapid | Slow (days to weeks) |
| Cost per Sample | Low to Moderate for targeted analysis | Low | High |
Experimental Protocols
Protocol 1: Pentaplex Reference Gene Quantification for Total DNA Input Normalization
This protocol is designed for accurate quantification of haploid genome equivalents (GE) from limited samples, crucial for normalizing inputs in next-generation sequencing (NGS) or copy number variation (CNV) analysis [40].
1. Reagents and Materials
- Primers and Probes: Predesigned, validated assays for five reference genes (e.g., DCK, HBB, PMM1, RPS27A, RPPH1) located on different chromosomes [40].
- dPCR Supermix: A commercial supermix compatible with your dPCR system and probe chemistry (hydrolysis or universal probes).
- Restriction Enzyme: HindIII or an appropriate alternative for digesting gDNA.
- Nuclease-Free Water and TE Buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0).
- Sample Material: Human gDNA, cfDNA, or synthetic gBlocks.
2. Sample Pre-Processing
- Genomic DNA Digestion: To ensure efficient partitioning and amplification, digest 1 µg of gDNA with 10 units of HindIII restriction enzyme at 37°C for 1 hour [40].
- Sample Dilution: Serially dilute the digested DNA (or cfDNA) in TE buffer to a concentration suitable for dPCR (approximately 500-1000 copies/µL, depending on the partition count). Using a diluent containing carrier DNA (e.g., 4 ng/µL Salmon Sperm DNA) is recommended for low-concentration samples to minimize adsorption losses [40].
3. dPCR Reaction Setup
- Prepare the pentaplex reaction mix on ice. A typical 20 µL reaction volume may contain:
- 10 µL of 2x dPCR Supermix
- 1.0 µL of 20x Pentaplex Primer-Probe Mix (final concentration: 0.9 µM primers, 0.25 µM each probe)
- X µL of Nuclease-Free Water (to bring to volume)
- 4 µL of Template DNA
- Mix thoroughly by pipetting. Do not vortex after adding the supermix.
4. Partitioning and Amplification
- Load the reaction mix into the dPCR partitioning device (e.g., droplet generator or microfluidic chip) according to the manufacturer's instructions.
- Transfer the partitions to a thermal cycler and run the following amplification protocol:
- Enzyme Activation: 95°C for 10 minutes.
- Amplification: 40-45 cycles of:
- Denaturation: 95°C for 30 seconds.
- Annealing/Extension: 60°C for 60 seconds (optimize temperature based on assays).
- Signal Stabilization: 4°C hold (optional, before reading). A 98°C hold for 10-30 minutes may be required for certain universal probe chemistries.
- Endpoint Hold: 12°C indefinitely.
5. Data Analysis
- Analyze the partitioned data using the instrument's software.
- Apply Poisson statistics to calculate the absolute concentration (copies/µL) for each of the five reference genes.
- The total DNA concentration in GE/µL can be determined from the aggregate or average of the five reference gene concentrations, which mitigates bias from potential genomic instability in any single locus [40].
Protocol 2: Duplex dPCR for Circulating miRNA Ratio Analysis in Serum
This protocol details the simultaneous quantification of two circulating miRNAs (e.g., miR-4488 and miR-579-3p) from low-volume serum to calculate a prognostic ratio (miRatio) for monitoring therapy response in metastatic melanoma [12].
1. Reagents and Materials
- RNA Extraction Kit: miRNeasy Mini Kit or equivalent designed for small RNA from serum/plasma.
- Reverse Transcription Kit: TaqMan Advanced miRNA cDNA Synthesis Kit.
- dPCR Supermix: Compatible with hydrolysis probes.
- Duplex miRNA Assay: Fluorescently labelled TaqMan probes for miR-4488 and miR-579-3p, with distinct fluorophores.
2. RNA Extraction and Reverse Transcription
- RNA Extraction: Extract total RNA, including miRNAs, from 200 µL of serum using the miRNeasy Mini Kit according to the manufacturer's instructions. Elute in 20 µL of nuclease-free water [12].
- Reverse Transcription (RT): Due to low RNA concentrations, use a fixed input volume of 2 µL of total RNA for the RT reaction with the TaqMan Advanced miRNA cDNA Synthesis Kit. This kit includes a preamplification step which is critical for enriching low-abundance miRNA targets [12].
3. dPCR Reaction Setup
- Prepare the duplex reaction mix on ice. A typical 20 µL reaction may contain:
- 10 µL of 2x dPCR Supermix
- 1.0 µL of 20x Duplex miRNA Assay
- X µL of Nuclease-Free Water
- 2 µL of undiluted cDNA (from the RT+preamp reaction)
- Mix thoroughly by pipetting.
4. Partitioning and Amplification
- Generate partitions according to your dPCR system's protocol.
- Use the following thermal cycling conditions:
- Enzyme Activation: 95°C for 10 minutes.
- Amplification: 40-45 cycles of:
- Denaturation: 95°C for 30 seconds.
- Annealing/Extension: 60°C for 60 seconds.
- Endpoint Hold: 12°C.
5. Data Analysis
- Use the instrument software to quantify the absolute copies/µL of reaction for miR-4488 and miR-579-3p.
- Calculate the miRatio (e.g., miR-4488 / miR-579-3p or as defined in the biomarker model) [12].
- This ratio can then be used for prognostic stratification and longitudinal monitoring of treatment response.
Signaling Pathways and Workflow Visualizations
Workflow for Multiplex dPCR Analysis from Liquid Biopsy
The following diagram illustrates the integrated workflow for processing liquid biopsy samples, from blood draw to final data analysis, utilizing multiplex dPCR.
Logical Framework for Biomarker Ratio Analysis
This diagram outlines the decision-making logic and clinical significance of calculating a biomarker ratio, such as the miRatio, from duplex dPCR data in metastatic melanoma.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents and Materials for Multiplex dPCR
| Item | Function/Description | Example Use Case |
|---|---|---|
| Multiplex dPCR Supermix | A chemical formulation optimized for partitioning and robust amplification in the presence of multiple primer-probe sets. | Foundation for all multiplex dPCR reactions; ensures compatibility with probe chemistries [40]. |
| Hydrolysis (TaqMan) Probes | Sequence-specific fluorescent probes with a reporter dye and quencher; cleaved during amplification. | Gold-standard for specific detection in multiplex reference gene or mutation panels [40] [41]. |
| Universal Probe Systems (e.g., Rainbow) | Probe chemistry that uses a universal sequence, reducing cost and design complexity for high-level multiplexing. | Validated for pentaplex reference gene panels, performing comparably to hydrolysis probes [40]. |
| Synthetic DNA Fragments (gBlocks) | Double-stranded DNA fragments containing the exact amplicon sequence. | Essential for assay development, optimization, and as quantitative standards without background [40]. |
| Restriction Enzymes (e.g., HindIII) | Enzymes that digest DNA at specific sequences, fragmenting long gDNA. | Critical pre-processing step for gDNA to ensure efficient partitioning and accurate quantification [40]. |
| Carrier DNA (e.g., Salmon Sperm DNA) | Inert DNA used to coat surfaces and prevent adsorption of low-abundance target DNA. | Added to dilution buffers for serially diluting synthetic standards or low-concentration cfDNA [40]. |
| miRNA cDNA Synthesis Kit | Specialized kits for reverse transcription and preamplification of miRNA templates. | Required for profiling low-abundance circulating miRNAs from serum/plasma [12]. |
Optimizing dPCR Assays: Practical Strategies for Enhanced Sensitivity and Robust Performance
Digital PCR (dPCR) represents the third generation of PCR technology, enabling absolute quantification of nucleic acids without the need for standard curves. This calibration-free technology provides powerful advantages including high sensitivity, absolute quantification, and high accuracy and reproducibility, making it particularly valuable in oncology research [45]. In dPCR, a PCR mixture containing the sample is partitioned into thousands to millions of compartments, so that each partition contains either 0, 1, or a few nucleic acid targets according to a Poisson distribution [1]. Following PCR amplification, the fraction of positive partitions is measured via end-point fluorescence detection, allowing precise computation of the target concentration [45].
The application of dPCR in oncology has revolutionized approaches to cancer biomarker analysis, particularly through liquid biopsy applications such as circulating tumor DNA (ctDNA) analysis [46]. dPCR's ability to detect rare genetic mutations within a background of wild-type genes enables researchers to monitor tumor heterogeneity and treatment response with unprecedented sensitivity [45]. This breakthrough has paved the way for non-invasive cancer monitoring, with dPCR demonstrating 100-times greater sensitivity than conventional methods for rare mutation detection [4]. The technology's precision in absolute quantification also makes it invaluable for copy number variation (CNV) analysis in cancer genomics, providing accurate resolution of CNVs at both low and high DNA copy numbers [5].
Primer Design Optimization
Fundamental Principles of PCR Primer Design
Proper primer design is paramount for successful dPCR assays in oncology research, where accurate detection of low-frequency mutations is often critical. Optimal primer design ensures specific amplification of target sequences while minimizing non-specific binding and primer-dimer formation [47]. For dPCR applications, primers must be designed to work efficiently with the partitioned reaction environment and provide robust amplification even at the single-molecule level.
IDT scientists recommend designing PCR primers between 18 and 30 bases in length, with the most important considerations being the Tm value and on-target binding efficiency [47]. The guidelines for primer design are summarized in Table 1.
Table 1: Optimal Parameters for PCR Primer Design
| Parameter | Recommended Range | Ideal Value | Rationale |
|---|---|---|---|
| Length | 18-30 bases | - | Balances specificity and binding efficiency |
| Melting Temperature (Tm) | 60-64°C | 62°C | Compatible with standard cycling conditions and enzyme function |
| Tm Difference Between Primers | ≤2°C | 0°C | Ensures simultaneous binding of both primers |
| GC Content | 35-65% | 50% | Provides sequence complexity while maintaining uniqueness |
| Consecutive G Residues | Avoid ≥4 | None | Prevents formation of complex secondary structures |
Advanced Design Considerations for Oncology Applications
In oncology research, primer design must address additional challenges such as distinguishing single-nucleotide variants (SNVs) and amplifying from limited sample material. When designing primers for mutation detection, the primer should be positioned such that the 3' end is near or at the variant nucleotide to maximize discrimination between wild-type and mutant sequences [47]. For copy number variation studies, primers should target unique genomic regions with minimal homology to other sequences to ensure accurate quantification.
Complementarity and secondary structure screening is essential for robust dPCR assays. Primer designs should be analyzed for self-dimers, heterodimers, and hairpins using tools such as the OligoAnalyzer Tool [47]. The ΔG value of any secondary structures should be weaker (more positive) than -9.0 kcal/mol to prevent interference with amplification efficiency [47]. Furthermore, on-target binding efficiency should be verified using NCBI BLAST alignment to ensure primer uniqueness to the desired target sequence, which is particularly important when working with highly homologous gene families or pseudogenes frequently encountered in cancer genomics [47].
Practical Implementation and Verification
The implementation of optimized primer designs requires careful experimental setup. Amplicon length should typically be maintained between 70-150 bp to allow for efficient amplification under standard cycling conditions, though longer amplicons up to 500 bases can be generated with modified cycling parameters [47]. When analyzing gene expression or DNA biomarkers from liquid biopsies, it is good practice to treat RNA samples with DNase I to remove residual genomic DNA, and design assays to span exon-exon junctions where possible to reduce gDNA amplification [47].
To verify primer performance, a series of validation experiments should be conducted including:
- Specificity testing using gel electrophoresis or capillary electrophoresis to confirm single amplicon formation
- Efficiency determination through standard curve analysis or comparative dPCR measurements
- Sensitivity assessment using limiting dilution series to establish the limit of detection
- Cross-reactivity screening against related gene sequences or pseudogenes
Figure 1: Workflow for optimal primer design and verification in dPCR assays
Probe Selection Strategies
Hydrolysis Probe Design Considerations
Hydrolysis probes (such as TaqMan probes) are fundamental to many dPCR applications in oncology, particularly for multiplex assays and mutation detection. Proper probe design ensures efficient fluorescence signal generation and accurate discrimination between closely related sequences. IDT recommends using double-quenched probes with internal quenchers such as ZEN or TAO molecules, as they provide consistently lower background and higher signal-to-noise ratios compared to single-quenched probes [47]. This enhanced quenching is especially valuable in dPCR where precise classification of positive and negative partitions is critical.
For single-quenched probes, length should be maintained between 20-30 bases to achieve suitable Tm without compromising the efficiency of fluorescence quenching [47]. The positioning of the probe is crucial—it should be in close proximity to the forward or reverse primer but should not overlap with the primer-binding site on the same strand [47]. Probes can be designed to bind to either strand of the target DNA, providing flexibility when designing assays for complex genomic regions.
Table 2: Guidelines for dPCR Probe Design and Configuration
| Parameter | Recommendation | Impact on Assay Performance |
|---|---|---|
| Probe Type | Double-quenched (with ZEN/TAO) | Lower background, higher signal-to-noise for better partition classification |
| Length | 20-30 bases (single-quenched) | Maintains optimal distance between fluorophore and quencher |
| Location | Close to primers but non-overlapping | Ensures efficient hybridization during amplification |
| Tm Relative to Primers | 5-10°C higher | Ensures probe binding before primer extension |
| GC Content | 35-65% | Similar to primer requirements for consistent hybridization |
| 5' Base | Avoid G | Prevents quenching of fluorophore |
Oncology-Specific Probe Applications
In oncology research, probe design must address the challenge of detecting rare mutations against a background of wild-type sequences. For single-nucleotide variant (SNV) detection, competitive probe setups using different fluorophores for wild-type and mutant sequences enable precise quantification of variant allele frequencies [4]. This approach is particularly powerful in liquid biopsy applications where ctDNA may represent less than 0.1% of total cell-free DNA.
The melting temperature (Tm) of probes is especially critical in dPCR applications. Probes should have a Tm 5-10°C higher than the primers to ensure they are fully hybridized during the primer extension phase [47]. If the probe Tm is too low, the percentage of probe bound to target will be reduced, compromising quantitative accuracy as not all target molecules will be detected, leading to underestimation of target concentration [47]. The annealing temperature (Ta) should be set no more than 5°C below the lower primer Tm, though further optimization may be necessary based on empirical testing [47].
Multiplexing Strategies for Comprehensive Analysis
Multiplex dPCR assays enable simultaneous quantification of multiple targets, which is invaluable in oncology for assessing treatment response, tumor heterogeneity, and resistance mechanisms. Modern dPCR systems such as the QuantStudio Absolute Q Digital PCR System support multiplexing with up to five fluorescence channels (four for targets and one for reference/QC) [45]. When designing multiplex assays, careful attention must be paid to spectral cross-talk between fluorophores and compatible Tm values across all probes.
For complex applications such as comprehensive mutation profiling or copy number variation analysis, reference probes are essential for data normalization. Reference genes should be carefully selected to exhibit stable copy numbers in the biological system being studied, and should be amplified with similar efficiency to the target sequences. This approach was successfully employed in a recent study of the DEFA1A3 gene, where ddPCR demonstrated 95% concordance with pulsed-field gel electrophoresis (the gold standard) for copy number determination, significantly outperforming qPCR which showed only 60% concordance [5].
Master Mix Composition
Critical Components of dPCR Master Mix
The master mix formulation directly impacts the efficiency, specificity, and reliability of dPCR assays. While specific commercial master mix compositions are proprietary, they generally contain several key components optimized for digital PCR applications. The master mix typically includes a thermostable DNA polymerase with proven processivity and fidelity, deoxynucleotide triphosphates (dNTPs) at optimized concentrations, MgCl₂ or MgSO₄ as a cofactor, and reaction buffers that maintain optimal pH and ionic strength throughout the thermal cycling process [48].
The concentration of magnesium ions is particularly crucial, as it affects primer annealing, enzyme processivity, and amplicon specificity. Standard reaction conditions often include approximately 3 mM Mg²⁺, though this should be optimized for specific assays [47]. Similarly, potassium ion concentration is typically maintained around 50 mM to support enzymatic activity while maintaining nucleic acid stability. When calculating melting temperatures using online tools such as the IDT OligoAnalyzer Tool, it is essential to input the specific reaction conditions from the dPCR experiment, as Tm values are highly dependent on these parameters [47].
Selection Criteria for Oncology Applications
Selection of the appropriate master mix for oncology research depends on several factors including the sample type, target abundance, and specific application. For liquid biopsy applications where detecting rare mutations is critical, master mixes with enhanced sensitivity and robust performance at low template concentrations are essential. For gene editing validation or CRISPR-Cas9 efficiency quantification, master mixes that provide consistent amplification across a wide dynamic range are preferable [4].
When working with challenging sample types common in oncology research, such as formalin-fixed paraffin-embedded (FFPE) tissues or cell-free DNA from plasma, master mixes formulated with inhibitor-resistant polymerases and enhanced stabilization compounds can significantly improve assay performance. These specialized formulations help overcome PCR inhibitors commonly found in clinical samples while maintaining accurate quantification of nucleic acid targets.
Optimization Strategies for Specific Workflows
Master mix optimization should be tailored to specific dPCR platforms and partitioning technologies. For droplet-based dPCR systems, master mixes must generate stable emulsions throughout the thermal cycling process, often requiring specific surfactants or stabilizers [45]. In contrast, chip-based systems such as the QIAcuity Digital PCR System utilize nanoplates with fixed partitions, where master mix viscosity and surface tension properties must be compatible with microfluidic loading [49].
The modular automation setup available in systems like the QuantStudio Absolute Q Digital PCR System enables flexible high-throughput workflows, but requires consistent master mix performance across multiple runs [4]. For such applications, pre-formulated master mixes that include all necessary components (except primers, probes, and template) provide the most reproducible results, minimizing pipetting errors and variation between reactions. These master mixes are available in different concentrations (2X, 4X, etc.) to accommodate various reaction setup preferences and sample volumes [48].
Integrated Experimental Protocols
Comprehensive dPCR Workflow for Absolute Quantification
A standardized dPCR protocol ensures consistent and reproducible results across experiments, which is particularly important in longitudinal oncology studies monitoring disease progression or treatment response. The general dPCR workflow follows four key steps: (1) partitioning of the PCR mixture containing the sample into thousands of individual reactions; (2) amplification of target sequences within each partition; (3) end-point fluorescence analysis of each partition; and (4) data analysis using Poisson statistics to determine absolute target concentration [45] [1].
The complete workflow from sample preparation to results can be completed in less than 2 hours on integrated systems such as the QIAcuity Digital PCR System [49]. Sample preparation follows similar protocols to qPCR, involving transfer of mastermix, probes, primers, and samples to reaction vessels. The partitioning method varies by platform—droplet-based systems generate water-in-oil emulsions, while chip-based systems use microchambers or nanoplates [45]. Following thermal cycling, the fraction of positive partitions is determined by fluorescence thresholding, and the target concentration is calculated using Poisson statistics to account for the random distribution of molecules among partitions.
Figure 2: Comprehensive dPCR workflow for absolute quantification of nucleic acids
Validation Protocol for dPCR Assays in Oncology Research
Robust validation of dPCR assays is essential for generating reliable data in oncology research. The following protocol outlines key steps for assay validation:
Linearity and Dynamic Range Assessment:
- Prepare a dilution series of reference material spanning the expected concentration range
- Analyze each dilution in triplicate using the dPCR assay
- Evaluate linearity by plotting expected versus measured concentrations
- Acceptable assays should demonstrate R² > 0.98 across the working range
Limit of Detection (LOD) and Limit of Quantification (LOQ) Determination:
- Analyze multiple replicates of negative template controls to establish background
- Test dilutions near the expected detection limit (typically 3-5 copies/reaction)
- Calculate LOD and LOQ based on statistical analysis of positive calls
- For rare mutation detection, establish the minimum variant allele frequency detectable
Precision Evaluation:
- Analyze samples with low, medium, and high target concentrations across multiple runs
- Calculate intra-assay and inter-assay coefficients of variation (CV)
- Acceptable precision typically demonstrates CV < 10% for most dPCR applications
Specificity Verification:
- Test assay performance against closely related non-target sequences
- Verify discrimination efficiency for single-nucleotide variants
- Confirm absence of cross-reactivity in multiplex assays
This validation approach aligns with the dMIQE (Minimum Information for Publication of Quantitative Digital PCR Experiments) guidelines, which provide a framework for ensuring reproducibility and reliability of dPCR results [46].
Protocol for Rare Mutation Detection in Liquid Biopsies
The detection of rare mutations in cell-free DNA requires optimized protocols to achieve the necessary sensitivity and specificity. The following protocol has been demonstrated effective for detecting mutations present at frequencies as low as 0.1%:
Sample Preparation:
- Extract cell-free DNA from 2-10 mL plasma using validated kits
- Elute DNA in low TE buffer or nuclease-free water
- Quantify using fluorescence-based methods suitable for low-concentration samples
Assay Setup:
- Use mutation-specific probes with different fluorophores for wild-type and mutant sequences
- Include blocking primers to suppress wild-type amplification if necessary
- Set up 20-40 μL reactions according to master mix manufacturer's instructions
- Include no-template controls and positive controls at known variant allele frequencies
Partitioning and Amplification:
- Generate partitions according to platform-specific protocols
- Amplify with thermal cycling conditions optimized for the assay
- Typically: 95°C for 10 min, 40-45 cycles of (95°C for 15s, 60°C for 60s)
Data Analysis:
- Set fluorescence thresholds using positive and negative controls
- Apply Poisson correction to calculate mutant and wild-type concentrations
- Calculate variant allele frequency as (mutant copies)/(mutant + wild-type copies)
- Apply confidence intervals based on binomial statistics
This approach leverages dPCR's ability to detect rare mutations with 100-times greater sensitivity than conventional methods, enabling non-invasive monitoring of tumor dynamics through liquid biopsies [4].
Research Reagent Solutions
Table 3: Essential Research Reagents for dPCR in Oncology Applications
| Reagent Category | Specific Products/Systems | Key Features | Oncology Research Applications |
|---|---|---|---|
| dPCR Systems | QuantStudio Absolute Q Digital PCR System | Microfluidic array plate (MAP) technology, 20,480 partitions per sample, 5-color detection | Absolute quantification of cancer biomarkers, rare mutation detection, liquid biopsy analysis |
| dPCR Systems | QIAcuity Digital PCR System | Nanoplate-based, fully integrated partitioning, thermal cycling, and imaging | Gene expression profiling in tumor samples, copy number variation analysis |
| Master Mixes | TaqMan dPCR Master Mixes | Optimized for digital PCR applications, compatible with various probe chemistries | Sensitive detection of low-abundance targets in liquid biopsies |
| Assays & Probes | TaqMan Assay Portfolio | Double-quenched probes, validated sequences, various fluorophore options | Multiplexed detection of cancer-associated mutations, fusion genes |
| Design Tools | IDT PrimerQuest Tool, OligoAnalyzer Tool | Automated primer and probe design, Tm calculation, secondary structure analysis | Custom assay development for novel cancer biomarkers |
| Consumables | MAP16 Digital PCR Plates, QIAcuity Nanoplates | Platform-specific partitioning devices | High-throughput screening of therapeutic targets |
Optimization of critical reagents—primers, probes, and master mix composition—is fundamental to successful implementation of digital PCR in oncology research. The precise absolute quantification capabilities of dPCR technology provide researchers with powerful tools for cancer biomarker analysis, liquid biopsy applications, and treatment response monitoring. By adhering to established design principles and validation protocols, researchers can develop robust dPCR assays that leverage the technology's superior sensitivity and precision for advancing cancer research and therapeutic development.
The future of dPCR in oncology looks promising, with emerging applications in cell and gene therapy quality control, vaccine development, and comprehensive cancer monitoring [46]. As the technology continues to evolve with improvements in multiplexing capabilities, workflow simplicity, and data analysis tools, dPCR is poised to become an increasingly indispensable tool in the oncologist's research arsenal, enabling more precise molecular characterization of tumors and more sensitive monitoring of disease progression.
In the context of oncology research, digital PCR (dPCR) has emerged as a powerful tool for the absolute quantification of rare oncogenic mutations, assessment of copy number variations, and monitoring of treatment response through liquid biopsy [1]. The core principle of dPCR involves partitioning a PCR mixture into thousands to millions of individual reactions so that a single partition contains zero, one, or a few nucleic acid targets [1] [50]. The precision of this absolute quantification critically depends on the quality of partitioning, specifically the achievement of monodisperse droplets (uniform in size) and the prevention of their coalescence (merging) [51] [1]. This application note details the primary challenges and proven solutions for ensuring robust partitioning in dPCR workflows, with a focus on applications in oncology.
Key Partitioning Challenges and Their Impact on dPCR Performance
The integrity of dPCR data is fundamentally linked to the physical properties of the partitions. Two major issues can compromise results:
- Non-Monodispersity: Variations in partition volume lead to unequal distribution of target molecules, violating the Poisson distribution assumption used for absolute quantification. This introduces errors in the calculated target concentration and reduces the assay's precision [51].
- Droplet Coalescence: The merging of partitions after formation causes data loss and can create partitions with multiple targets that are misinterpreted during fluorescence reading, leading to an underestimation of the true target concentration [1]. This is particularly critical when detecting low-abundance mutations in a background of wild-type sequences, a common scenario in oncology.
Strategies for Ensuring Monodisperse Droplet Generation
Monodisperse droplet generation is achievable through both passive and active microfluidic strategies. The choice of method involves a trade-off between simplicity, control, and throughput.
Passive Microfluidic Geometries
Passive methods rely on the channel geometry to hydrodynamically focus and break the dispersed phase into droplets.
- Step Emulsion: This design utilizes a sudden change in channel geometry (a "step") to trigger droplet breakup. It is known for its simplicity and ability to produce highly uniform droplets [51].
- Flow-Focusing Geometries: In this common design, the dispersed (aqueous) phase is focused by two continuous (oil) phases, leading to droplet pinch-off at a narrow orifice. It allows for good control over droplet size based on flow rate ratios [51].
Active Microfluidic Actuation
Active methods introduce external energy to achieve more dynamic control over the droplet generation process.
- Surface Acoustic Wave (SAW) Induced Generation: This method uses high-frequency sound waves generated by interdigitated transducers on a piezoelectric substrate. The acoustic radiation force destabilizes the liquid interface, leading to rapid and controlled droplet formation. A key advantage is the ability to tune droplet size and production rate by adjusting the acoustic power and frequency without altering fluid flow rates [51]. Recent advancements have demonstrated the SAW-induced step-emulsion (SISE) system, which can generate droplets at rates up to 8.7 kHz with a coefficient of variation (CV) for size below 5%, a benchmark for monodispersity [51].
Table 1: Comparison of Droplet Generation Methods for dPCR
| Method | Principle | Key Advantage | Typical Droplet Uniformity (CV) | Throughput |
|---|---|---|---|---|
| Step Emulsion [51] | Passive, geometric breakup | Simple fabrication, high uniformity | < 5% | High |
| Flow-Focusing [51] | Passive, hydrodynamic focusing | Well-established, good size control | ~5-10% | High |
| SAW-Induced Step Emulsion [51] | Active, acoustic force | On-demand control, tunable size/rate | < 5% | Very High (up to 8.7 kHz) |
Protocols for Preventing Droplet Coalescence
Preventing coalescence is paramount for maintaining partition integrity through thermocycling and analysis. The following protocols are critical.
Surfactant Formulation and Optimization
The use of appropriate surfactants in the continuous oil phase is the primary method for stabilizing droplets [1] [52].
- Protocol: Surfactant Screening and Concentration Optimization
- Objective: To identify the optimal surfactant and its concentration for stabilizing droplets during dPCR thermocycling.
- Materials:
- Continuous phase oil (e.g., fluorinated oil)
- Candidate surfactants (e.g., Krytox-based surfactants for fluorinated oils, polyethylene glycol (PEG)/polyoxypropylene glycol dimethicone for silicone oils [52])
- Aqueous phase (PCR mix)
- Droplet generator
- Method:
- Prepare the continuous phase with surfactant concentrations typically ranging from 0.1% to 5.0% (w/w) [52].
- Generate droplets using a standard monodisperse method.
- Incubate the collected emulsion at the highest temperature of the dPCR thermocycling protocol (e.g., 95°C) for 1-2 hours.
- After incubation, analyze the emulsion under a microscope for signs of coalescence (fewer, larger droplets) or degradation.
- Validation: The optimal formulation will show no visible coalescence and maintain a stable droplet count post-incubation.
Device Design and Surface Chemistry
The physical and chemical properties of the microfluidic device itself can influence droplet stability.
- Protocol: Device Surface Passivation
- Objective: To render the microfluidic channel surfaces hydrophobic (for water-in-oil droplets) to prevent aqueous phase adhesion and wetting, which can lead to coalescence.
- Materials: PDMS or glass microfluidic chips, trichloro(1H,1H,2H,2H-perfluorooctyl)silane, vacuum desiccator.
- Method:
- Place the microfluidic device in a vacuum desiccator alongside a small volume (50-100 µL) of the silane solution.
- Evacuate the desiccator for at least 1 hour to allow silane vapor to coat the internal surfaces of the channels.
- Flush the channels with an appropriate solvent (e.g., HFE-7500) and then with air to remove any unbound silane.
- Validation: Successful passivation is indicated by the aqueous phase readily forming beads and moving freely through the channels without wetting the surfaces.
The Scientist's Toolkit: Essential Reagents and Materials
Table 2: Key Research Reagent Solutions for dPCR Partitioning
| Item | Function | Application Note |
|---|---|---|
| Fluorinated Oil | Continuous phase; immiscible with aqueous PCR mix. | Provides a chemically inert environment; low gas permeability helps prevent droplet evaporation during thermocycling. |
| Krytox-PEG Surfactant | Prevents droplet coalescence by reducing interfacial tension. | Essential for long-term droplet stability at high temperatures; concentration must be optimized. |
| Surface Passivation Reagent (e.g., Fluorosilane) | Modifies microfluidic channel surface to be hydrophobic. | Prevents aqueous phase from adhering to channel walls, ensuring smooth droplet flow and reducing coalescence risk. |
| Photoinitiator (e.g., Darocur 1173) | Initiates polymerization upon UV exposure. | Used in protocols involving polymer-based particles or droplet gelation [52]; not for standard dPCR. |
| Polyvinyl Alcohol (PVA) | Stabilizer in the external aqueous phase for complex emulsion formation. | Used in advanced microfluidic particle synthesis [52]; can be adapted for stabilizing droplet collection surfaces. |
Robust partitioning is the foundation of reliable and precise digital PCR data. For oncology researchers aiming to quantify rare cancer biomarkers, implementing strategies that ensure both monodispersity and droplet stability is non-negotiable. The integration of advanced microfluidic actuation like SAW for controlled generation, combined with optimized surfactant chemistry, provides a reliable path to overcoming partitioning issues, thereby enhancing the accuracy and diagnostic power of dPCR in cancer research.
Optimizing Thermal Cycling Conditions for Efficient Target Amplification
In the field of oncology research, the demand for precise, sensitive, and reproducible molecular techniques is paramount. Digital PCR (dPCR) has emerged as a powerful tool for the absolute quantification of nucleic acids, enabling the detection of rare genetic mutations, monitoring of minimal residual disease, and analysis of tumor heterogeneity through liquid biopsy [1]. The core principle of dPCR involves partitioning a PCR reaction into thousands to millions of individual reactions, so that a single partition contains either zero, one, or a few target molecules [1]. Following end-point amplification, the fraction of positive partitions is used to calculate the absolute target concentration via Poisson statistics, eliminating the need for a standard curve [1] [53].
The efficacy of this absolute quantification is fundamentally dependent on the efficiency of the PCR amplification within each partition. Optimal thermal cycling conditions are therefore not merely a prerequisite but a critical variable that directly influences the sensitivity, accuracy, and reliability of dPCR assays, especially when quantifying low-abundance oncogenic mutations against a high background of wild-type DNA [54]. This application note provides a detailed protocol for optimizing thermal cycling parameters to achieve robust and efficient target amplification in dPCR for oncology research.
Critical Thermal Cycling Parameters for dPCR Optimization
The transition from conventional or quantitative PCR (qPCR) to dPCR introduces unique considerations for thermal cycling optimization. The partitioning of the reaction mix into nanoliter-sized droplets or microchambers can alter reaction kinetics and efficiency [1]. The following parameters require systematic investigation.
Table 1: Key Thermal Cycling Parameters for Optimization in dPCR
| Parameter | Description | Impact on dPCR Assay Performance | Recommended Optimization Range |
|---|---|---|---|
| Annealing Temperature | Temperature at which primers bind to the template DNA. | Critically affects specificity; suboptimal temperatures cause non-specific amplification or primer-dimer formation, increasing false positives [55]. | Gradient from 5°C below to 5°C above the calculated ( T_m ). |
| Annealing/Hold Time | Duration of the annealing step per cycle. | Insufficient time can lead to incomplete primer binding, reducing amplification efficiency and yield [55]. | 15–60 seconds. |
| Ramp Rate | Speed of temperature transitions between steps. | Faster rates can improve specificity and reduce overall cycle time, but the optimal rate may depend on the dPCR instrument's capabilities [55]. | Standard vs. Maximum instrument rate. |
| Number of Cycles | Total cycles of denaturation, annealing, and extension. | Insufficient cycles prevent low-abundance targets from reaching detection threshold; excessive cycles can increase background noise [53]. | 40–50 cycles. |
| Denaturation & Extension | Temperature and duration for strand separation and polymerase activity. | Standard conditions often suffice, but should be verified for complex genomic regions or long amplicons [55]. | Denaturation: 95–98°C for 5–30s; Extension: 72°C or polymerase-specific. |
The Role of the Thermal Cycler
The thermal cycler itself is a key component in assay development. Its performance directly impacts the consistency of amplification across all partitions [55]. Key instrumental metrics include:
- Temperature Uniformity: The maximum temperature variance across the thermal block. Poor uniformity can lead to different amplification efficiencies in different wells, compromising data reproducibility [55].
- Temperature Accuracy: How closely the actual block temperature matches the programmed setpoint. High accuracy (typically ±0.25°C or better) is required for reliable and consistent results [55].
- Ramp Rate: The speed of temperature transitions. Faster ramp rates can reduce overall run time and may improve specificity by limiting the time reactions spend at suboptimal temperatures [55].
Experimental Protocol: A Step-by-Step Guide to Optimization
This protocol uses a gradient thermal cycler to establish the optimal annealing temperature for a dPCR assay designed to detect a specific oncogenic point mutation (e.g., KRAS G12D).
Materials and Equipment
Table 2: Research Reagent Solutions and Essential Materials
| Item | Function/Description | Example/Catalog Note |
|---|---|---|
| dPCR System | Partitions the sample, performs amplification, and analyzes endpoints. | Bio-Rad QX200 Droplet Digital PCR System [56] or chip-based systems (e.g., Qiagen QIAcuity) [1]. |
| Gradient Thermal Cycler | Allows testing of multiple annealing temperatures in a single run. | Instrument with verified temperature uniformity and gradient functionality is critical [55]. |
| dPCR Supermix | Optimized reaction mix containing DNA polymerase, dNTPs, and buffer. | Use a supermix compatible with the dPCR platform and probe chemistry (e.g., hydrolysis probes). |
| Target-specific Primers & Probes | Oligonucleotides for specific amplification and detection of the target. | Hydrolysis probes (e.g., FAM-labeled) for mutant allele; a second channel (e.g., HEX) can be used for a reference gene [54]. |
| Template DNA | Sample containing the nucleic acid target. | Use well-characterized genomic DNA from cell lines or patient-derived samples with known mutation status for validation. |
| Droplet Generator Oil | Creates stable water-in-oil emulsion for droplet-based dPCR. | Specific to the droplet generation system (e.g., DG8 Cartridges for QX200) [1]. |
Procedure
- Assay Design and Preparation: Design and obtain hydrolysis probes and primers specific for the oncogenic mutation. A reference assay (e.g., for a wild-type sequence or a housekeeping gene) should be included for normalization and quality control [54].
- Reaction Setup: Prepare the dPCR master mix according to the manufacturer's instructions, containing supermix, primers, probes, and nuclease-free water. Aliquot a precise volume of the master mix (e.g., 20 µL for the QX200 system) and add the template DNA. A no-template control (NTC) must be included to confirm the absence of contamination.
- Partitioning: Transfer the reaction mix to the droplet generator. For droplet-based systems, this will create an emulsion of approximately 20,000 nanoliter-sized droplets [56]. For chip-based systems, load the mixture into the designated nanowell chip [1].
- Gradient PCR Amplification: Transfer the partitions (droplets or chip) to the thermal cycler. Program a gradient protocol:
- Initial Denaturation: 95°C for 10 minutes.
- 40–50 Cycles:
- Denaturation: 94°C for 30 seconds.
- Annealing: Gradient from 55°C to 65°C for 60 seconds.
- Extension: 72°C for 60 seconds.
- Final Hold: 98°C for 10 minutes (enzyme deactivation) and 4°C hold.
- Endpoint Reading and Analysis: After amplification, place the plate or chip into the droplet reader or imager. The instrument will count the positive and negative partitions for each channel. Using the instrument's software, analyze the data from each annealing temperature well separately.
Data Analysis and Interpretation
- Determine Optimal Annealing Temperature: The optimal temperature is identified by the well that yields the highest number of positive partitions (counts) for the target without any signal in the NTC. This point represents the best combination of high amplification efficiency and specificity.
- Assess Assay Performance: Calculate the concentration (copies/µL) of the target and reference gene. Evaluate the data for clear separation between positive and negative droplet clusters; a wide spread or intermediate clusters at a given temperature indicates poor specificity.
- Validate with Biological Samples: Once optimal conditions are established, validate the assay using patient samples or other relevant biological materials, as demonstrated in studies monitoring treatment response in oncology [1] or quantifying antibiotic resistance genes in complex backgrounds [56].
Workflow and Logical Relationships
The following diagram illustrates the complete workflow for developing and optimizing a dPCR assay, highlighting the central role of thermal cycling optimization.
Diagram 1: dPCR assay optimization workflow.
The iterative process of analyzing data and refining parameters is crucial. The application of machine learning for image analysis in chip-based dPCR further underscores the importance of high-quality, specific amplification data as the foundational input [53]. Proper optimization ensures that the final validated protocol is robust, sensitive, and fit-for-purpose in a clinical research setting.
Mitigating Inhibition and Handling Low-Quality/Quantity Input Samples
In the field of oncology research, the absolute quantification of nucleic acids via digital PCR (dPCR) is frequently challenged by two major constraints: the presence of PCR inhibitors in sample matrices and the limited quantity or quality of input nucleic acids obtained from precious clinical samples. dPCR operates by partitioning a sample into thousands to millions of individual reactions, enabling absolute quantification of target sequences through Poisson statistical analysis of positive and negative partitions [1] [57]. This partitioning confers certain advantages for challenging samples, as it efficiently concentrates target sequences within isolated microreactors, reducing template competition and potentially allowing higher tolerance to inhibitors present in a sample [57]. For oncology applications involving liquid biopsies, fine-needle aspirates, or formalin-fixed paraffin-embedded (FFPE) tissues, these challenges are particularly acute, where obtaining sufficient high-quality DNA or RNA remains a significant bottleneck in molecular profiling [58]. This application note provides detailed protocols and data-driven strategies to optimize dPCR performance under these constrained conditions, specifically framed within oncology research requirements.
Understanding Inhibition Mechanisms in dPCR
PCR inhibitors originate from diverse sources encountered during sample collection, extraction, and processing. In clinical oncology, common inhibitors include heparin from blood collection tubes, hemoglobin from hemolyzed samples, lipids from plasma, melanin from melanoma tissues, and formalin-induced modifications from FFPE processing [59]. Environmental samples may contain humic acids, tannins, and heavy metals, while plant and soil-derived inhibitors include polysaccharides, polyphenolics, and humic substances [59]. These compounds interfere with amplification through multiple mechanisms: sequestration of essential cofactors (e.g., Mg²⁺), direct interaction with DNA polymerase, or degradation of nucleic acids [59].
In dPCR, inhibition manifests through several observable effects on partition fluorescence profiles. The primary manifestations include:
- Reduced fluorescence amplitude in positive partitions due to impaired amplification efficiency [60]
- Increased background fluorescence in negative partitions caused by auto-fluorescent compounds [59]
- Poor cluster separation between positive and negative populations, resulting in increased "rain" [60]
- Underestimation of target concentration when inhibition severity compromises amplification [59]
dPCR vs. qPCR Inhibition Resilience
Comparative studies demonstrate that dPCR generally exhibits higher resilience to inhibition than quantitative PCR (qPCR). Research on Pepper mild mottle virus detection in complex matrices showed that reverse transcription dPCR (RT-ddPCR) maintained accurate quantification in the presence of inhibitors where RT-qPCR significantly underestimated targets [59]. This enhanced tolerance stems from the partitioning process which effectively dilutes inhibitors across thousands of reactions, with each partition acting as an individual microreactor where amplification can proceed even when inhibitors are present at concentrations that would compromise bulk PCR [57] [59].
Table 1: Comparative Performance of dPCR and qPCR in Inhibitor-Rich Matrices
| Matrix/Inhibitor | Effect on qPCR | Effect on dPCR | Mechanism |
|---|---|---|---|
| Seed extracts | Significant underestimation | Moderate underestimation | Fluorescent compounds increase background |
| Tannic acid | Complete inhibition at high concentrations | Dose-dependent reduction in positive partitions | Polymerase inhibition |
| Humic acids | Reduced efficiency, increased Cq | Minimal impact on quantification | DNA binding, polymerase inhibition |
| Plant polysaccharides | Delayed amplification, reduced efficiency | Slight reduction in fluorescence amplitude | Polymerase competition |
| Wastewater effluent | Significant quantification bias | Maintained accuracy at low dilutions | Multiple mechanisms |
Protocols for Low-Input Sample Analysis
NSCLC Case Study: Experimental Design
A 2024 study directly compared dPCR performance with next-generation sequencing (NGS) for non-small cell lung cancer (NSCLC) biomarker detection using limited inputs [58]. The experimental design evaluated the ChromaCode HDPCR NSCLC Panel against the Illumina TSO500 NGS assay across serially diluted DNA and RNA inputs from FFPE reference standards and clinical samples. Inputs were titrated from 40 ng down to 0.5 ng for DNA and from 20 ng to 0.25 ng for RNA to simulate conditions of sample exhaustion [58].
The dPCR methodology utilized the QIAcuity platform with 26K nanoplate 24-well plates. For DNA targets, the master mix combined 10.5 μL of QIAcuity Probe Master Mix, 8.4 μL of HDPCR Mix, and variable volumes of molecular-grade water. For RNA fusion detection, the reaction included 10.5 μL of QIAcuity OneStep Advance Probe Master Mix, 0.45 μL of OneStep RT Mix, 8.4 μL of HDPCR Mix, and variable water volumes [58]. Sample inputs were adjusted to achieve the desired mass in each reaction, with a total reaction volume of 39 μL loaded per well.
Low-Input Performance Data
The NSCLC panel study generated comprehensive quantitative data on assay performance across input ranges, summarized in Table 2 below.
Table 2: dPCR Performance Metrics Across DNA/RNA Input Ranges in NSCLC Detection [58]
| Input (DNA/RNA) | Sensitivity (%) | Specificity (%) | PPA (%) | NPA (%) | Success Rate (%) |
|---|---|---|---|---|---|
| 40/20 ng | 100 | 100 | 100 | 100 | 100 |
| 20/10 ng | 100 | 100 | 100 | 100 | 100 |
| 15/7.5 ng | >95 | 100 | >95 | 100 | 100 |
| 10/5 ng | >95 | 100 | >95 | 100 | 100 |
| 5/2.5 ng | 85-90 | 100 | 85-90 | 100 | 90 |
| 2/1 ng | 70-75 | 100 | 70-75 | 100 | 75 |
| NGS (100/100 ng) | ~80 (at low inputs) | 100 | ~80 (at low inputs) | 100 | ~80 |
The data demonstrated that dPCR maintained high sensitivity (>95%) and positive percent agreement (PPA) at inputs as low as 10 ng DNA and 5 ng RNA, while NGS sensitivity declined by up to 86% at comparable low inputs [58]. This performance advantage is particularly valuable for NSCLC samples where tissue quantity is often limiting, such as with fine-needle aspirates or core needle biopsies, which have reported failure rates of approximately 20.2% with NGS methods [58].
Figure 1: Experimental workflow for handling low-input and low-quality samples in oncology dPCR applications.
Mitigation Strategies for PCR Inhibition
Sample Preparation and Purification
Effective inhibition mitigation begins with optimized sample preparation. For FFPE tissues, increased incubation times with proteinase K (overnight at 56°C) improves nucleic acid recovery and reduces cross-linking artifacts [58]. For liquid biopsies, volume-defined extraction methods consistently yield higher DNA quantities compared to bead-based isolation. Incorporating inhibitor removal steps, such as the use of Qiagen DNeasy PowerClean Pro Cleanup kits for environmental samples or OneStep PCR Inhibitor Removal columns for clinical specimens, significantly reduces co-purified inhibitors [59].
Dilution strategies represent a straightforward approach to mitigation, but require validation for each sample type. In wastewater surveillance, a 1:10 dilution typically restores amplification efficiency without substantially compromising sensitivity [59]. For blood-derived samples, dilution in TE buffer containing 0.5% Tween-20 can neutralize heparin inhibition without significant template dilution.
Data Analysis Solutions for Inhibition
Advanced data analysis methods can computationally correct for inhibition effects. A 2022 study introduced a double threshold approach to address both inhibition-induced fluorescence reduction and artifactual high-fluorescence droplets ("stars") [60]. This method approximates positive and negative droplet distributions as normal distributions and establishes separate thresholds for classification.
The double threshold method involves:
- Exporting raw fluorescence amplitude data from all partitions
- Calculating mean (μ) and standard deviation (σ) for negative population
- Setting lower threshold at μnegative + 5σnegative to exclude "rain"
- Calculating mean (μ) and standard deviation (σ) for positive population
- Setting upper threshold at μpositive - 3σpositive to exclude "stars"
- Applying both thresholds for accurate positive/negative classification
This approach demonstrated superior performance compared to single-threshold methods in environmental samples with varying inhibitor levels, reducing both false negatives and false positives [60].
Figure 2: Comprehensive strategy for identifying and mitigating PCR inhibition in dPCR workflows.
Detailed Experimental Protocols
Protocol 1: Low-Input NSCLC Mutation Detection
This protocol adapts the ChromaCode HDPCR NSCLC Panel for minimal input samples [58]:
Materials:
- QIAcuity Nanoplate 26K 24-well plates
- QIAcuity Probe Master Mix (DNA targets)
- QIAcuity OneStep Advance Probe Master Mix (RNA fusions)
- OneStep RT Mix
- HDPCR NSCLC Panel reagents
- Molecular-grade water
- Extracted DNA/RNA samples
Procedure:
- Sample Quantification: Quantify DNA using Qubit dsDNA BR Assay and RNA using Qubit RNA BR Assay. Record DNA Integrity Number (DIN) and RNA Integrity Number (RIN) if possible.
- Master Mix Preparation (DNA Wells):
- Combine 10.5 μL QIAcuity Probe Master Mix
- Add 8.4 μL HDPCR Mix
- Include variable volume of molecular-grade water (calculated based on sample volume)
- Add sample DNA (adjust volume to achieve 10-40 ng total input)
- Mix thoroughly by pipetting
- Master Mix Preparation (RNA Wells):
- Combine 10.5 μL QIAcuity OneStep Advance Probe Master Mix
- Add 0.45 μL OneStep RT Mix
- Add 8.4 μL HDPCR Mix
- Include variable volume of molecular-grade water
- Add sample RNA (adjust volume to achieve 5-20 ng total input)
- Mix thoroughly by pipetting
- Plate Loading: Pipette 39 μL of each reaction mix into designated wells of the QIAcuity Nanoplate.
- Partitioning and Amplification: Run plate on QIAcuity according to manufacturer's instructions with the following cycling conditions:
- Initial denaturation: 95°C for 2 minutes
- 50 cycles of: 95°C for 15 seconds, 60°C for 30 seconds, 72°C for 30 seconds
- Final extension: 72°C for 5 minutes
- Hold: 4°C
- Data Analysis: Analyze using ChromaCode Cloud platform, which reports detected targets and estimated mutant allele fraction (MAF).
Validation: Include positive controls with known mutation status and no-template controls in each run. For inputs below 10 ng DNA/5 ng RNA, increase cycle number to 55 and expect slightly reduced precision.
Protocol 2: Inhibition-Resistant ddPCR for Challenging Matrices
This protocol is adapted from environmental detection studies [60] [59] and optimized for inhibitory clinical samples:
Materials:
- Bio-Rad QX200 Droplet Generator
- DG8 Cartridges and Gaskets
- Droplet Reader Oil
- EvaGreen Supermix or ddPCR Supermix for Probes
- PCR-grade water
- Inhibitor-resistant DNA polymerase (optional)
Procedure:
- Inhibition Assessment:
- Run undiluted sample and 1:5 diluted sample in parallel
- Compare absolute quantification values - significant differences indicate inhibition
- Check droplet clustering quality - poor separation suggests inhibition
- Reaction Setup:
- Prepare 20 μL reactions containing:
- 11 μL EvaGreen Supermix
- 200 nM primers (final concentration)
- 1-5 μL template DNA (adjust based on expected concentration)
- PCR-grade water to 20 μL
- For inhibited samples, supplement with:
- 0.5 μg/μL BSA (final concentration)
- 0.5% Tween-20 (final concentration)
- Prepare 20 μL reactions containing:
- Droplet Generation:
- Transfer 20 μL reaction mix to DG8 cartridge well
- Add 70 μL Droplet Generation Oil to appropriate well
- Place gasket and cartridge in QX200 Droplet Generator
- Generate droplets according to manufacturer's protocol
- Transfer and Amplification:
- Carefully transfer 40 μL emulsified reaction to 96-well PCR plate
- Seal plate with foil heat seal
- Amplify using the following conditions:
- 95°C for 5 minutes
- 50 cycles of: 95°C for 30 seconds, 60°C for 1 minute
- 4°C hold
- Ramp rate: 2°C/second
- Droplet Reading and Analysis:
- Load plate in QX200 Droplet Reader
- Analyze using QuantaSoft software with manual thresholding or export raw data for double-threshold analysis
Double-Threshold Analysis:
- Export raw fluorescence data (CSV format)
- Calculate mean (μneg) and standard deviation (σneg) of negative droplet population
- Set lower threshold at μneg + 5×σneg
- Identify positive population above lower threshold
- Calculate mean (μpos) and standard deviation (σpos) of positive population
- Set upper threshold at μpos - 3×σpos to exclude "stars"
- Count positive droplets between lower and upper thresholds
- Calculate concentration using Poisson statistics
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Key Research Reagent Solutions for Inhibition and Low-Input Challenges
| Reagent/Material | Function | Application Notes |
|---|---|---|
| QIAcuity Probe Master Mix | Optimized chemistry for nanoplate dPCR | Provides consistent partitioning and amplification in 26K-96K formats [58] |
| OneStep RT Mix | Reverse transcription for RNA targets | Enables cDNA synthesis directly in partition reaction [58] |
| HDPCR NSCLC Panel | Multiplexed detection of NSCLC mutations | Designed for low-input FFPE samples (10 ng DNA, 5 ng RNA) [58] |
| BSA (Fraction V) | Inhibition mitigation | Binds inhibitors, stabilizes polymerase (0.1-0.5 μg/μL final) [60] |
| Tween-20 | Surfactant for inhibition resistance | Reduces surface adhesion of inhibitors (0.1-0.5% final) [60] |
| Droplet Generation Oil | Creates water-in-oil emulsion | Formulated with stabilizers to prevent coalescence during cycling [60] |
| Qubit dsDNA/RNA BR Assays | Accurate nucleic acid quantification | Essential for low-input workflow planning [58] |
| Inhibitor-Resistant Polymerase | Enhanced amplification efficiency | Engineered enzymes with higher tolerance to clinical inhibitors [59] |
Effective management of inhibition and low-input challenges is essential for reliable dPCR quantification in oncology research. The protocols and strategies presented here demonstrate that through optimized sample preparation, reaction reformulation, and advanced data analysis, dPCR can deliver precise absolute quantification even with suboptimal samples. The resilience of dPCR to these challenges positions it as a valuable tool for precious oncology specimens where material is limited and results have direct implications for therapeutic decision-making. As dPCR technology continues to evolve with higher partition densities and more robust chemistry, its utility for challenging clinical samples will further expand, solidifying its role in precision oncology research.
Digital PCR (dPCR) represents a significant advancement in nucleic acid quantification by enabling absolute target measurement without standard curves. This application note details established protocols for threshold setting and Poisson correction in dPCR data analysis, with particular emphasis on oncology research applications. We provide comprehensive validation data and step-by-step methodologies to ensure precise quantification of rare genetic variants, including cancer-associated mutations, at frequencies as low as 0.001% [61]. The protocols outlined herein support reliable molecular monitoring for therapeutic assessment and minimal residual disease detection in hemato-oncology and solid tumor management.
Digital PCR operates through sample partitioning into thousands of individual reactions, effectively creating a binary readout (positive or negative) for each partition [62]. The fundamental principle involves statistical application of Poisson distribution to these binary outcomes to calculate absolute target concentration [63]. This method offers distinctive advantages over quantitative PCR (qPCR), particularly for low-abundance targets, as it eliminates requirement for standard curves and demonstrates enhanced resistance to PCR inhibitors [61]. In oncology research, these characteristics prove invaluable for detecting rare somatic mutations, monitoring treatment response, and detecting minimal residual disease with exceptional precision [64] [61].
The accuracy of dPCR quantification hinges on two critical analytical components: appropriate fluorescence threshold setting to distinguish positive from negative partitions, and proper application of Poisson statistics to account for multiple target molecules occupying individual partitions. This document addresses both components through optimized protocols and validation data from oncology research applications.
Threshold Setting Methodologies
Theoretical Framework
Threshold setting establishes the fluorescence amplitude that discriminates between positive partitions (containing the target sequence) and negative partitions (lacking the target) [63]. Improper threshold placement introduces quantification bias, particularly impactful for low-frequency targets common in oncology applications such as circulating tumor DNA analysis.
Experimental Approach to Threshold Optimization
We recommend a empirical strategy for threshold determination using experimental controls:
- Control Samples: Include no-template controls (NTC) and known negative biological samples to establish the negative cluster baseline [63].
- Manual Threshold Adjustment: Set the threshold at the minimum point between clearly defined positive and negative populations [63].
- Validation Samples: Use samples with known target concentrations across the expected range, including low-abundance targets (0.01-0.1%) relevant to cancer mutation detection [61].
Table 1: Threshold Setting Performance Metrics Using Different Methodologies
| Methodology | Precision (CV%) | Accuracy (%) | Best Use Cases |
|---|---|---|---|
| Empirical Manual Setting | 1-5% | 95-105% | Clear cluster separation |
| Automated Algorithm | 2-7% | 90-110% | High-throughput screening |
| Cluster-based | 1-3% | 98-102% | Low-abundance targets |
Impact of Threshold Variability on Quantification
In a validation study examining BCR-ABL fusion detection, threshold variations of ±10% resulted in concentration changes of ≤5% for high-copy targets (>100 copies/μL) but up to 15% variation for low-copy targets (<10 copies/μL) [61]. This highlights the critical importance of consistent threshold application in minimal residual disease monitoring where detection sensitivity below 0.001% may be required [61].
Poisson Correction Implementation
Mathematical Foundation
The Poisson distribution accounts for the probability of multiple target molecules residing in a single partition. The fundamental equation for absolute quantification is:
λ = -ln(1 - p)
Where λ represents the average number of target molecules per partition and p is the ratio of positive partitions to total partitions [63] [62]. This correction becomes increasingly important as target concentration increases, as multiple copies in partitions would otherwise lead to underestimation without proper statistical adjustment.
Protocol: Poisson Correction Application
Materials Required:
- dPCR system with partitioning capability (droplet or chip-based)
- Nucleic acid sample with known concentration for validation
- Appropriate assay reagents (primers, probes, master mix)
Procedure:
- Partition Generation: Create 10,000-20,000 partitions per reaction [63] [61].
- Amplification: Perform PCR amplification with endpoint detection.
- Partition Counting: Identify positive and negative partitions.
- Calculate Positive Fraction: Determine p (positive partitions/total partitions).
- Apply Poisson Correction: Calculate λ = -ln(1 - p).
- Determine Concentration: Convert to copies/μL using partition volume.
Validation Criteria:
- Optimal positive fraction range: 0.05-0.95 [63]
- Acceptable positive fraction range: 0.01-0.99 [62]
- For rare target detection (<0.1%), ensure sufficient total partitions to capture adequate positive events [61]
Table 2: Impact of Positive Fraction on Quantification Accuracy
| Positive Fraction | Without Poisson Correction | With Poisson Correction | Recommended Application |
|---|---|---|---|
| <0.05 | Underestimation by 1-10% | Accurate quantification | Rare target detection |
| 0.1-0.9 | Minimal error (<1%) | Accurate quantification | Routine quantification |
| >0.9 | Significant underestimation | Accurate quantification | High concentration targets |
Limitations and Considerations
The Poisson model assumes random distribution of target molecules and perfect amplification efficiency. In practice, factors including sample viscosity, partition uniformity, and amplification inhibitors can affect these assumptions [63]. For rare allele detection in oncology (e.g., <0.01%), ensure sufficient partitions to achieve statistically significant detection - typically requiring >100,000 partitions for reliable detection at 0.001% frequency [61].
Experimental Validation Protocol
Objective
Validate dPCR performance for detection and quantification of cancer-associated genetic variants using appropriate threshold setting and Poisson correction.
Materials and Equipment
Table 3: Research Reagent Solutions for dPCR Oncology Applications
| Reagent/Material | Function | Example Specifications |
|---|---|---|
| dPCR Master Mix | Provides optimized reaction components | Contains polymerase, dNTPs, buffer |
| Assay-specific Primers/Probes | Target sequence detection | Hydrolysis probes for target and control |
| Partitioning Oil/Matrix | Creates nanoreactors | Generates 20,000 droplets per reaction |
| Reference DNA | Positive control | Certified copy number concentration |
| Nucleic Acid Purification Kits | Sample preparation | Removes PCR inhibitors from blood/bone marrow |
Step-by-Step Procedure
Sample Preparation:
- Extract nucleic acids from patient samples (blood, bone marrow, tissue) [61].
- Quantify DNA using fluorescence methods.
- Dilute to working concentration (10-100 ng/μL).
Reaction Setup:
- Prepare master mix containing: 10 μL dPCR supermix, 1 μL assay-specific primers/probes (20X), 50 ng template DNA, nuclease-free water to 20 μL total volume [63].
- Include no-template controls and positive controls with known concentration.
Partitioning and Amplification:
Data Acquisition and Analysis:
- Read endpoint fluorescence for all partitions.
- Set discrimination threshold using negative control population.
- Record counts of positive and negative partitions.
- Apply Poisson correction to calculate absolute concentration.
Application in Oncology Research
Rare Mutation Detection
dPCR enables precise quantification of mutant alleles in a background of wild-type DNA, critical for oncology applications where mutation frequencies may be extremely low (0.001-0.1%) [61]. The technology demonstrates superior sensitivity and precision for targets such as BCR-ABL fusions in leukemia and NPM1 mutations in AML compared to qPCR, particularly at very low allele frequencies [61].
Method Performance Characteristics
Validation studies for hemato-oncology applications demonstrate:
- Detection Sensitivity: Reliable detection down to 0.001% mutant allele frequency [61]
- Precision: Coefficient of variation <10% even at low copy numbers [63]
- Dynamic Range: 1 to 100,000 copies/μL without dilution [62]
- Inhibitor Tolerance: Robust performance with inhibitors that typically affect qPCR [61]
Multiplex Detection Strategies
dPCR facilitates multiplex detection of multiple mutation types in single reactions, enabled by compartmentalization that reduces inter-assay competition [61]. This capability proves particularly valuable in oncology for comprehensive mutation profiling with limited sample material.
Troubleshooting Guide
Table 4: Common Data Analysis Challenges and Solutions
| Issue | Potential Cause | Recommended Solution |
|---|---|---|
| Poor cluster separation | Suboptimal amplification efficiency | Validate assay design; optimize annealing temperature |
| High negative control signal | Contamination or non-specific amplification | Implement rigorous contamination controls; redesign probes |
| Inconsistent results between replicates | Partition quality issues or pipetting errors | Verify partition uniformity; improve pipetting technique |
| Significant deviation from expected values | Improper threshold setting or Poisson application | Validate with reference material; review threshold placement |
Proper implementation of threshold setting and Poisson correction methodologies ensures accurate absolute quantification in dPCR applications. These protocols enable reliable detection and measurement of rare genetic variants in oncology research, supporting advancements in cancer diagnostics, treatment monitoring, and minimal residual disease assessment. The experimental approaches outlined provide a framework for robust dPCR assay implementation with performance characteristics suitable for the most demanding research and clinical validation applications.
Benchmarking dPCR: Analytical Validation and Performance Comparison with qPCR and NGS
Digital PCR (dPCR) represents a third generation of polymerase chain reaction technology, enabling the absolute quantification of nucleic acids without the need for a standard curve [1]. This calibration-free technology operates by partitioning a PCR reaction mixture into thousands to millions of individual reactions, so that each partition contains either 0, 1, or a few nucleic acid targets [1]. Following amplification, the fraction of positive partitions is counted via end-point fluorescence detection, and the absolute concentration of the target molecule is calculated using Poisson statistics [1] [65]. This partitioning approach provides dPCR with powerful advantages for oncology research, including exceptional sensitivity, absolute quantification, high accuracy, and reproducibility [1].
In the context of oncology, dPCR has emerged as a pivotal tool for detecting rare genetic mutations, monitoring treatment response, and quantifying minimal residual disease (MRD) [1] [65]. The technology's ability to provide both a qualitative (yes/no) call and precise quantitative measurement in a single assay makes it particularly valuable for tracking genomic biomarkers throughout the course of cancer treatment [65]. For clinical researchers and drug development professionals, establishing robust analytical validation parameters—specifically sensitivity, specificity, and precision—is fundamental to ensuring the reliability of dPCR data for both research and eventual clinical applications.
Core Analytical Validation Parameters
The implementation of dPCR in regulated research and potential clinical diagnostics necessitates rigorous analytical validation. Key performance parameters must be experimentally established to ensure the technology meets the required standards for accuracy and reliability.
Sensitivity
Sensitivity in dPCR encompasses both the ability to detect low-abundance targets (limit of detection) and to accurately quantify them (limit of quantification). A 2025 study developing a reverse transcription dPCR (RT-dPCR) for Hepatitis D virus (HDV) RNA quantification demonstrated an exceptionally low limit of detection (LOD) of 0.7 copies/mL and a limit of quantification (LOQ) of 10 copies/mL [66]. This sensitivity is particularly crucial in oncology for applications like liquid biopsy, where circulating tumor DNA (ctDNA) can be present at very low frequencies.
The superior sensitivity of dPCR is further highlighted in direct comparisons with quantitative PCR (qPCR). In a study comparing multiplex dPCR to qPCR for detecting periodontal pathobionts, dPCR demonstrated superior sensitivity, detecting lower bacterial loads that resulted in qPCR false negatives and a 5-fold underestimation of pathogen prevalence at low concentrations (< 3 log10 Geq/mL) [67]. This enhanced detection capability for rare alleles and low-copy transcripts enables dPCR to identify molecular recurrence months before radiologic relapse in oncology settings [65].
Specificity
Specificity refers to the ability of a dPCR assay to correctly distinguish and quantify the intended target without cross-reactivity or interference. The high specificity of dPCR is achieved through precise primer and probe design, optimized reaction conditions, and the physical separation of targets during partitioning.
In a 2025 study developing a droplet digital PCR (ddPCR) assay for determining FRS2 gene copy number in bladder cancer, researchers demonstrated 100% sensitivity and 100% specificity when compared to fluorescence in situ hybridization (FISH) as a reference method, with a kappa value of 1 indicating perfect agreement [68]. The assay showed clear separation between positive and negative droplets in one-dimensional fluorescence amplitude plots, and duplex detection of both the target gene (FRS2) and reference gene (RPP30) within the same reaction exhibited no interference between primers or probes [68].
Assay specificity can be further enhanced through sophisticated probe design and melting-curve analysis. For instance, one study combined multiplex dPCR with melting-curve analysis to improve ctDNA detection efficiency, lowering the limit of detection to below 0.2% variant allele frequency while accurately genotyping KRAS mutations in pancreatic cancer [65].
Precision
Precision describes the reproducibility of measurements under defined conditions, encompassing both repeatability (intra-assay precision) and reproducibility (inter-assay precision). Digital PCR exhibits remarkable precision due to its partitioning technology and absolute quantification approach.
A comprehensive validation of the Bio-Rad QX200 Droplet dPCR system using a multifactorial experimental design demonstrated high precision, sensitivity, uniformity, and robustness [69]. Most experimental factors tested—including the operator, primer/probe system, and addition of restriction enzymes—had no relevant effect on DNA copy number quantification, confirming system robustness [69]. However, the study identified that the choice of ddPCR master mix was a critical factor affecting accuracy across the working range [69].
In the FRS2 copy number assay, precision was quantitatively evaluated through calculation of coefficients of variation (CV%). The assay demonstrated excellent repeatability and precision, with intra-assay CV% of 2.58% and 3.75%, and inter-assay CV% of 2.68% and 3.79%, across 20 ng and 2 ng input DNA levels, respectively [68]. These low CV values indicate high reproducibility essential for reliable longitudinal monitoring in oncology research.
Table 1: Summary of Key Analytical Performance Parameters from Recent Studies
| Parameter | Performance Metric | Experimental Context | Reference |
|---|---|---|---|
| Sensitivity | LOD: 0.7 copies/mL; LOQ: 10 copies/mL | HDV RNA quantification | [66] |
| Sensitivity | Detected 31% of samples negative by RT-qPCR | Clinical HDV samples at low concentrations | [66] |
| Sensitivity | 5-fold higher pathogen detection vs qPCR | Periodontal pathobiont detection | [67] |
| Specificity | 100% sensitivity and specificity vs FISH | FRS2 copy number in bladder cancer | [68] |
| Precision | Intra-assay CV: 2.58-3.75%; Inter-assay CV: 2.68-3.79% | FRS2 copy number validation | [68] |
| Precision | Lower intra-assay variability (median CV: 4.5%) vs qPCR | Periodontal pathobiont quantification | [67] |
| Robustness | Operator, primer/probe system, enzymes had no relevant effect | Multifactorial validation of ddPCR system | [69] |
Table 2: Comparison of Digital PCR Performance Across Applications
| Application Area | Key Advantage | Performance Evidence | Reference |
|---|---|---|---|
| Copy Number Variation | 95% concordance with PFGE (gold standard) | DEFA1A3 CNV analysis vs PFGE | [5] |
| Viral Load Quantification | Absolute quantification without standard curve | HDV RNA detection and monitoring | [66] |
| Liquid Biopsy / ctDNA | Detection of <0.2% variant allele frequency | KRAS mutations in pancreatic cancer | [65] |
| Cancer Biomarker Detection | 100% concordance with FISH | FRS2 amplification in bladder cancer | [68] |
| Pathogen Detection | Superior sensitivity and precision vs qPCR | Periodontal pathobiont quantification | [67] |
Experimental Protocols for Validation
Protocol 1: Determining Limit of Detection (LOD) and Limit of Quantification (LOQ)
This protocol follows the approach used in the HDV RNA validation study [66]:
Prepare Dilution Series: Create fourteen two-fold serial dilutions of the target nucleic acid, starting from a concentration known to be detectable.
Replicate Measurements: For each dilution, perform twenty replicates in four separate runs with five technical replicates each. For the last two dilutions (lowest concentrations), measure fifty replicates in five separate runs with ten technical replicates.
Include Controls: In each run, include negative template controls (NTCs) with at least one NTC per 8-well strip.
Calculate LOD and LOQ: Use the RT-dPCR values to determine the assigned copy number for each dilution. Calculate LOD and LOQ concentrations as described by international standards, such as the method referenced in [66].
Protocol 2: Establishing Precision (Intra-assay and Inter-assay)
This protocol is adapted from the FRS2 copy number validation study [68]:
Sample Preparation: Use genomic DNA extracted from appropriate sources (e.g., healthy donor samples for baseline, cancer cell lines for elevated copy numbers).
Intra-assay Precision:
- Test two different input DNA amounts (e.g., 20 ng and 2 ng)
- Perform each measurement in triplicate within the same run
- Calculate CV% for each DNA concentration
Inter-assay Precision:
- Using the same DNA amounts (20 ng and 2 ng)
- Measure in triplicate per day for five consecutive days
- Calculate intra-assay CV% and inter-assay CV% according to CLSI EP05-A3 guidelines
Define Minimum Reliable Input: Determine the lowest DNA input that results in a CV% less than 5% as the minimum reliable input for the assay.
Protocol 3: Assessing Specificity Against Reference Methods
This protocol follows the approach used for validating FRS2 copy number detection against FISH [68]:
Sample Selection: Obtain clinical samples with known status (e.g., FFPE bladder cancer tissues with varying FRS2 amplification status).
Reference Method Testing: Perform FISH analysis using dual-probe assays (target gene probe and centromeric reference probe). Count signals from 25+ randomly selected tumor nuclei per sample.
dPCR Testing: Run the same samples using the optimized ddPCR assay with target and reference genes.
Statistical Comparison: Calculate sensitivity, specificity, and kappa coefficient to measure agreement between methods. A case is considered positive by FISH if the target/reference ratio is ≥ 2.0 and the average target copy number per cell is ≥ 4.0.
Signaling Pathways and Workflows
Diagram 1: Digital PCR Analytical Validation Workflow
Diagram 2: Relationship Between Analytical Validation Parameters
Research Reagent Solutions
Table 3: Essential Materials for Digital PCR Validation
| Reagent/Equipment | Function | Example Products/Brands |
|---|---|---|
| dPCR Master Mix | Critical component affecting accuracy; contains DNA polymerase, dNTPs, buffer | Bio-Rad "Supermix for Probes (no dUTP)" [69], One-Step RT-ddPCR Advanced Kit [66] |
| Primers/Probes | Target-specific amplification and detection; must be optimized for specificity | LNA (Locked Nucleic Acid) Mutation Assays [70], Custom-designed primers [68] |
| Partitioning System | Creates thousands of individual reactions for absolute quantification | Bio-Rad QX200 [66], Stilla Technologies Naica System [66], QIAGEN QIAcuity [71] |
| Reference Gene Assay | Normalization control for copy number variation studies | RPP30 reference gene [68], other stable single-copy genes |
| DNA Extraction Kits | Obtain high-quality nucleic acids from various sample types | INSTANT virus RNA/DNA kit [66], QIAamp Viral RNA Mini Kit [66] |
| Restriction Enzymes | May be used to fragment genomic DNA for improved partitioning | Various (validation studies show minimal effect on quantification) [69] |
Within oncology research, the precise quantification of nucleic acids is fundamental for advancing personalized medicine, from monitoring treatment response to detecting minimal residual disease. Quantitative real-time PCR (qPCR) has long been the cornerstone technique for such applications. However, digital PCR (dPCR), the third generation of PCR technology, presents a paradigm shift by enabling absolute quantification without the need for a standard curve [45] [1]. This application note provides a structured, evidence-based comparison of dPCR and qPCR, focusing on two critical performance parameters for molecular diagnostics in oncology: analytical sensitivity and tolerance to PCR inhibitors. Data and protocols are contextualized within the framework of absolute quantification needs in oncology research and drug development.
The following table synthesizes key performance metrics from recent comparative studies across various sample types, highlighting the advantages of dPCR in sensitive detection and robust quantification.
Table 1: Head-to-Head Comparison of dPCR and qPCR Performance Across Various Applications
| Application / Target | Key Performance Metric | qPCR Performance | dPCR/ddPCR Performance | Reference & Context |
|---|---|---|---|---|
| Periodontal Pathobionts (P. gingivalis, A. actinomycetemcomitans) | Sensitivity (Detection Rate) | Lower sensitivity; false negatives at low concentrations (< 3 log10Geq/mL) | Superior sensitivity; 5-fold higher prevalence estimate for A. actinomycetemcomitans | [72] Subgingival plaque samples |
| Precision / Reproducibility | Higher intra-assay variability (Median CV%) | Lower intra-assay variability (Median CV: 4.5%) | [72] | |
| Multi-Strain Probiotics (B. lactis Bl-04, etc.) | Limit of Detection (LoD) | Higher LoD | 10 to 100-fold lower LoD | [73] Fecal DNA samples |
| Plant Pathogen (P. nicotianae) | Positive Detection Rate | 83.9% | 96.4% | [74] Tobacco root and soil samples |
| Inhibitor Tolerance | Quantification in soil samples | Affected by inhibitors | Better accuracy for low concentrations; superior tolerance | [74] Complex soil matrices |
| Circulating miRNAs (miR-4488, miR-579-3p) | Sensitivity for Low-Abundance Targets | Lower sensitivity for miR-4488 | Superior sensitivity for low-abundance miRNAs | [3] Serum liquid biopsy |
| Viral Detection (Plum Pox Virus) | Sensitivity with Purified RNA | Lower sensitivity | Higher sensitivity than RT-qPCR | [75] Plant RNA and crude extract |
Essential Workflows and Understanding dPCR Advantages
The fundamental difference between the two technologies lies in their approach to quantification. qPCR relies on monitoring amplification in real-time and comparing the cycle threshold (Cq) to a standard curve for relative quantification [45]. In contrast, dPCR partitions a sample into thousands of nanoliter-scale reactions, performs endpoint amplification, and uses Poisson statistics to provide absolute quantification without a standard curve [45] [1].
Diagram: Workflow and Conceptual Comparison of qPCR and dPCR
This partitioning confers two main advantages in oncology research:
- Superior Sensitivity and Precision for Rare Targets: Partitioning a sample effectively enriches low-abundance targets, allowing for detection of rare mutations or minimal residual disease present at fractions as low as 0.001% [45] [3].
- Enhanced Tolerance to PCR Inhibitors: By distributing inhibitors across thousands of partitions, the amplification in inhibitor-free partitions remains efficient, making dPCR robust for complex samples like serum, plasma, and stool [74] [73].
The Scientist's Toolkit: Key Research Reagent Solutions
The following table outlines essential reagents and their critical functions for setting up dPCR experiments, particularly in an oncology context.
Table 2: Essential Reagents for dPCR Assay Development
| Reagent / Material | Function / Description | Application Note |
|---|---|---|
| dPCR Master Mix | Provides core components for amplification (polymerase, dNTPs, buffer). | Critical for performance. "Supermix for Probes (no dUTP)" confirmed for accurate quantification [69]. |
| Primers & Hydrolysis Probes | Sequence-specific oligonucleotides for target amplification and detection. | Often labeled with FAM, HEX/VIC, TAMRA/Atto550, Cy5 [45]. Same designs often transferable from qPCR [73]. |
| Restriction Enzyme | Redes DNA viscosity by cutting long strands; may improve partition uniformity. | e.g., Anza 52 PvuII; studies show it is not always a critical factor [72] [69]. |
| RNase Inhibitor | Protects RNA templates from degradation during reverse transcription steps. | Essential for RT-dPCR of miRNA/RNA; improves droplet cluster separation [75]. |
| Microfluidic Plates/Cartridges | Solid chips with pre-formed wells for partition generation. | Used in systems like QIAcuity [72]. Provides high reproducibility. |
| Droplet Generation Oil/Surfactant | Creates stable water-in-oil emulsion for droplet-based dPCR systems. | Essential for droplet stability; prevents coalescence during thermocycling [45] [1]. |
Detailed Experimental Protocols
Protocol: Duplex dPCR for Circulating miRNA Quantification in Liquid Biopsy
This protocol is adapted from a metastatic melanoma study for simultaneous quantification of two miRNAs in serum [3].
1. Sample Preparation and RNA Extraction:
- Process serum samples (200 µL) using the miRNeasy Mini Kit (Qiagen) or equivalent.
- Elute RNA in a small volume (e.g., 20 µL) of nuclease-free water. Note that concentration may be below standard spectrophotometer detection limits.
2. Reverse Transcription and Preamplification:
- Use the TaqMan Advanced miRNA cDNA Synthesis Kit.
- Perform reverse transcription with a fixed input volume (e.g., 2 µL of eluted RNA), as concentration is often too low for accurate quantification.
- Include a preamplification step per the kit protocol to enrich target miRNA sequences.
3. Duplex dPCR Reaction Setup:
- Reaction Mix (40 µL total volume, adjust based on system):
- 10 µL of 4x Probe PCR Master Mix
- 0.4 µM of each specific forward and reverse primer
- 0.2 µM of each specific probe (e.g., labeled with FAM and HEX/VIC)
- 2-5 µL of preamplified cDNA template
- Nuclease-free water to 40 µL
- Critical Step: Include an RNase inhibitor in the mix to prevent RNA degradation and reduce "rain" [75].
4. Partitioning and Thermocycling:
- Load the reaction mix into a dPCR plate or cartridge for partitioning (e.g., generating ~20,000 droplets or partitions).
- Thermocycling Conditions:
- Enzyme activation: 95°C for 2 minutes
- 45 cycles of:
- Denaturation: 95°C for 15 seconds
- Annealing/Extension: 58-60°C for 1 minute
- Signal stabilization: 98°C for 10 minutes (for droplet systems)
- Hold at 4°C
5. Imaging and Data Analysis:
- Read the plate on an appropriate dPCR instrument (e.g., QIAcuity Four, QX200 Droplet Reader).
- Set fluorescence thresholds for each channel to distinguish positive and negative partitions.
- Use manufacturer's software (e.g., QuantaSoft, QIAcuity Software Suite) for absolute quantification based on Poisson statistics.
- Calculate the expression ratio (miRatio) of the two miRNAs directly from the copies/µL values provided by the software [3].
Protocol: dPCR Assay for Absolute Quantification of Vector Copy Number in CAR-T Cells
This protocol outlines a robust method for assessing the safety of genetically engineered T-cell products [76].
1. Genomic DNA (gDNA) Extraction:
- Extract high-quality gDNA from CAR-T or TCR-T cell products using a standard silica-column or magnetic bead-based kit.
- Accurately quantify gDNA using a fluorometric method (e.g., Qubit dsDNA HS Assay).
2. Assay Design:
- Design two assays:
- Target Assay: Specific to the transgene integrated into the T-cell genome.
- Reference Assay: Specific to a single-copy endogenous human gene (e.g., RPP30).
3. ddPCR Reaction Setup:
- Reaction Mix (20 µL total volume for droplet systems):
- 10 µL of 2x ddPCR Supermix for Probes (no dUTP)
- 1 µL of each primer/probe mix (final concentration 500 nM/250 nM)
- 50-100 ng of input gDNA
- Nuclease-free water to 20 µL
- Partitioning: Generate droplets using an Automated Droplet Generator.
4. Thermocycling and Reading:
- Amplify the droplets using a standard thermal cycler with the following conditions:
- Initial denaturation: 95°C for 10 minutes
- 40-45 cycles of:
- Denaturation: 94°C for 30 seconds
- Annealing/Extension: 58-60°C for 1 minute
- Enzyme deactivation: 98°C for 10 minutes
- Hold at 4°C
- Read the droplets using a Droplet Reader.
5. Data Analysis and VCN Calculation:
- The software provides the absolute concentration (copies/µL) for both the target (transgene) and reference (endogenous gene) assays.
- Calculate the bulk VCN (VCN~bulk~):
VCN~bulk~ = (Transgene copies/µL) / (Endogenous gene copies/µL)
- Calculate the adjusted VCN (VCN~adj~): To account for transduction efficiency and provide a more accurate VCN within transduced cells, apply a correction based on Poisson distribution: VCN~adj~ = VCN~bulk~ / [1 - e^(-VCN~bulk^)] This adjustment is validated against sorted transgene-positive cell populations [76].
The collective evidence demonstrates that dPCR offers tangible advantages over qPCR for specific applications in oncology research, particularly where absolute quantification, high sensitivity for rare targets, and robustness in complex matrices are required. The provided protocols for liquid biopsy miRNA analysis and CAR-T cell VCN quantification serve as templates that can be adapted for various targets, empowering researchers to implement this powerful technology in their pursuit of precise and reliable molecular data.
Circulating tumor DNA (ctDNA), a fraction of cell-free DNA (cfDNA) shed into the bloodstream by tumor cells, has emerged as a transformative biomarker in oncology. Its analysis enables minimally invasive liquid biopsies for cancer detection, molecular profiling, and monitoring treatment response. The two predominant technologies for ctDNA analysis are digital PCR (dPCR) and next-generation sequencing (NGS). dPCR provides unparalleled sensitivity for the absolute quantification of known mutations, while NGS offers a broad, hypothesis-free screening of genomic alterations. Recognizing that these are not competing but complementary technologies is crucial for deploying them effectively within clinical research and drug development programs. This application note details their performance characteristics, provides structured experimental protocols, and outlines a strategic framework for their complementary use in solid tumor profiling.
Technology Comparison: Performance and Application Scope
Quantitative Performance and Key Characteristics
Table 1: Comparative Performance of dPCR and NGS in ctDNA Analysis
| Feature | Digital PCR (dPCR) | Next-Generation Sequencing (NGS) |
|---|---|---|
| Principle | Partitioning, end-point PCR, and absolute quantification via Poisson statistics [1]. | Massive parallel sequencing of clonally amplified DNA fragments [77]. |
| Primary Strength | Ultra-sensitive detection and absolute quantification of known targets [1] [77]. | Comprehensive profiling of known and novel variants across multiple genes [78] [79]. |
| Typical VAF LOD | Can detect mutant allele frequencies as low as 0.001% (0.01% typical) [80] [77]. | Limit of Detection (LOD) typically ~0.5%; improvements pushing to 0.1% [78] [81]. |
| Detection Type | Target-specific; requires prior knowledge of the mutation sequence [77]. | Untargeted; can discover novel mutations, indels, CNVs, and fusions [78] [81]. |
| Quantification | Absolute quantification, without need for standard curves [1] [77]. | Relative quantification; requires complex bioinformatic analysis [78]. |
| Throughput | Low to medium throughput; ideal for tracking few known mutations in many samples [79]. | High throughput; suited for analyzing many genomic regions across multiple samples [79]. |
| Cost & Workflow | Lower operational cost and simpler, faster workflow [80] [77]. | Higher cost and more complex workflow, including library prep and bioinformatics [80] [78]. |
| Key Application | Monitoring MRD and treatment response for known mutations [80] [1]. | Comprehensive genomic profiling for therapy selection and clinical trial enrollment [78] [79]. |
A 2025 study in rectal cancer directly compared these technologies, finding that ddPCR detected ctDNA in 58.5% (24/41) of baseline plasma samples, significantly outperforming a targeted NGS panel which detected ctDNA in 36.6% (15/41) of the same samples (p=0.00075) [80] [82]. This underscores dPCR's superior analytical sensitivity for detecting known mutations, a critical factor in minimal residual disease (MRD) assessment.
Strategic Complementarity in Research Workflows
The choice between dPCR and NGS is not mutually exclusive but rather sequential and complementary. A typical integrated workflow involves:
- Discovery and Profiling Phase: Using NGS to identify tumor-specific somatic mutations from tissue or baseline liquid biopsies. This provides a comprehensive molecular profile of the tumor [79] [77].
- Informed Monitoring Phase: Designing patient-specific dPCR assays for the mutations identified by NGS. This tumor-informed approach allows for ultra-sensitive, cost-effective longitudinal monitoring of treatment response and MRD [80] [79].
NGS faces sensitivity challenges in low-ctDNA contexts. Its effective sensitivity is constrained by sequencing depth and the need for unique molecular identifiers (UMIs) to eliminate PCR and sequencing errors. Achieving a 99% detection probability for a variant at a 0.1% Variant Allele Frequency (VAF) requires a sequencing depth of approximately 10,000x, which remains prohibitively expensive for routine use [78]. Furthermore, the absolute quantity of input ctDNA is a fundamental limitation; in low-shedding tumors, the total number of mutant DNA molecules in a sample may be too low for reliable statistical detection by any method [78].
Experimental Protocols for ctDNA Analysis
Tumor-Informed dPCR for MRD Monitoring
This protocol enables ultra-sensitive tracking of one or two known mutations following their discovery via NGS.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Brief Description |
|---|---|
| Streck Cell-Free DNA BCT Tubes | Stabilizes blood samples for up to 14 days, preventing genomic DNA contamination and preserving ctDNA profile [80]. |
| cfDNA Extraction Kit | Silica membrane or magnetic bead-based kits optimized for recovery of short-fragment cfDNA from plasma. |
| TaqMan Assay Probes (FAM/HEX) | Hydrolysis probes designed against the specific mutant and wild-type sequences for target detection in dPCR [4]. |
| ddPCR Droplet Generator & Oil | Partitions the PCR reaction into ~20,000 nanodroplets for absolute quantification [80] [1]. |
| Droplet Reader (Fluorescence Detector) | Performs end-point fluorescence measurement of each droplet to determine positive/negative partitions [1]. |
Workflow:
Sample Collection & Processing:
- Collect patient blood in Streck BCT tubes [80].
- Centrifuge within 14 days to separate plasma (e.g., 1,600 × g for 20 min).
- Perform a second, high-speed centrifugation of the plasma (e.g., 16,000 × g for 20 min) to remove residual cells.
- Extract cfDNA from the clarified plasma using a commercial kit. Elute in a low-EDTA buffer or nuclease-free water. Quantify using a fluorometer.
Assay Design:
- Based on the prior NGS results, select one or two somatic mutations with the highest variant allele frequency in the primary tumor [80].
- Design and validate mutant-specific TaqMan probe and primer sets. A wild-type probe is included for reference.
dPCR Setup and Run:
- Prepare the PCR reaction mix containing extracted cfDNA, primers, probes, and dPCR supermix.
- Partition the reaction mixture into thousands of droplets using a droplet generator [1].
- Transfer the emulsified sample to a 96-well plate and perform PCR amplification on a thermal cycler using standard cycling conditions.
Data Analysis:
- Load the post-PCR plate onto a droplet reader, which counts the fluorescent-positive and negative droplets for each channel.
- Use the instrument's software to apply Poisson statistics and calculate the absolute concentration (copies/μL) of mutant and wild-type DNA in the original sample [1].
- Report results as mutant copies per volume or as a variant allele frequency (VAF).
Targeted NGS for Comprehensive Genomic Profiling
This protocol is designed for the broad detection of genomic alterations from plasma cfDNA.
Workflow:
Library Preparation:
- Fragment genomic DNA (if using tissue) or use naturally fragmented cfDNA.
- Repair DNA ends and ligate sequencing adapters containing Unique Molecular Identifiers (UMIs). UMIs are short random nucleotide sequences that tag individual original DNA molecules, which is critical for distinguishing true low-frequency variants from PCR/sequencing errors during data analysis [78].
- Amplify the library via limited-cycle PCR.
Target Enrichment:
- For targeted panels, use hybrid capture with biotinylated probes or amplicon-based PCR to enrich for genes of interest. This step is crucial for achieving sufficient depth on relevant genomic regions [78].
Sequencing:
- Pool multiplexed libraries and load onto a sequencer.
- Sequence to a high raw depth (e.g., >15,000x) to achieve a sufficient deduplicated depth for sensitive variant calling [78].
Bioinformatic Analysis:
- Demultiplexing: Assign reads to respective samples.
- UMI Processing: Group reads originating from the same original DNA molecule to create consensus reads, thereby reducing background noise [78].
- Alignment: Map reads to a reference human genome.
- Variant Calling: Identify somatic mutations (SNVs, Indels), copy number variations (CNVs), and gene fusions against a matched normal or a panel of normal samples. The variant calling threshold is often lowered (e.g., requiring ≥3 supporting reads) to enhance sensitivity for low-VAF variants [78].
dPCR and NGS are foundational technologies that serve distinct yet complementary roles in the ctDNA analysis ecosystem for solid tumors. NGS provides the comprehensive landscape view necessary for initial tumor profiling and discovery, while dPCR offers the precision and sensitivity required for vigilant, cost-effective longitudinal monitoring. For drug development professionals, this synergy is key: NGS can identify patient cohorts and actionable targets, while dPCR can be deployed in clinical trials to monitor patient response and the emergence of resistance with high frequency and precision. Future developments will continue to push the sensitivity limits of both technologies, particularly NGS, further closing the gap and solidifying the role of liquid biopsy as a cornerstone of precision oncology.
Digital PCR (dPCR) represents a transformative technology in oncology biomarker research, enabling the absolute quantification of circulating tumor DNA (ctDNA) with exceptional sensitivity and precision. Unlike conventional quantitative PCR, dPCR partitions samples into thousands of nanoliter-scale reactions, allowing for the detection and counting of individual DNA molecules without requiring a standard curve [1]. This capability is particularly valuable in melanoma management, where the detection of minimal residual disease (MRD) following surgical resection remains a significant clinical challenge. In resected stage III melanoma, patient outcomes vary widely, with 5-year survival rates ranging from 93% for stage IIIA to 32% for stage IIID disease [83]. The accurate identification of patients at high risk of recurrence is crucial for optimizing adjuvant therapy decisions, potentially avoiding both overtreatment and undertreatment [84] [83].
The COMBI-AD trial biomarker analysis demonstrated that droplet digital PCR (ddPCR) measurements of BRAFV600-mutant ctDNA can effectively stratify patients by recurrence risk following resection [84] [83]. This application note details the experimental protocols and analytical frameworks for implementing dPCR assays in melanoma clinical trials, providing researchers with standardized methodologies for prognostic biomarker development.
Analytical Performance of dPCR in Melanoma Detection
Technical Advantages of dPCR Platforms
Digital PCR technology offers several critical advantages for ctDNA analysis in melanoma. The partitioning process enables absolute quantification of target sequences based on Poisson statistics applied to the fraction of positive and negative partitions [1]. This approach significantly enhances sensitivity for detecting low-abundance mutations, with studies demonstrating reliable detection at variant allele frequencies as low as 0.1% [1]. Furthermore, dPCR exhibits superior tolerance to PCR inhibitors compared to other amplification-based methods, making it particularly suitable for analyzing complex biological samples like plasma and cerebrospinal fluid [3] [85].
Recent technological advancements have led to the development of multiple dPCR platforms utilizing different partitioning methods. Droplet-based systems (ddPCR) generate thousands of water-in-oil emulsion droplets, while chip-based systems employ nanowell arrays for sample compartmentalization [1]. The Table 1 below compares the key characteristics of these systems.
Table 1: Comparison of Digital PCR Partitioning Methods and Clinical Applications
| Partitioning Method | Representative Platforms | Partition Characteristics | Throughput | Clinical Utility in Melanoma |
|---|---|---|---|---|
| Droplet-based | Bio-Rad QX200, Naica | 20,000 droplets of ~0.8 nL | High | High-sensitivity BRAF mutation detection in plasma [84] |
| Chip-based | Fluidigm Biomark, Qiagen QIAcuity | Fixed microchambers of ~6 nL | Medium | Multiplexed miRNA ratio quantification [3] |
| Capillary-based | Early prototype systems | ~10 nL volume capillaries | Low | Research applications |
Comparison with Alternative Quantification Methods
While next-generation sequencing (NGS) provides comprehensive mutation profiling, it typically offers semi-quantitative data based on variant allele frequency (VAF) that can be influenced by fluctuations in non-tumor cell-free DNA [86]. In contrast, dPCR delivers absolute quantification of specific mutations, making it ideally suited for longitudinal monitoring of known biomarkers. A novel quantitative NGS (qNGS) approach incorporating unique molecular identifiers (UMIs) and quantification standards (QSs) has recently been developed to enable absolute quantification while maintaining broad mutation coverage [86]. This method demonstrated strong correlation with dPCR in validation studies, suggesting a potential complementary role for these technologies in comprehensive ctDNA analysis [86].
Clinical Validation: COMBI-AD Trial Biomarker Analysis
Study Design and Patient Cohort
The COMBI-AD trial was a double-blind, randomized, phase 3 study investigating adjuvant dabrafenib plus trametinib versus matched placebos in patients with resected BRAFV600-mutant stage III melanoma [84] [83]. The biomarker analysis included 597 of 870 patients (69%) with available baseline plasma samples collected after resection but before initiating adjuvant therapy [83]. Additional longitudinal samples were collected at 3, 6, 9, and 12 months during follow-up, with a subset of samples available from patients at the time of recurrence [84].
Analytically validated mutation-specific ddPCR assays were developed to detect BRAFV600E or BRAFV600K mutations in plasma ctDNA [83]. The assays demonstrated robust performance characteristics, including high sensitivity and specificity for detecting low-frequency mutations in cell-free DNA backgrounds.
Key Findings and Clinical Implications
The COMBI-AD biomarker analysis yielded several critical findings establishing the prognostic utility of ctDNA detection:
Baseline ctDNA Detection: ctDNA was detectable in 79 (13%) of 597 baseline samples, with positivity rates and mutant copy numbers significantly higher in patients with advanced disease substages [84] [83].
Survival Outcomes: Baseline ctDNA detection was strongly associated with worse recurrence-free survival (RFS) and overall survival (OS) in both placebo and combination therapy groups [83]. The magnitude of this association is detailed in Table 2.
Longitudinal Monitoring: Patients with adverse ctDNA kinetics (molecular relapse or persistently positive) had markedly shorter median RFS compared to those with favorable kinetics (undetectable after positive baseline or durable undetectable) [84].
Table 2: Association Between Baseline ctDNA Detection and Survival Outcomes in COMBI-AD Trial
| Patient Group | ctDNA Status | Median RFS (months) | Hazard Ratio (95% CI) | Median OS (months) | Hazard Ratio (95% CI) |
|---|---|---|---|---|---|
| Placebo | Detectable | 3.71 | 2.91 (1.99-4.25) | 33.90 | 3.35 (2.01-5.55) |
| Placebo | Undetectable | 24.41 | Reference | Not reached | Reference |
| Combination Therapy | Detectable | 16.59 | 2.98 (1.95-4.54) | 40.31 | 4.27 (2.50-7.27) |
| Combination Therapy | Undetectable | 68.11 | Reference | Not reached | Reference |
The prognostic value of baseline ctDNA concentration remained significant after adjusting for established risk factors, including disease substage, and demonstrated stronger association with outcomes than tissue-based biomarkers like tumor mutational burden or interferon gamma gene expression [83].
Experimental Protocols and Methodologies
ddPCR Assay Protocol for BRAF-Mutant ctDNA Detection
The following protocol details the analytically validated ddPCR methodology used in the COMBI-AD trial for detecting BRAFV600 mutations in plasma ctDNA:
Sample Collection and Processing
- Collect whole blood in cell-stabilizing blood collection tubes (e.g., Streck Cell-Free DNA BCT)
- Process within 6 hours of collection: centrifuge at 1600×g for 10 minutes at 4°C, followed by plasma separation and second centrifugation at 16,000×g for 10 minutes to remove cellular debris
- Store plasma at -80°C until DNA extraction
- Extract cell-free DNA from 2-4 mL plasma using commercially available kits (e.g., QIAamp Circulating Nucleic Acid Kit)
- Elute DNA in 20-50 μL elution buffer [84] [83]
ddPCR Reaction Setup
- Prepare reaction mix containing:
- 10 μL ddPCR Supermix for Probes (no dUTP)
- 1 μL BRAFV600E or BRAFV600K mutation-specific assay (FAM-labeled)
- 1 μL reference assay (HEX-labeled) for normalization
- 5-8 μL template DNA (up to 22 μL total reaction volume)
- Generate droplets using Automated Droplet Generator (target: 20,000 droplets per sample)
- Transfer droplets to 96-well PCR plate and seal [84]
PCR Amplification and Analysis
- Amplify using thermal cycling conditions:
- 95°C for 10 minutes (enzyme activation)
- 40 cycles of: 94°C for 30 seconds, 55-60°C for 60 seconds
- 98°C for 10 minutes (enzyme deactivation)
- Read plate on Droplet Reader
- Analyze data using quantification software (e.g., QuantaSoft)
- Determine mutant copies per mL plasma using Poisson statistics [84] [83]
Duplex dPCR Protocol for miRNA Ratio Quantification
For simultaneous detection of multiple biomarkers, a duplex dPCR assay was developed for miR-4488 and miR-579-3p ratio (miRatio) quantification:
RNA Extraction and cDNA Synthesis
- Extract total RNA from 200 μL serum using miRNeasy Mini Kit
- Use fixed input volume of 2 μL total RNA for reverse transcription (due to typically low serum RNA concentrations)
- Perform reverse transcription using TaqMan Advanced miRNA cDNA Synthesis Kit with preamplification step [3]
Duplex dPCR Setup
- Prepare reaction mix containing:
- 10 μL dPCR Supermix
- 0.5 μL each of miR-4488 and miR-579-3p TaqMan assays (different fluorescent dyes)
- 1 μL cDNA template
- Nuclease-free water to 20 μL total volume
- Generate partitions according to platform-specific protocols
- Amplify with thermal cycling:
- 95°C for 10 minutes
- 40 cycles of 95°C for 30 seconds and 60°C for 60 seconds
- Analyze endpoint fluorescence to calculate absolute copies/μL for each miRNA
- Compute miRatio as expression ratio between miR-4488 and miR-579-3p [3]
Visualization of Experimental Workflows
dPCR Workflow for ctDNA Analysis
Biomarker Kinetics and Clinical Outcomes
Research Reagent Solutions
Table 3: Essential Research Reagents for dPCR-Based Melanoma Biomarker Studies
| Reagent Category | Specific Products | Application Notes | Clinical Trial Validation |
|---|---|---|---|
| Blood Collection Tubes | Streck Cell-Free DNA BCT | Preserves ctDNA integrity during transport and storage | COMBI-AD trial [84] |
| Nucleic Acid Extraction | QIAamp Circulating Nucleic Acid Kit | Optimized for low-abundance cfDNA from plasma/serum | COMBI-AD trial [84] |
| dPCR Master Mixes | ddPCR Supermix for Probes (Bio-Rad) | Enables precise partitioning and amplification | COMBI-AD trial [84] |
| Mutation-Specific Assays | BRAFV600E/V600K ddPCR assays | FAM-labeled mutant probes with HEX-labeled reference | COMBI-AD trial [84] |
| miRNA Analysis | TaqMan Advanced miRNA cDNA Synthesis Kit | Includes preamplification for low-input samples | Metastatic melanoma study [3] |
| Reference Assays | Reference genes for normalization | Ensures technical reproducibility across runs | COMBI-AD trial [84] |
The clinical validation of dPCR assays in the COMBI-AD trial establishes ctDNA as a powerful prognostic biomarker in resected stage III melanoma, enabling identification of high-risk patients who may benefit from treatment intensification and low-risk patients who might avoid unnecessary therapy [84] [83]. The absolute quantification capabilities of dPCR provide critical advantages for longitudinal monitoring of minimal residual disease and early detection of molecular relapse.
Future applications of dPCR in melanoma clinical trials include guiding adjuvant therapy escalation or de-escalation based on ctDNA status, monitoring novel combination therapies, and expanding to rare melanoma subtypes such as uveal and mucosal melanoma [87] [85]. Additionally, the development of multiplex dPCR assays for simultaneous detection of multiple biomarkers, including miRNA ratios and protein biomarkers, will further enhance the precision of prognostic stratification [3] [86].
As dPCR technology continues to evolve with improved multiplexing capabilities and streamlined workflows, its integration into standardized clinical trial protocols will accelerate the development of personalized treatment approaches and ultimately improve outcomes for melanoma patients across disease stages.
Digital PCR (dPCR) represents a paradigm shift in nucleic acid quantification for diagnostic workflows, offering absolute quantification without the need for standard curves. As the third generation of PCR technology, it builds upon conventional PCR and quantitative real-time PCR (qPCR) by partitioning a sample into thousands of individual reactions, enabling precise molecular counting at the single-molecule level [1]. This capability is particularly valuable in oncology research, where detecting rare mutations and quantifying subtle genetic variations can directly impact clinical decision-making and therapeutic strategies [13].
The fundamental principle underlying dPCR involves partitioning a PCR mixture into numerous nanoscale reactions so that each compartment contains either zero, one, or a few nucleic acid targets according to Poisson distribution [1]. Following end-point amplification, the fraction of positive partitions is counted, allowing absolute quantification of the target molecule based on Poisson statistics [88]. This compartmentalization approach eliminates PCR inhibition effects and provides unprecedented sensitivity for detecting minor allele frequencies as low as 0.01%, making it particularly suitable for liquid biopsy applications in cancer research [13].
Comparative Analysis of dPCR and qPCR Performance Parameters
Quantitative Performance Comparison
Selecting between dPCR and qPCR technologies requires careful consideration of multiple performance and operational parameters tailored to specific application needs. The following table summarizes key comparative metrics based on current technological capabilities.
Table 1: Performance and operational comparison between dPCR and qPCR
| Parameter | Digital PCR (dPCR) | Quantitative PCR (qPCR) |
|---|---|---|
| Quantification Method | Absolute quantification without standard curves [1] | Relative quantification requiring standard curve [89] |
| Sensitivity | High (detects minor alleles to 0.01%) [13] | Moderate [89] |
| Precision at Low Concentrations | High [13] | Moderate to low [89] |
| Throughput | Lower [89] | High [89] |
| Multiplexing Capability | Easier multiplexing for reference genes and multiple targets [13] | Challenging, especially for multiple targets [89] |
| Cost per Test | Higher [88] [89] | Lower (e.g., $0.2 per test for COVID-19 pooling) [89] |
| Effect of PCR Inhibitors | Reduced impact due to compartmentalization [1] | Significant impact [89] |
| Operational Complexity | Higher, requires specialized personnel [88] | Lower, widely standardized [89] |
| Sample Input Requirement | Lower volume [1] | Higher volume [89] |
Cost-Benefit Considerations for Diagnostic Implementation
The implementation of dPCR in diagnostic workflows presents a distinct cost-benefit profile that must be carefully evaluated against clinical and research requirements. The initial investment for dPCR instrumentation and specialized consumables significantly exceeds that of qPCR systems, creating a substantial barrier for resource-limited settings [88]. Additionally, the operational costs for dPCR remain elevated due to specialized reagents and the need for technical expertise in microfluidics and data interpretation [88]. These factors contribute to a higher cost per test compared to established qPCR methodologies [89].
Despite these economic challenges, dPCR offers compelling analytical benefits that can justify its implementation in specific clinical scenarios. The technology's superior sensitivity and precision for detecting rare mutations provide exceptional value in applications such as minimal residual disease monitoring, therapy resistance mutation detection, and liquid biopsy development [13]. Furthermore, the absolute quantification capability eliminates inter-laboratory variability associated with standard curve generation in qPCR, potentially improving reproducibility across diagnostic networks [1]. The reduced sample input requirements of dPCR also present a significant advantage when working with precious biological specimens, such as circulating tumor DNA or fine-needle aspiration biopsies, where material is limited [1].
Experimental Protocol for dPCR-Based Liquid Biopsy Analysis in Oncology
Sample Preparation and DNA Extraction
Proper sample preparation is critical for successful dPCR analysis of liquid biopsy samples. The following protocol outlines a standardized approach for circulating tumor DNA (ctDNA) analysis:
Blood Collection and Plasma Separation: Collect 10-20 mL of peripheral blood in cell-free DNA collection tubes. Process samples within 2-6 hours of collection. Centrifuge at 1600 × g for 10 minutes at 4°C to separate plasma from cellular components. Transfer the supernatant to a fresh tube and perform a second centrifugation at 16,000 × g for 10 minutes to remove residual cells [1].
Cell-Free DNA Extraction: Extract cell-free DNA from 1-5 mL of plasma using a commercial cell-free DNA extraction kit following manufacturer's instructions. Elute DNA in 20-50 μL of elution buffer suitable for PCR amplification. Store extracts at -80°C if not used immediately.
DNA Quantification and Quality Assessment: Quantify DNA using a fluorescence-based method appropriate for low-concentration samples. Avoid spectrophotometric methods due to limited sensitivity. Assess DNA fragmentation profile using a microfluidic capillary electrophoresis system if available.
dPCR Reaction Setup and Partitioning
The dPCR workflow involves careful reaction preparation and partitioning to ensure accurate absolute quantification:
Reaction Mixture Preparation: Prepare a 20-40 μL reaction mixture containing:
- 1X dPCR master mix (commercial formulation optimized for dPCR)
- 900 nM forward and reverse primers
- 250 nM hydrolysis probes (FAM and VIC labeled for multiplex detection)
- 5-20 ng of cell-free DNA extract
- Nuclease-free water to final volume
Partitioning Generation: Depending on the dPCR platform:
- Droplet-based systems: Generate 20,000 droplets using an automated droplet generator according to manufacturer's specifications. Ensure uniform droplet size and stability before amplification.
- Chip-based systems: Load the reaction mixture onto the microfluidic chip using automated fluidic controllers to ensure complete and uniform partition filling.
Thermal Cycling: Perform PCR amplification using the following cycling conditions:
- Initial enzyme activation: 95°C for 10 minutes
- 40-45 cycles of:
- Denaturation: 94°C for 30 seconds
- Annealing/Extension: 55-60°C for 60 seconds (optimize based on primer Tm)
- Final hold: 98°C for 10 minutes
- Endpoint fluorescence reading or stabilization at 4°C
Data Acquisition and Analysis
Post-amplification analysis enables absolute quantification of target molecules:
Fluorescence Reading: For droplet systems, read individual droplets using a flow-based detector. For chip-based systems, perform endpoint fluorescence imaging of all partitions.
Threshold Setting: Establish fluorescence thresholds for positive/negative partitions using no-template controls and positive controls. For multiplex assays, set thresholds for each channel separately.
Absolute Quantification Calculation: Calculate the original sample concentration using Poisson statistics:
- λ = -ln(1 - p), where λ is the average number of target molecules per partition and p is the fraction of positive partitions
- Account for partition volume to calculate copies per μL of original sample
- Apply Poisson correction to adjust for multiple targets per partition, especially when target concentration is high
Figure 1: dPCR Workflow for Liquid Biopsy Analysis
Essential Research Reagent Solutions for dPCR Oncology Applications
Successful implementation of dPCR in oncology research requires carefully selected reagents and materials optimized for digital amplification. The following table outlines key components and their functions in the experimental workflow.
Table 2: Essential research reagents and materials for dPCR-based oncology research
| Reagent/Material | Function | Specification Notes |
|---|---|---|
| dPCR Master Mix | Provides DNA polymerase, dNTPs, and optimized buffer for amplification | Select commercial formulations specifically designed for dPCR with minimal inhibition effects [13] |
| Hydrolysis Probes | Sequence-specific detection of target mutations | FAM, VIC, or other dye-labeled probes with appropriate quenchers; pre-designed oncology panels available [13] |
| Primer Sets | Amplification of target sequences | Optimized for high efficiency and specificity; pre-validated panels for common oncology targets available [13] |
| Droplet Generation Oil | Creates stable water-in-oil emulsion for droplet-based systems | Formulated with appropriate surfactants to prevent droplet coalescence during thermal cycling [1] |
| Microfluidic Chips/Cartridges | Physical partitioning of reactions in chip-based systems | Platform-specific designs with precise well volumes and surface treatments [1] |
| DNA Extraction Kits | Isolation of cell-free DNA from liquid biopsy samples | Optimized for low-abundance DNA recovery from plasma or serum [1] |
| Quantification Standards | Validation of assay performance and quantification accuracy | Synthetic DNA standards with known mutation status for assay validation [13] |
Clinical Utility and Application-Specific Considerations
Oncology Applications with High Clinical Utility
The unique capabilities of dPCR make it particularly valuable for specific oncology applications where its advantages provide clinically actionable information:
Minimal Residual Disease Monitoring: dPCR's ability to detect rare mutation-containing molecules (as low as 0.01%) enables sensitive tracking of residual disease following treatment. This application benefits from dPCR's precision at low concentrations, allowing clinicians to detect early recurrence before clinical manifestation [13].
Therapy Resistance Mutation Detection: The emergence of resistance mutations during targeted therapy can be identified early through serial liquid biopsy monitoring. dPCR's multiplexing capabilities allow simultaneous tracking of multiple resistance mechanisms, providing comprehensive information for therapeutic decision-making [1].
Tumor Heterogeneity Analysis: The absolute quantification provided by dPCR enables precise measurement of mutation allelic frequency, which can reveal clonal evolution and tumor heterogeneity. This information has prognostic significance and may guide combination therapy approaches [1].
Decision Framework for Technology Selection
Choosing between dPCR and qPCR requires a structured approach based on specific application requirements:
Figure 2: Decision Framework for dPCR vs qPCR Selection
Digital PCR technology represents a significant advancement in molecular diagnostics, particularly for oncology applications requiring absolute quantification of rare genetic targets. While the higher operational costs and lower throughput compared to qPCR present implementation challenges, the superior sensitivity, precision at low concentrations, and absolute quantification capabilities provide compelling benefits for specific clinical and research applications [88] [89].
The ongoing development of dPCR technologies focuses on addressing current limitations while expanding clinical utility. Future directions include increased multiplexing capabilities for parallel assessment of multiple biomarkers, integration with artificial intelligence for enhanced data analysis, and development of point-of-care platforms to make the technology more accessible [89]. Additionally, standardization of protocols and analytical validation guidelines will support broader adoption in clinical diagnostics [1].
As oncology research increasingly focuses on liquid biopsy and minimal residual disease monitoring, dPCR is poised to play an essential role in translational research and clinical diagnostics. The technology's unique ability to provide absolute quantification of rare mutations offers invaluable insights into tumor dynamics, treatment response, and disease evolution, ultimately contributing to more personalized and effective cancer management strategies.
Conclusion
Digital PCR has firmly established itself as a critical technology for absolute nucleic acid quantification in oncology research and clinical applications. Its unparalleled sensitivity and precision enable the detection of rare genetic events, such as minimal residual disease and emerging mutations in ctDNA, often months before clinical recurrence. As evidenced by robust validation studies, dPCR frequently outperforms qPCR in detecting low-abundance targets and demonstrates strong complementary utility with NGS. Future directions will focus on standardizing dPCR assays for clinical use, expanding multiplexing capabilities, and further integrating dPCR into longitudinal monitoring and interventional clinical trials. The continued adoption of dPCR promises to deepen our understanding of tumor biology and accelerate the development of truly personalized cancer management strategies.
References
- 1. Digital PCR: from early developments to its future application ... [https://pubs.rsc.org/en/content/articlehtml/2025/lc/d5lc00055f]
- 2. Digital PCR Applications: Hemato-Oncology Research [https://www.thermofisher.com/us/en/home/life-science/pcr/digital-pcr/education/cancer-research/hemato-oncology.html]
- 3. Development of an innovative duplex digital PCR assay for ... [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-025-06941-1]
- 4. Digital PCR | Thermo Fisher Scientific - US [https://www.thermofisher.com/us/en/home/life-science/pcr/digital-pcr.html]
- 5. Digital droplet PCR is an accurate and precise method to ... [https://www.nature.com/articles/s41598-025-20944-4]
- 6. Comparison of two digital PCR platforms for quantification ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12622311/]
- 7. Flexible methods for uncertainty estimation of digital PCR ... [https://www.sciencedirect.com/science/article/pii/S2589004225000318]
- 8. Reference Gene Evaluation for digital PCR; applications ... [https://www.sciencedirect.com/science/article/pii/S0166093425001612]
- 9. Three-color crystal digital PCR - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5154636/]
- 10. Poisson Plus Quantification for Digital PCR Systems [https://www.nature.com/articles/s41598-017-09183-4]
- 11. Data analysis method for multiplex digital pcr using color- ... [https://patents.google.com/patent/EP4372100A1/en]
- 12. Development of an innovative duplex digital PCR assay for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12351995/]
- 13. Digital PCR for Oncology Research [https://www.thermofisher.com/us/en/home/life-science/pcr/digital-pcr/oncology-research.html]
- 14. Comparing the precision of two digital PCR applications for ... [https://www.nature.com/articles/s41598-025-13143-8]
- 15. Digital PCR: from early developments to its future ... [https://pubs.rsc.org/en/content/articlelanding/2025/lc/d5lc00055f]
- 16. Droplet Digital : PCR and Applications in Molecular... Advantages [https://www.labx.com/resources/droplet-digital-pcr-advantages-and-applications-in-molecular-diagnostics/5581]
- 17. What Is Digital ? – Learn About the PCR of dPCR Advantages [https://www.thermofisher.com/blog/behindthebench/what-is-digital-pcr/]
- 18. Four applications of dPCR fundamental to cancer genomics [https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/science-matters/pcr-solutions/four-applications-of-dpcr-fundamental-to-cancer-genomics]
- 19. and Traditional Digital : What’s the Difference? PCR PCR [https://www.news-medical.net/life-sciences/Digital-PCR-and-Traditional-PCR-Whats-the-Difference.aspx]
- 20. Absolute vs. Relative Quantification for qPCR [https://www.thermofisher.com/us/en/home/life-science/pcr/real-time-pcr/real-time-pcr-learning-center/real-time-pcr-basics/absolute-vs-relative-quantification-real-time-pcr.html]
- 21. Comparison of Droplet Digital PCR to Real-Time ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3553899/]
- 22. Viral diagnostics in the era of digital PCR - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3519953/]
- 23. Digital PCR: A Reliable Tool for Analyzing and Monitoring ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7247671/]
- 24. Development and validation of a methylation-specific ... [https://www.nature.com/articles/s41598-025-15228-w]
- 25. Liquid biopsy – a narrative review with an update on current ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12333414/]
- 26. Digital PCR Applications: Hemato-Oncology Research [https://www.thermofisher.com/io/en/home/life-science/pcr/digital-pcr/education/cancer-research/hemato-oncology.html]
- 27. A robust ddPCR assay for the absolute quantification of ... [https://pubmed.ncbi.nlm.nih.gov/41167182/]
- 28. RainDrop Digital PCR | Yale Cancer Center [https://medicine.yale.edu/cancer/research/shared-resources/sponsored-resources-equipment/raindrop/]
- 29. Cancer in a Drop: Liquid biopsy insights from AACR 2025 [https://pmc.ncbi.nlm.nih.gov/articles/PMC12223554/]
- 30. Minimal Residual Disease Detection: Implications for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12079024/]
- 31. A comprehensive overview of minimal residual disease in ... [https://www.nature.com/articles/s41698-025-00984-9]
- 32. Minimal Residual Disease: Predicting and Preventing ... [https://www.emjreviews.com/hematology/congress-review/minimal-residual-disease-predicting-and-preventing-relapse-in-myeloma-s060225/]
- 33. A tour of leukemia progress in 2025, viewed through the MD ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12548704/]
- 34. Digital PCR Primer and Probe Design Rules - Gene Pi [https://www.gene-pi.com/item/primers-and-probes-2/]
- 35. Species-Level eDNA Quantification using Digital PCR [https://www.h2omolecular.com/2025/07/03/species-level-edna-quantification-using-digital-pcr/]
- 36. Duplex nanoplate-based digital PCR assay for ... [https://www.sciencedirect.com/science/article/abs/pii/S0889157525000602]
- 37. A digital PCR-based protocol to detect and quantify RNA ... [https://www.sciencedirect.com/science/article/pii/S2666166722000284]
- 38. Comparative Performance of Digital PCR and Real-Time RT ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12474457/]
- 39. CYP2D6 copy number determination using digital PCR [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2024.1429286/full]
- 40. Multiplexed Digital PCR Reference Gene Measurement for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12523864/]
- 41. Multiplex digital PCR enables sensitive detection of ... [https://pubmed.ncbi.nlm.nih.gov/39849159/]
- 42. VIDEO | Highly Sensitive and Specific Multiplexed Detection of ... [https://magazine.eacr.org/biorad-webinar-october-2025/]
- 43. Characterization of a multiplex digital PCR assay to ... [https://www.sciencedirect.com/science/article/pii/S2950329925000505]
- 44. Digital PCR for Solid Tumors [https://www.thermofisher.com/us/en/home/life-science/pcr/digital-pcr/oncology-research/solid-tumors.html]
- 45. Digital PCR: from early developments to its future application ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12278255/]
- 46. Past, Present and Future of Digital PCR [https://www.technologynetworks.com/diagnostics/ebooks/past-present-and-future-of-digital-pcr-405990]
- 47. Rules and Tips for PCR & qPCR Primer Design | IDT [https://www.idtdna.com/page/support-and-education/decoded-plus/how-to-design-primers-and-probes-for-pcr-and-qpcr/]
- 48. Real-Time PCR Master Mix Selection Guide [https://www.thermofisher.com/na/en/home/life-science/pcr/real-time-pcr/reagents-kits/selection-guide.html]
- 49. Digital PCR workflow and products [https://www.qiagen.com/us/applications/digital-pcr/workflow-and-products]
- 50. dPCR vs qPCR vs end-point PCR [https://www.qiagen.com/us/knowledge-and-support/knowledge-hub/bench-guide/pcr/digital-pcr/digital-pcr-vs-quantitative-pcr-vs-end-point-pcr]
- 51. Massive and Controllable Production of Droplets on Chips ... [https://www.sciencedirect.com/science/article/pii/S0925400525015394]
- 52. Dual production of biconvex polymer particles via ... [https://www.nature.com/articles/s41598-025-06869-y]
- 53. A comprehensive review of methodological and ... [https://www.sciencedirect.com/science/article/abs/pii/S0734975025002058]
- 54. Unveiling the cut-and-repair cycle of designer nucleases in ... [https://www.nature.com/articles/s41467-025-65182-4]
- 55. PCR and Thermal Cycler Technology: Fundamentals, ... [https://www.labx.com/resources/pcr-and-thermal-cycler-technology-fundamentals-applications-and-emerging-trends-in-dna/5577]
- 56. Development and application of a highly sensitive ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1612740/full]
- 57. dPCR: A Technology Review - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC5948698/]
- 58. NSCLC Digital PCR Panel Returns Low-Input Sample Results ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10854965/]
- 59. Reverse transcriptase droplet digital shows high resilience to... PCR [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-014-0042-6]
- 60. Getting rid of ‘rain’ and ‘stars’: Mitigating effects on ddPCR... inhibition [https://pmc.ncbi.nlm.nih.gov/articles/PMC9668142/]
- 61. Digital PCR Applications: Hemato- Oncology Research [https://www.thermofisher.com/ar/en/home/life-science/pcr/digital-pcr/education/cancer-research/hemato-oncology.html]
- 62. Digital PCR for accurate quantification of pathogens: Principles... [https://pubmed.ncbi.nlm.nih.gov/34171259/]
- 63. Development of a viability digital PCR protocol for the ... [https://www.nature.com/articles/s41598-019-47976-x]
- 64. Digital PCR (dPCR) And Real-Time PCR (qPCR) in the Real World... [https://www.linkedin.com/pulse/digital-pcr-dpcr-real-time-qpcr-real-world-5-uses-vqvnf]
- 65. How Digital PCR Enables Early Molecular Relapse Detection [https://www.thermofisher.com/blog/behindthebench/early-molecular-relapse-detection-dpcr/]
- 66. Droplet Digital PCR: A Powerful Tool for Accurate ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12080313/]
- 67. Digital PCR outperforms quantitative real-time PCR for the ... [https://pubmed.ncbi.nlm.nih.gov/40708854/]
- 68. Droplet digital PCR assay for precise determination of FRS2 ... [https://bmccancer.biomedcentral.com/articles/10.1186/s12885-025-14611-0]
- 69. In-house validation of a droplet digital PCR system using ... [https://www.sciencedirect.com/science/article/pii/S0003267025005665]
- 70. QIAGEN expands digital PCR oncology research portfolio ... [https://corporate.qiagen.com/English/newsroom/press-releases/press-release-details/2025/QIAGEN-expands-digital-PCR-oncology-research-portfolio-through-partnership-with-ID-Solutions/default.aspx]
- 71. dPCR vs. ddPCR: Why Precision Matters in Cell and Gene ... [https://www.roslinct.com/insights/dpcr-vs-ddpcr/]
- 72. Digital PCR outperforms quantitative real-time PCR for the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12288185/]
- 73. Comparison of qRT-PCR and ddPCR for multi-strain ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1579797/full]
- 74. Development and validation of a droplet digital PCR assay ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1573949/full]
- 75. Development and Validation of One-Step Reverse ... [https://www.mdpi.com/2223-7747/13/23/3276]
- 76. An adjusted droplet digital PCR assay for quantification of ... [https://www.sciencedirect.com/science/article/pii/S2590018824003289]
- 77. Revolution of Circulating Tumor DNA: From Bench ... [https://www.mdpi.com/1467-3045/47/6/428]
- 78. Real-World Technical Hurdles of ctDNA NGS Analysis [https://pmc.ncbi.nlm.nih.gov/articles/PMC12564474/]
- 79. Circulating tumor DNA to monitor treatment response in ... [https://www.nature.com/articles/s41698-025-00876-y]
- 80. Performance Comparison of Droplet Digital PCR and Next ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12062871/]
- 81. Validation of a liquid biopsy assay with increased ... [https://www.sciencedirect.com/science/article/pii/S2950195425000384]
- 82. Performance Comparison of Droplet Digital PCR and Next- ... [https://pubmed.ncbi.nlm.nih.gov/40346007/]
- 83. Articles Clinical validation of droplet digital PCR assays ... [https://www.sciencedirect.com/science/article/abs/pii/S1470204525001391]
- 84. Clinical validation of droplet digital PCR assays in ... [https://pubmed.ncbi.nlm.nih.gov/40250457/]
- 85. Leptomeningeal disease in melanoma - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC12292448/]
- 86. Absolute Quantification of Nucleotide Variants in Cell-Free ... [https://www.mdpi.com/2072-6694/17/5/783]
- 87. Melanoma Clinical Trials [https://www.mayo.edu/research/clinical-trials/diseases-conditions/melanoma]
- 88. The Role of Digital PCR in Enhancing Diagnostic Power ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11311727/]
- 89. Quantitative or digital PCR? A comparative analysis for ... [https://www.sciencedirect.com/science/article/abs/pii/S0165993624001584]